Abstract
The cellular machinery regulating microRNA biogenesis and maturation relies on a small number of simple steps and minimal biological requirements and is broadly conserved in all eukaryotic cells. The same holds true in disease. This allows for a substantial degree of freedom in the engineering of transgenes capable of simultaneously expressing multiple microRNAs of choice, allowing a more comprehensive modulation of microRNA landscapes, the study of their functional interaction, and the possibility of using such synergism for gene therapy applications. We have previously engineered a transgenic cluster of functionally associated microRNAs to express a module of suppressed microRNAs in brain cancer for therapeutic purposes. Here, we provide a detailed protocol for the design, cloning, delivery, and utilization of such artificial microRNA clusters for gene therapy purposes. In comparison with other protocols, our strategy effectively decreases the requirements for molecular cloning, because the nucleic acid sequence encoding the combination of the desired microRNAs is designed and validated in silico and then directly synthesized as DNA that is ready for subcloning into appropriate delivery vectors, for both in vitro and in vivo use. Sequence design and engineering require 4–5 h. Synthesis of the resulting DNA sequence requires 4–6 h. This protocol is quick and flexible and does not require special laboratory equipment or techniques, or multiple cloning steps. It can be easily executed by any graduate student or technician with basic molecular biology knowledge.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data that support the findings of this study but are not directly available within the paper and its Supplementary Methods files are available from the corresponding author upon reasonable request. This includes DNA sequences of all transgenes used in this study.
References
Wu, Y. E., Parikshak, N. N., Belgard, T. G. & Geschwind, D. H. Genome-wide, integrative analysis implicates microRNA dysregulation in autism spectrum disorder. Nat. Neurosci. 19, 1463–1476 (2016).
Esteller, M. Non-coding RNAs in human disease. Nat. Rev. Genet. 12, 861–874 (2011).
Moradifard, S., Hoseinbeyki, M., Ganji, S. M. & Minuchehr, Z. Analysis of microRNA and gene expression profiles in Alzheimer’s Disease: a meta-analysis approach. Sci. Rep. 8, 4767 (2018).
Calin, G. A. et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 99, 15524–15529 (2002).
Croce, C. M. Causes and consequences of microRNA dysregulation in cancer. Nat. Rev. Genet. 10, 704–714 (2009).
Ambros, V. microRNAs: tiny regulators with great potential. Cell 107, 823–826 (2001).
Treiber, T., Treiber, N. & Meister, G. Regulation of microRNA biogenesis and its crosstalk with other cellular pathways. Nat. Rev. Mol. Cell Biol. 20, 5–20 (2019).
Krek, A. et al. Combinatorial microRNA target predictions. Nat. Genet. 37, 495–500 (2005).
He, L. et al. A microRNA polycistron as a potential human oncogene. Nature 435, 828–833 (2005).
Barroso-del Jesus, A., Lucena-Aguilar, G. & Menendez, P. The miR-302-367 cluster as a potential stemness regulator in ESCs. Cell Cycle 8, 394–398 (2009).
Landais, S., Landry, S., Legault, P. & Rassart, E. Oncogenic potential of the miR-106-363 cluster and its implication in human T-cell leukemia. Cancer Res. 67, 5699–5707 (2007).
Kunjapur, A. M., Pfingstag, P. & Thompson, N. C. Gene synthesis allows biologists to source genes from farther away in the tree of life. Nat. Commun. 9, 4425 (2018).
Bhaskaran, V. et al. The functional synergism of microRNA clustering provides therapeutically relevant epigenetic interference in glioblastoma. Nat. Commun. 10, 442 (2019).
Jin, H. Y. et al. Transfection of microRNA mimics should be used with caution. Front. Genet. 6, 340 (2015).
Beg, M. S. et al. Phase I study of MRX34, a liposomal miR-34a mimic, administered twice weekly in patients with advanced solid tumors. Invest. New Drugs 35, 180–188 (2017).
