Journal of Hazardous Materials ( IF 12.2 ) Pub Date : 2021-05-05 , DOI: 10.1016/j.jhazmat.2021.126027 Sai Xu 1 , Muhammad Zeeshan Qasim 2 , Tao Zhang 3 , Ruyue Wang 2 , Chao Li 4 , Shijian Ge 2
|
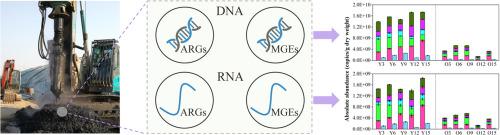
Landfills are the hotspots for the occurrence of antibiotic resistance genes (ARGs) in the environment. However, limited information is available on the profile of ARGs in response to the varying age of refuse in landfills. In this study, the diversity, abundance and expression of ARGs in a Chinese landfill were assessed by high-throughput quantitative PCR. A total of 154 ARGs were detected and 66% of them were transcriptionally active. The total abundance of ARG transcripts was one magnitude lower than that of ARGs. The ermT-01, tetX, sul2, aadA-02 and aadA2–03 genes were found to be the most abundant ARGs (ARG transcripts) and their sum abundance showed a linear relation with the total abundance of ARGs (ARG transcripts). The total abundance of ARGs (ARG transcripts) in young refuse was significantly higher than that in old refuse (p < 0.01) and the profile of ARGs (ARG transcripts) between the old and young refuse was distinct as revealed by the principal coordinates analysis. The variation partitioning analysis showed heavy metals (mainly Cr and Zn) were the major drivers that affect the profile of ARGs (ARG transcripts). These findings provided new insights into the ARGs in landfills and indicated their potential threats should not be neglected.











































 京公网安备 11010802027423号
京公网安备 11010802027423号