当前位置:
X-MOL 学术
›
Mol. Genet. Genomic Med.
›
论文详情
Our official English website, www.x-mol.net, welcomes your
feedback! (Note: you will need to create a separate account there.)
Next‐generation sequencing identifies rare pathogenic and novel candidate variants in a cohort of Chinese patients with syndromic or nonsyndromic hearing loss
Molecular Genetics & Genomic Medicine ( IF 1.5 ) Pub Date : 2020-10-23 , DOI: 10.1002/mgg3.1539 Yan-Bao Xiang 1 , Chen-Yang Xu 1 , Yun-Zhi Xu 1 , Huan-Zheng Li 1 , Li-Li Zhou 1 , Xue-Qin Xu 1 , Zi-Hui Chen 2 , Shao-Hua Tang 1, 2
Molecular Genetics & Genomic Medicine ( IF 1.5 ) Pub Date : 2020-10-23 , DOI: 10.1002/mgg3.1539 Yan-Bao Xiang 1 , Chen-Yang Xu 1 , Yun-Zhi Xu 1 , Huan-Zheng Li 1 , Li-Li Zhou 1 , Xue-Qin Xu 1 , Zi-Hui Chen 2 , Shao-Hua Tang 1, 2
Affiliation
|
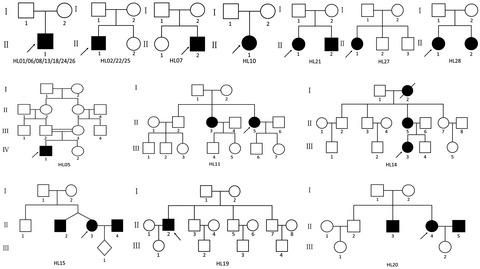
Hearing loss (HL) is a common sensory disorder in humans characterized by extreme clinical and genetic heterogeneity. In recent years, next‐generation sequencing (NGS) technologies have proven to be highly effective and powerful tools for population genetic studies of HL. Here, we analyzed clinical and molecular data from 21 Chinese deaf families who did not have hotspot mutations in the common deafness genes GJB2, SLC26A4, GJB3, and MT‐RNR1.
更新日期:2020-12-27











































 京公网安备 11010802027423号
京公网安备 11010802027423号