Van Zandwijk, N. et al. Safety and activity of microRNA-loaded minicells in patients with recurrent malignant pleural mesothelioma: a first-in-man, phase 1, open label, dose-escalation study. Lancet Oncol. 18, 1386–1396 (2017).
Liu, Y. P., Haasnoot, J., Brake, O., Berkhout, B. & Konstantinova, P. Inhibition of HIV-1 by multiple shRNAs expressed from a single microRNA polycistron. Nucleic Acid Res. 36, 2811–2824 (2008).
Chen, S. C., Stern, P., Guo, Z. & Chen, J. Expression of multiple artificial microRNAs from a chicken miRNA126-based lentiviral vector. PLoS ONE 6, e22437 (2011).
Askou, A. L. et al. Multigenic lentiviral vectors for combined and tissue-specific expression of miRNA and protein-based antiangiogenic factors. Mol. Ther. Methods Cin. Dev. 2, 14064 (2015).
Yang, X., Marcucci, K., Anguela, X. & Couto, L. B. Preclinical evaluation of an anti-HCV miRNA cluster for treatment of HCV infection. Mol. Ther. 21, 588–601 (2013).
Wang, T., Xie, Y., Tan, A., Li, S. & Xie, Z. Construction and characterization of a synthetic microRNA cluster for multiplex RNA interference in mammalian cells. ACS Synth. Biol. 5, 1193–1200 (2016).
Brate, J. et al. Unicellular origin of the animal microRNA machinery. Curr. Biol. 28, 3288–3295.e5 (2018).
Alexopoulou, A. N., Couchman, J. R. & White, J. R. The CMV early enhancer/chicken β actin (CAG) promoter can be used to drive transgene expression during the differentiation of murine embryonic stem cells into vascular progenitors. BMC Cell Biol. 9, 2 (2008).
ZenG, Y. & Cullen, B. R. Efficient processing of primary microRNA Hairpins by Drosha requires flanking non structured RNA sequences. J. Biol. Chem. 280, 27595–27603 (2005).
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 125, 887–901 (2006).
Basu, S., Campbell, H. M., Dittel, B. N. & Ray, A. Purification of specific cell population by fluorescence activated cell sorting (FACS). J. Vis. Exp. 41, 1546 (2010).
Chiba, Y. Kaplan-Meier curves for survivor causal effects with time-to-event outcomes. Clin. Trials 10, 515–521 (2013).
Zhang, N. et al. Engineering artificial microRNAs for multiplex gene silencing and simplified transgenic screen. Plant Physiol. 178, 989–1001 (2018).
Acknowledgements
Human primary glioma stem-like cells (GSCs) (GBM34 and GBM30) were kindly donated by E. A. Chiocca’s laboratory at Brigham and Women’s Hospital. This work was supported by NINDS grant K08NS101091 to P.P.
Author information
Authors and Affiliations
Contributions
V.B. performed the experiments, analyzed the data, crafted the figures, and helped with manuscript preparation. Y.Y. and F.B. produced the AAV vectors. P.P. conceived the protocol, designed the transgenes, analyzed the data, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Carlo Croce and the other, anonymous, reviewer(s) for their contribution to the peer review of this work
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related link
Key reference using this protocol
Bhaskaran, V. et al. Nat. Commun. 10, 442 (2019): https://doi.org/10.1038/s41467-019-08390-z
Integrated supplementary information
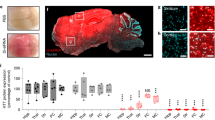
Supplementary Fig. 1 Histologic analysis of AAV-infected brain tumors.
Representative confocal microscope images of intracranial human GBM in nude mice three days after stereotactic intratumoral inoculation of 5x109 AAV particles encoding Cluster 3 transgene. In the upper row are displayed images at 1x magnification (scale bar, 1 mm); In the lower row is displayed the 20x magnification of the corresponding white boxed section (scale bar, 100 μm). Hematoxylin and Eosin (H&E) stain shows the brain/tumor interface, and the absence of grossly evident brain tissue necrosis or damage after virus infection. Infected cells are Green Fluorescent Protein (GFP)-positive. Tumor cells are Red Fluorescent Protein (RFP)-positive. Yellow line denotes the edges of the entire sectioned brain in the GFP and RFP channels.
