Abstract
Key message
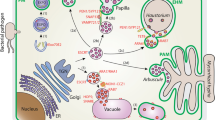
Both OsIPPI1 and OsIPPI2 enzymes are found in the endoplasmic reticulum, providing novel important insights into the role of this compartment in the synthesis of MVA pathway isoprenoids.
Abstract
Isoprenoids are synthesized from the precursor’s isopentenyl diphosphate (IPP) and dimethylallyl diphosphosphate (DMAPP), which are interconverted by the enzyme isopentenyl diphosphate isomerase (IPPI). Many plants express multiple isoforms of IPPI, the only enzyme shared by the mevalonate (MVA) and non-mevalonate (MEP) pathways, but little is known about their specific roles. Rice (Oryza sativa) has two IPPI isoforms (OsIPPI1 and OsIPPI2). We, therefore, carried out a comprehensive comparison of IPPI gene expression, protein localization, and isoprenoid biosynthesis in this species. We found that OsIPPI1 mRNA was more abundant than OsIPPI2 mRNA in all tissues, and its expression in de-etiolated leaves mirrored the accumulation of phytosterols, suggesting a key role in the synthesis of MVA pathway isoprenoids. We investigated the subcellular localization of both isoforms by constitutively expressing them as fusions with synthetic green fluorescent protein. Both proteins localized to the endoplasmic reticulum (ER) as well as peroxisomes and mitochondria, whereas only OsIPPI2 was detected in plastids, due to an N-terminal transit peptide which is not present in OsIPPI1. Despite the plastidial location of OsIPPI2, the expression of OsIPPI2 mRNA did not mirror the accumulation of chlorophylls or carotenoids, indicating that OsIPPI2 may be a redundant component of the MEP pathway. The detection of both OsIPPI isoforms in the ER indicates that DMAPP can be synthesized de novo in this compartment. Our work shows that the ER plays an as yet unknown role in the synthesis of MVA-derived isoprenoids, with important implications for the metabolic engineering of isoprenoid biosynthesis in higher plants.







Similar content being viewed by others
References
Bannai H, Tamada Y, Maruyama O, Nakai K, Miyano S (2002) Extensive feature detection of N-terminal protein sorting signals. Bioinformatics 18:298–305
Busquets A, Keim V, Closa M, del Arco A, Boronat A, Arro M, Ferrer A (2008) Arabidopsis thaliana contains a single gene encoding squalene synthase. Plant Mol Biol 67:25–36
Campos N, Boronat A (1995) Targeting and topology in the membrane of plant 3-hydroxy-3-methylglutaryl coenzyme A reductase. Plant Cell 7:2163–2174
Capell T, Christou P (2004) Progress in plant metabolic engineering. Curr Opin Biotechnol 15:148–154
Clastre M, Papon N, Courdavault V, Giglioli-Guivarc’h N, St-Pierre B, Simkin AJ (2011) Subcellular evidence for the involvement of peroxisomes in plant isoprenoid biosynthesis. Plant Signal Behav 6:2044–2046
Cunningham FX Jr, Gantt E (2000) Identification of multi-gene families encoding isopentenyl diphosphate isomerase in plants by heterologous complementation in Escherichia coli. Plant Cell Physiol 41:119–123
Davies BH (1976) Carotenoids. Chemistry and Biochemistry of Plant Pigments. Academic, London, pp 38–366
Emanuelsson O, Nielsen H, von Heijne G (1999) ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci 8:978–984
Emanuelsson O, Brunak S, von Heijne G, Nielsen H (2007) Locating proteins in the cell using TargetP, SignalP and related tools. Nat Protoc 2:953–971
Enfissi EM, Barneche F, Ahmed I, Lichtlé C, Gerrish C, McQuinn RP, Giovannoni JJ, Lopez-Juez E, Bowler C, Bramley PM, Fraser PD (2010) Integrative transcript and metabolite analysis of nutritionally enhanced DE-ETIOLATED1 downregulated tomato fruit. Plant Cell 22:1190–1215
Errampalli D, Fletcher J (1993) Production of monospecific polyclonal antibodies against aster yellows mycoplasmalike organism-associated antigen. Phytopathol 83:1279–1282
