Abstract
Interstitial deletions of 1q23.3q24.1 are rare. Here, chromosomal microarray testing identified a de novo microdeletion of arr[GRCh37]1q23.3q24.1(164816055_165696996) × 1 in a patient with moderate developmental delay, hearing loss, cryptorchidism, and other distinctive features. The clinical features were common to those previously reported in patients with overlapping deletions. The patient’s deletion size was 881 kb—the smallest yet reported. This therefore narrowed down the deletion responsible for the common clinical features. The deleted region included seven genes; deletion of LMX1A, RXRG, and ALDH9A1 may have caused our patient’s neurodevelopmental delay.
Similar content being viewed by others
Introduction
Interstitial microdeletions neighboring the 1q24 region are rare, and approximately 10 patients with such deletions have been reported to date1,2. One genotype–phenotype correlation study suggested that several regions may be responsible for the phenotypic features. Chatron et al.1 reported that patients with 1q24q25 microdeletions have distinctive phenotypes, including growth deficiency; they suggested that a 1.9-Mb region at 1q24.3q25.1 was the shortest region of overlap (SRO). Moreover, they proposed that deletions of two of the genes within the SRO, namely, the dynamin 3 gene (DNM3) and the centromere protein L gene (CENPL), were potential causes of intellectual disability and growth deficiency, respectively. However, some patients have microdeletions in the region proximal to the 1q24q25 region1.
Here, we report the smallest microdeletion yet found in the 1q23q24 region, and gene deletions that may be related to clinical features such as moderate developmental delay, hearing loss, and cryptorchidism, in a young boy.
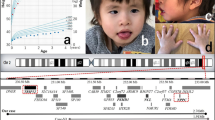
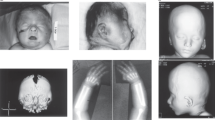
The patient was a boy aged 2 years and 8 months who had been born at full term with a birth weight of 2818 g and a normal spontaneous delivery. The pregnancy was uneventful. His parents were not consanguineous. Soon after birth, congenital hearing loss and cryptorchidism were found. At 4 months, poor weight gain and developmental delay were noted. At 10 months, the boy was referred to our hospital for further examination; at this time, he could turn over but could not sit unsupported. He had distinctive features, including mild hypotelorism, low-set ear, a tented upper lip, and a small jaw. Neurological examination revealed axial hypotonia. There were no abnormalities on routine laboratory examination, including plasma amino acids and urinary organic acids. Chromosomal G-banding showed a normal male karyotype of 46,XY. Brain magnetic resonance imaging (MRI) at 19 months revealed T2 high intensity in the right genu of the internal capsule (Fig. 1b, d); this had not been noted at 12 months (Fig. 1a, c). This area was not enhanced by gadolinium administration (Fig. 1c, d). Serum tumor markers were negative, with alpha-fetoprotein 8.0 ng/mL and human chorionic gonadotropin beta < 0.1 ng/mL.
Axial T2-weighted images were taken at a, c 12 months and b, d 19 months. c, d A T2 high-intensity area found in the right genu of the internal capsule was not enhanced by gadolinium administration. e The deleted regions in our patient and in patients reported previously are shown by red bars. f Results of chromosomal microarray analysis of our patient
At the time of writing, the boy was 82.0 cm tall ( <third percentile); he weighed 9.9 kg ( <third percentile) and had an occipitofrontal circumference of 48 cm (10th to 25th percentile), indicating growth impairment but not microcephaly. The boy can walk with support but has no meaningful vocabulary, indicating moderate developmental delay.
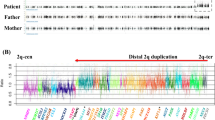
This study was approved by the ethical committees of Osaka City University and Tokyo Women’s Medical University. After obtaining written informed consent from the patient’s family, we performed chromosomal microarray testing using the Agilent microarray 60 K (Agilent Technologies, Santa Clara, CA, USA), as described previously3. A genomic copy number loss was identified at the chromosome region of 1q23.3q24.1 with minimum and maximum extents of 164,816,055–165,696,996 (881-kb) and 164,761,881–165,739,928 (978-kb), respectively [GRCh37/hg19]. (Fig. 1e, f). The same testing of both parents revealed that they did not have this deletion, indicating its de novo origin.
We identified a 1q23.3q24.1 microdeletion in a patient with developmental delay, growth impairment, hearing loss, and cryptorchidism. In the literature, there are reports of at least 10 patients who have shown deletions overlapping with that in our patient1,2,4,5,6,7. The clinical features of these patients are summarized in Table 1. Developmental delay, intellectual disability, or both were observed in all patients. Growth impairment, hypotonia, and distinctive features were also noted in most of the patients (Table 1). Depiction of the deletion regions of these patients on a genome map shows that the deletion identified in our patient is the smallest yet found (Fig. 1e). The genomic coordinates of the copy number variation in our patient delimit the maximum extent (Fig. 1f). In contrast, in the comparison cases reported by Chatron et al.1, the coordinates delimit the minimum extent, and in the patient reported by Al-Bughaili4, they were exact.
