Current Bioinformatics ( IF 2.4 ) Pub Date : 2021-01-31 , DOI: 10.2174/1574893615999200601122328 Shulin Zhao 1 , Ying Ju 2 , Xiucai Ye 3 , Jun Zhang 4 , Shuguang Han 5
|
Background: Bioluminescence is a unique and significant phenomenon in nature. Bioluminescence is important for the lifecycle of some organisms and is valuable in biomedical research, including for gene expression analysis and bioluminescence imaging technology. In recent years, researchers have identified a number of methods for predicting bioluminescent proteins (BLPs), which have increased in accuracy, but could be further improved.
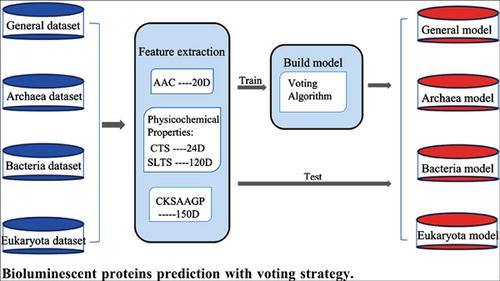
Methods: In this study, a new bioluminescent proteins prediction method, based on a voting algorithm, is proposed. Four methods of feature extraction based on the amino acid sequence were used. 314 dimensional features in total were extracted from amino acid composition, physicochemical properties and k-spacer amino acid pair composition. In order to obtain the highest MCC value to establish the optimal prediction model, a voting algorithm was then used to build the model. To create the best performing model, the selection of base classifiers and vote counting rules are discussed.
Results and Conclusion: The proposed model achieved 93.4% accuracy, 93.4% sensitivity and 91.7% specificity in the test set, which was better than any other method. A previous prediction of bioluminescent proteins in three lineages was also improved using the model building method, resulting in greatly improved accuracy.











































 京公网安备 11010802027423号
京公网安备 11010802027423号