Fukasawa Y, Tsuji J, Fu SC, Tomii K, Horton P, Imai K (2015) MitoFates: improved prediction of mitochondrial targeting sequences and their cleavage sites. Mol Cell Proteom 14:1113–1126
Green TR, Dennis DT, West CA (1975) Compartmentation of isopentenyl pyrophosphate isomerase and prenyl transferase in developing castor bean endosperm. Biochem Biophys Res Commun 64:976–982
Guirimand G, Guihur A, Phillips MA, Oudin A, Glevarec G, Melin C, Papon N, Clastre M, St-Pierre B, Rodriguez-Concepcion M, Burlat V, Courdavault V (2012) A single gene encodes isopentenyl diphosphate isomerase isoforms targeted to plastids, mitochondria and peroxisomes in Catharanthus roseus. Plant Mol Biol 79:443–459
Heinig U, Gutensohn M, Dudareva N, Aharoni A (2013) The challenges of cellular compartmentalization in plant metabolic engineering. Curr Opin Biotechnol 24:239–246
Jin X, Bai C, Bassie L, Nogareda C, Romagosa I, Twyman RM, Christou P, Zhu C (2018) ZmPBF and ZmGAMYB transcription factors independently transactivate the promoter of the maize (Zea mays) β-carotene hydroxylase 2 gene. New Phytol 11:1–14
Jung KH, Lee J, Dardick C, Seo YS, Cao P, Canlas P, Phetsom J, Xu X, Ouyang S, An K, Cho YJ, Lee GC, Lee Y, An G, Ronald PC (2008) Identification and functional analysis of light-responsive unique genes and gene family members in rice. PLoS Genet 4:e1000164
Kajiwara S, Fraser PD, Kondo K, Misawa N (1997) Expression of an exogenous isopentenyl diphosphate isomerase gene enhances isoprenoid biosynthesis in Escherichia coli. Biochem J 324:421–426
Kaundal R, Raghava GPS (2009) RSLpred: an integrative system for predicting subcellular localization of rice proteins combining compositional and evolutionary information. Proteomics 9:2324–2342
Leivar P, González VM, Castel S, Trelease RN, López-Iglesias C, Arró M, Boronat A, Campos N, Ferrer A, Fernàndez-Busquets X (2005) Subcellular localization of arabidopsis 3-hydroxy-3-methylglutaryl-coenzyme A reductase. Plant Physiol 137:57–69
Martin D, Piulachs MD, Cunillera N, Ferrer A, Belles X (2007) Mitochondrial targeting of farnesyl diphosphate synthase is a widespread phenomenon in eukaryotes. Biochim Biophys Acta 1773:419–426
Misawa N, Satomi Y, Kondo K, Yokoyama A, Kajiwara S, Saito T, Ohtani T, Miki W (1995) Structure and functional analysis of a marine bacterial carotenoid biosynthesis gene cluster and astaxanthin biosynthetic pathway proposed at the gene level. J Bacteriol 177:6575–6584
Nakamura A, Shimada H, Masuda T, Ohta H, Takamiya K (2001) Two distinct isopentenyl diphosphate isomerases in cytosol and plastid are differentially induced by environmental stresses in tobacco. FEBS Lett 506:61–64
Neuberger G, Maurer-Stroh S, Eisenhaber B, Hartig A, Eisenhaber F (2003) Motif refinement of the peroxisomal targeting signal 1 and evaluation of taxon-specific differences. J Mol Biol 328:567–579
Nogueira M, Mora L, Enfissi EM, Bramley PM, Fraser PD (2013) Subchromoplast sequestration of carotenoids affects regulatory mechanisms in tomato lines expressing different carotenoid gene combinations. Plant Cell 25:4560–4579
Nogueira M, Enfissi EM, Almeida J, Fraser PD (2018) Creating plant molecular factories for industrial and nutritional isoprenoid production. Curr Opin Biotechnol 49:80–87
Okada K, Kasahara H, Yamaguchi S, Kawaide H, Kamiya Y, Nojiri H, Yamane H (2008) Genetic evidence for the role of isopentenyl diphosphate isomerases in the mevalonate pathway and plant development in Arabidopsis. Plant Cell Physiol 49:604–616
Page JE, Hause G, Raschke M, Gao W, Schmidt J, Zenk MH, Kutchan TM (2004) Functional analysis of the final steps of the 1-deoxy-D-xylulose 5-phosphate (DXP) pathway to isoprenoids in plants using virus-induced gene silencing. Plant Physiol 134:1401–1413