Supplementary Fig. 2 Temporal pattern of transgenic microRNA expression in vivo.
Real Time RT-PCR quantification of mature microRNA expression in human G30 GBMs recovered from mice brains at the times labelled in the X axis. Day 7 and Day 19 refers to number of days after the intracranial infection with the AAV vectors. Control refers to ex-vivo tumors infected with control AAV expressing scrambled microRNA transgene and analyzed at Day 7 after infection. Represented are Mean ± SD from n=3 independent experiments. *** = p<0.001 (Student’s t-test, 2 tails).
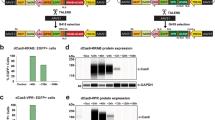
Supplementary Fig. 3 Strategy to verify transgene expression.
a: Cartoon schematizing the genomic configuration of the lentiviral vector used to overexpress microRNAs. The GFP transgene is downstream of an EF1-responsive element, while the Cluster 3 primary transcript is downstream of the CMV promoter. Colored arrows represent primer sequences used for the RT-qPCR. Green arrows represent primers for the GFP sequence. Black arrows represent primers for the microRNAs cluster transgene (note that the forward primer is in the 5’ flanking region, while the reverse primer is in the miR-128 hairpin sequence). b: Relative quantification of microRNA expression at different timepoints after intracranial implantation. G34 cells previously implanted intracranially in athymic mice were isolated from the brain at time of mouse euthanasia (either at day 12 or at day 25) and the expression of cluster 3 transgene was measured against that of cells expressing negative control and against parental Cluster 3 cells at time of implantation (day 1). Reported are Mean ± SD from three independent experiments. c: RT-qPCR showing expression of GFP transgene from cells in panel b. d: RT-qPCR showing expression of primary Cluster 3 sequence from cells in panel b. For both c and d, reported are mean + SE from one representative experiment in technical triplicates. *=p<0.05; ***=p<0.001 (Student’s t-test, 2 tails). While the expression of GFP remains stable across all time points, the expression of the microRNA transgene decreases over time, reflecting the observed progressive decrease in mature microRNA overexpression. (Adapted from Bhaskaran et al.13 under a Creative Commons Attribution 4.0 license (https://creativecommons.org/licenses/by/4.0/legalcode).
Supplementary Fig. 4 Effect of microRNA clusters on common targets.
Representative western blot from G34 GBM cells after stable expression of either negative control, single microRNAs or their combination by lentiviral infection. SP1 and JAG1 are GBM-relevant oncogenes that are independently targeted by each one of the three microRNAs. While each microRNA, as expected, decreases the level of both proteins, the combination of microRNAs does not further increase the downregulation obtained by single targeting. (Adapted from Bhaskaran et al.13 under a Creative Commons Attribution 4.0 license (https://creativecommons.org/licenses/by/4.0/legalcode).
Supplementary information
Supplementary Information
Supplementary Figures 1–4 and Supplementary Methods
Rights and permissions
About this article
Cite this article
Bhaskaran, V., Yao, Y., Bei, F. et al. Engineering, delivery, and biological validation of artificial microRNA clusters for gene therapy applications. Nat Protoc 14, 3538–3553 (2019). https://doi.org/10.1038/s41596-019-0241-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0241-8
This article is cited by
-
Variants of the adeno-associated virus serotype 9 with enhanced penetration of the blood–brain barrier in rodents and primates
Nature Biomedical Engineering (2022)
-
Circular RNAs: characteristics, biogenesis, mechanisms and functions in liver cancer
Journal of Hematology & Oncology (2021)
-
Ensuring accurate resource identification
Nature Protocols (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.