Journal of the American Society for Mass Spectrometry ( IF 3.1 ) Pub Date : 2018-10-15 , DOI: 10.1007/s13361-018-2073-0 Måns Ekelöf 1 , Kenneth P. Garrard 1 , Rika Judd 2 , Elias P. Rosen 3 , De-Yu Xie 2 , Angela D. M. Kashuba 3 , David C. Muddiman 1, 2, 4
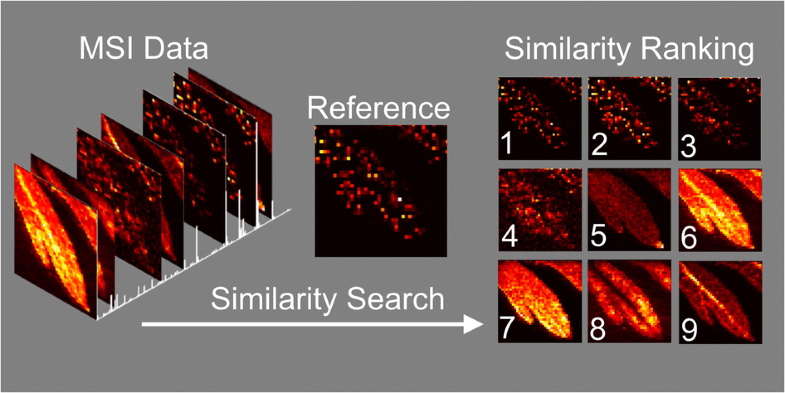
Analyzing mass spectrometry imaging data can be laborious and time consuming, and as the size and complexity of datasets grow, so does the need for robust automated processing methods. We here present a method for comprehensive, semi-targeted discovery of molecular distributions of interest from mass spectrometry imaging data, using widely available image similarity scoring algorithms to rank images by spatial correlation. A fast and powerful batch search method using a MATLAB implementation of structural similarity (SSIM) index scoring with a pre-selected reference distribution is demonstrated for two sample imaging datasets, a plant metabolite study using Artemisia annua leaf, and a drug distribution study using maraviroc-dosed macaque tissue.
ᅟ
中文翻译:

用于质谱成像数据分析的数字图像识别方法的评估
分析质谱成像数据可能既费力又费时,并且随着数据集的大小和复杂性的增长,对健壮的自动化处理方法的需求也越来越大。我们在这里提出一种用于从质谱成像数据中全面,半目标地发现目标分子分布的方法,该方法使用广泛可用的图像相似性评分算法通过空间相关性对图像进行排名。针对两个样本成像数据集,使用青蒿叶的植物代谢物研究和使用maraviroc进行药物分布研究,展示了使用MATLAB实现的结构相似性(SSIM)指数评分和预选参考分布的快速强大的批量搜索方法猕猴组织。

ᅟ











































 京公网安备 11010802027423号
京公网安备 11010802027423号