Pankratov I, McQuinn R, Schwartz J, Bar E, Fei Z, Lewinsohn E, Zamir D, Giovannoni JJ, Hirschberg J (2016) Fruit carotenoid-deficient mutants in tomato reveal a function of the plastidial isopentenyl diphosphate isomerase (IDI1) in carotenoid biosynthesis. Plant J 88:82–94
Phillips MA, D’Auria JC, Gershenzon J, Pichersky E (2008) The Arabidopsis thaliana type I isopentenyl diphosphate isomerases are targeted to multiple subcellular compartments and have overlapping functions in isoprenoid biosynthesis. Plant Cell 20:677–696
Pulido P, Perello C, Rodriguez-Concepcion M (2012) New insights into plant isoprenoid metabolism. Mol Plant 5:964–967
Rodriguez-Concepcion M (2010) Supply of precursors for carotenoid biosynthesis in plants. Arch Biochem Biophys 504:118–122
Sapir-Mir M, Mett A, Belausov E, Tal-Meshulam S, Frydman A, Gidoni D, Eyal Y (2008) Peroxisomal localization of Arabidopsis isopentenyl diphosphate isomerases suggests that part of the plant isoprenoid mevalonic acid pathway is compartmentalized to peroxisomes. Plant Physiol 148:1219–1228
Stamellos KD, Shackelford JE, Shechter I, Jiang G, Conrad D, Keller GA, Krisans SK (1993) Subcellular localization of squalene synthase in rat hepatic cells. J Biol Chem 268:12818–12824
Street IP, Coffman HR, Baker JA, Poulter CD (1994) Identification of Cys139 and Glu207 as catalytically important groups in the active site of isopentenyl diphosphate:dimethylallyl diphosphate isomerase. Biochem 33:4212–4217
Thibodeaux CJ, Liu HW (2017) The type II isopentenyl diphosphate:dimethylallyl diphosphate isomerase (IDI-2): a model for acid/base chemistry in flavoenzyme catalysis. Arch Biochem Biophys 632:47–58
Vranova E, Coman D, Gruissem W (2012) Structure and dynamics of the isoprenoid pathway network. Mol Plant 5:318–333
Vranova E, Coman D, Gruissem W (2013) Network analysis of the MVA and MEP pathways for isoprenoid synthesis. Annu Rev Plant Biol 64:665–700
Wang M, Wang D, Zhang Q, Chai J, Peng Y, Cai X (2017) Identification and cytochemical immunolocalization of acetyl-CoA acetyltransferase involved in the terpenoid mevalonate pathway in Euphorbia helioscopia laticifers. Bot Stud 58:62
Zhu C, Sanahuja G, Yuan D, Farre G, Arjo G, Berman J, Zorrilla-Lopez U, Banakar R, Bai C, Perez-Massot E, Bassie L, Capell T, Christou P (2013) Biofortification of plants with altered antioxidant content and composition: genetic engineering strategies. Plant Biotechnol J 11:129–141
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31870278); the Spanish Ministry of Economy and Competitiveness (MINECO), Spain (RTI2018-097613-B-I00; PGC2018-097655-B-I00); and in part by the European Union Framework Program DISCO (613513) “from DISCOvery to products: a next-generation pipeline for the sustainable generation of high-value plant products”, the European Cooperation in Science and Technology project EUROCAROTEN (OC-2015-1-19780), Generalitat de Catalunya Grant 2017 SGR 828 to the Agricultural Biotechnology and Bioeconomy Unit (ABBU), and the International Science and Technology Cooperation Project 20190201013JC (from Jilin Provincial Science and Technology Department, China).
Funding
CZ conceived and designed the research. XJ, CB, LG, VM, MD, XN, YS, LS, TC, PDF, and CZ conducted the experiments. XJ, CB, LG, VM, MD, XN, YS, LS, TC, PDF, PC, and CZ analyzed the data. XJ, PC, and CZ wrote the manuscript. All authors read and approved the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Stefan Schillberg.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jin, X., Baysal, C., Gao, L. et al. The subcellular localization of two isopentenyl diphosphate isomerases in rice suggests a role for the endoplasmic reticulum in isoprenoid biosynthesis. Plant Cell Rep 39, 119–133 (2020). https://doi.org/10.1007/s00299-019-02479-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-019-02479-x