Among the genes included in the region deleted in our patient (Fig. 1e), seven are included in the Online Mendelian Inheritance in Man database (OMIM; https://omim.org/), and mutations or deletions of two—namely, the pre-B-cell leukemia homeobox 1 gene (PBX1; MIM #176310) and the transmembrane and coiled-coil domains 1 gene (TMCO1; MIM #614123)—have been associated with diseases (see Supplementary information). PBX1 is one of the three-amino-acid loop extension (TALE) homeodomain-containing transcription factors and is involved in early development in mammals8. Mutations or deletions of PBX1 are responsible for congenital anomalies of the kidney and urinary tract9. Although our patient had no renal abnormalities, eight other patients had renal malformation1,5,6,7 (Table 1). This would be derived from incomplete penetrance of phenotypic features. Mackenroth et al.5 proposed disruption of PBX1 as a potential cause of renal malformation, albeit perhaps with incomplete penetrance, as there was no renal malformation in patient 7 in the series of Chatron et al.1.
TMCO1 is a member of the DUF841 superfamily of several eukaryotic proteins with unknown function10. Xin et al.10 reported that a homozygous frameshift mutation in TMCO1 causes craniofacial dysmorphism, skeletal anomalies, and developmental delay. Because these clinical conditions are associated with autosomal recessive traits9, this gene would not be related to the clinical features of our patient.
The LIM homeobox transcription factor 1A gene (LMX1A) is essential for neuronal differentiation into dopaminergic neurons in the midbrain11, and the retinoic acid receptor RXR-gamma isoform C gene (RXRG), expressed mainly in the striatum, encodes a nuclear retinoic acid receptor12. Mackenroth et al.5 proposed that disruption of these genes was a cause of developmental delay. In addition, the SRO includes the gene encoding aldehyde dehydrogenase 9 family member A1 (ALDH9A1), which catalyzes the dehydrogenation of gamma-aminobutyraldehyde to gamma-aminobutyric acid. Bartlett et al.13 suggested that ALDH9A1 disruption may be related to autism, so the gene may also be associated with brain development. The analysis of more patients will be needed for a precise genotype–phenotype correlation study.
Our patient incidentally had an abnormal MRI finding in the basal ganglia that had not been apparent earlier. To rule out a malignant tumor, a detailed examination was performed, and a spot with high T2 intensity was found that was not enhanced by gadolinium. We were unable to reach a final conclusion about the significance of this lesion. Continuous follow-up of this finding will be needed.
HGV Database
The relevant data from this Data Report are hosted at the Human Genome Variation Database at
https://doi.org/10.6084/m9.figshare.hgv.2627
https://doi.org/10.6084/m9.figshare.hgv.2630
https://doi.org/10.6084/m9.figshare.hgv.2633
https://doi.org/10.6084/m9.figshare.hgv.2636
https://doi.org/10.6084/m9.figshare.hgv.2639
References
Chatron, N. et al. Refinement of genotype-phenotype correlation in 18 patients carrying a 1q24q25 deletion. Am. J. Med. Genet. A 167A, 1008–1017 (2015).
Burkardt, D. D. et al. Distinctive phenotype in 9 patients with deletion of chromosome 1q24-q25. Am. J. Med. Genet A 155A, 1336–1351 (2011).
Yamamoto, T. et al. An emerging phenotype of Xq22 microdeletions in females with severe intellectual disability, hypotonia and behavioral abnormalities. J. Hum. Genet. 59, 300–306 (2014).
Al-Bughaili, M. et al. de novo 1q23.3-q24.2 deletion combined witha GORAB missense mutation causes a distinctivephenotype with cutis laxa. J. Hum. Genet. 62, 325–328 (2017).
Mackenroth, L. et al. Interstitial 1q23.3q24.1 deletion in a patient with renal malformation, congenital heart disease, and mild intellectual disability. Am. J. Med. Genet. A 170, 2394–2399 (2016).
Lam, F. & Morris, C. Nine year old boy with chromosome 1q23.3-q25.1 deletion. Am. J. Med. Genet. A 170, 3013–3017 (2016).
Della Monica, M. et al. A case of autism with an interstitial 1q deletion (1q23.3–24.2) and a de novo translocation of chromosomes 1q and 5q. Am. J. Med. Genet. A 143A, 2733–2737 (2007).
Sonnet, W., Rezsohazy, R. & Donnay, I. Characterization of TALE genes expression during the first lineage segregation in mammalian embryos. Dev. Dyn. 241, 1827–1839 (2012).
Heidet, L. et al. Targeted exome sequencing identifies PBX1 as involved in monogenic congenital anomalies of the kidney and urinary tract. J. Am. Soc. Nephrol. 28, 2901–2914 (2017).
Xin, B. et al. Homozygous frameshift mutation in TMCO1 causes a syndrome with craniofacial dysmorphism, skeletal anomalies, and mental retardation. Proc. Natl Acad. Sci. USA 107, 258–263 (2010).
Andersson, E. et al. Identification of intrinsic determinants of midbrain dopamine neurons. Cell 124, 393–405 (2006).
Mangelsdorf, D. J. et al. Characterization of three RXR genes that mediate the action of 9-cis retinoic acid. Genes Dev. 6, 329–344 (1992).
Bartlett, C. W., Goedken, R. & Vieland, V. J. Effects of updating linkage evidence across subsets of data: reanalysis of the autism genetic resource exchange data set. Am. J. Hum. Genet. 76, 688–695 (2005).
Acknowledgements
This study was supported partially by the Practical Research Project for Rare/Intractable Diseases, Japan Agency for Medical Research and Development; a grant from the Ministry of Health, Labor and Welfare, Japan; and a grant-in-aid for Scientific Research C from the Japan Society for the Promotion of Science (to T.Y.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hoshina, T., Seto, T., Shimono, T. et al. Narrowing down the region responsible for 1q23.3q24.1 microdeletion by identifying the smallest deletion. Hum Genome Var 6, 47 (2019). https://doi.org/10.1038/s41439-019-0079-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41439-019-0079-1