Abstract
A recently disclosed Erk-induced PARP1 activation mediates the expression of immediate early genes (IEG) in response to a variety of extra- and intra-cellular signals implicated in memory acquisition, development and proliferation. Here, we review this mechanism, which is initiated by stimulation-induced binding of PARP1 to phosphorylated Erk translocated into the nucleus. Their binding maintains their long-lasting activity in a synergism, which offers a new pattern for targeted therapy.
Similar content being viewed by others
Introduction
Activated polyADP-ribose polymerase-1 (PARP1) catalyzes posttranslational modification of nuclear proteins by adding a series of negatively charged ADP-ribose moieties (poly-ADP-ribosylation).1,2 PARP1 substrates include PARP1 itself, histones, high mobility group proteins, topoisomerases, gyrases, DNA methyltransferase and demethylases, and the insulator protein CTCF (CCCTC-binding factor).1,3,4,5,6,7,8 Poly-ADP-ribosylation modulates the interaction of these substrates with the negatively charged DNA and with other chromatin-bound proteins.1,2 Poly-ADP-ribosylation of DNA methyltransferase has been explored for its epigenetic effect, and for its possible role in de novo methylation in the central nervous system.9,10,11,12,13
PARP1 is activated by binding to DNA breaks, and its poly-ADP-ribosylation is implicated in single-strand and double-strand DNA break repair.1,14,15 DNA-bound PARP1 poly-ADP-ribosylates chromatin-bound proteins, causing chromatin loosening near sites of DNA damage. In addition, ADP-ribose polymers on the activated PARP1 bind and recruit XRCC1 (X-ray repair cross-complementing protein 1), which acts as a scaffold for DNA repair proteins (DNA ligase 3, polynucleotide kinase-3-phosphatase and aprataxin).1,14,15 In double-strand break repair, activated and poly-ADP-ribosylated PARP1 is implicated to participate in DNA end resection for homologous recombination (HR) and in nonhomologous end joining (NHEJ) repair by activating the DNA-dependent kinase.14,15
Recent findings have revealed other mechanisms of PARP1 activation not involving its binding to DNA breaks. PARP1 is activated by interaction with the transcription factor Yin Yang 1 (YY1), which either up- or down-regulates gene expression.16 In addition, PARP1 is activated via a variety of signal-transduction mechanisms in the absence of stress conditions causing DNA breaks. PARP1 is activated by Ca2+ via CAMKII activation17 or via IP3-induced Ca2+ release into the nucleoplasm.18 Additionally, PARP1 becomes activated downstream in the MAP kinase phosphorylation cascade by binding to phosphorylated Erk, without involving the kinase activity.19,20,21 In this mechanism, activated PARP1 mediates Erk-induced expression of immediate early genes (IEGs), which are implicated in a variety of mechanisms unrelated to DNA repair.
IEG expression is independent of de novo-synthesized transcription factors or other protein mediators.22,23,24 IEGs are rapidly expressed in response to signals activating transcription factors bound to their promoters, including RNAPolII that is ready to act22,23,24,25 Many signal transduction pathways inducing IEG expression are mediated by phosphorylation of the mitogen-activated protein kinase (MAPK) cascade22,26,27,28,29,30 PARP1 activation is implicated in MAP kinase-induced expression of oncogenes that promote proliferation.31 Additionally, stimulation-induced PARP1 activation-mediated Erk-induced IEG expression that is implicated in synaptic potentiation and memory acquisition20,32 Here, we summarize findings indicating synergistic activity between PARP1 and phosphorylated Erk that mediates IEG expression. This mechanism reveals new targets of therapeutic significance.
PARP1 acts as an anchoring protein for phosphorylated Erk
Erk is bound to MEK in the cytoplasm of unstimulated cells at specific docking sites.33,34 Erk-MEK binding is disrupted by signals inducing MEK and Erk phosphorylation, and phosphorylated Erk is translocated apparently as a homodimer into the nucleus.33,34,35,36 Phosphorylated Erk homodimers do not diffuse freely into the nucleus. They are apparently translocated by transportins,33 although the modalities and regulation of Erk transfer and accumulation in the nucleus are not completely understood. In the absence of a nuclear localization signal (NLS), phosphorylated Erk could shuttle between the cytoplasm and the nucleus.33,34,35,36 However, relatively long-lasting activity of phosphorylated Erk in the nucleus has been documented in both quiescent and proliferating cells.29,30,34,37 This activity could be attributable to a possible Erk binding to nuclear protein(s) that retains its activity in the nucleus.29,34 Nuclear phosphatases, specifically MKPs, could be possible candidates.29,35 These phosphatases are activated by signals phosphorylating the MAP kinase cascade, and their activity is simultaneously regulated with the activity of phosphorylated Erk.35 However, MKPs are mainly expressed in proliferating cells, and only stress-inducing stimuli induce MKP expression in quiescent cells.35 However, long-lasting Erk activity has been measured in neurons under physiological conditions in the absence of stress-inducing stimulation.21,34 Recently, another candidate for anchoring phosphorylated Erk in the nuclei of both quiescent and proliferating cells under a variety of types of physiological stimulation has emerged.19,38 Docking sites of phosphorylated Erk have been identified in the abundant nuclear proteinPARP1.20,39,40,41,42 In addition, stimuli inducing Erk phosphorylation and translocation into the nucleus also induce the binding of phosphorylated Erk to PARP1,20 and PARP1 is required to maintain the activity of phosphorylated Erk in the nucleus for hours.20,37
PARP1 binding to phosphorylated Erk induces PARP1 activation
Binding to phosphorylated Erk induces PARP1 activation and poly-ADP-ribosylation.19,20,37 In a cell-free system, recombinant phosphorylated Erk-induced poly-ADP-ribosylation of recombinant PARP1 in the presence of NAD without implicating the kinase activity of Erk.19 Accordingly, PARP activation is dependent on MEK activity in stimulated cerebral neurons, cardiomyocytes and mouse embryonic fibroblasts (MEFs).19,20,37,43 Additionally, PARP1 has been found to be activated as long as it is bound to phosphorylated Erk, and poly-ADP-ribosylation does not interfere with this binding.19,20,38
Consensus docking sites of phosphorylated Erk have been identified in PARP1. These include four sites that partially match the known docking motifs of phosphorylated Erk in its various substrates: 633KYPKK637, 683KK684, 747KKPPLL752 and 1007FNF1009.39,40,41,42 All the sites are located in the WGR domain, helical domain (HD), and catalytic (CAT) domain of PARP1 (aa 556–1014)44 (Fig. 1a). Additionally, a negatively charged protein-binding domain in Erk (CRS/CD region) is involved in its binding to the docking sites in PARP1.19,20
a A ribbon structural model for the open conformation of PARP1 with optional consensus docking sites for phosphorylated Erk. Erk2 monomers in a homodimer (formed after Erk2 phosphorylation) are indicated by dark and light gray ribbons. Optional Erk-binding motifs on the HD, WGR and the CAT domain of PARP1 are indicated by orange spheres. The CRS/CD protein-binding region on Erk2 and the optional Erk-binding motifs on PARP1 are highlighted by red and blue shadows, respectively (from ref. 20) b The modeled conformation of PARP1 bound to damaged DNA indicating the occluded docking sites of phosphorylated Erk (from Ref # 20). c Autoradiograms presenting a comparison between the dose-dependent [32P]poly-ADP-ribosylation of recombinant PARP1 bound to recombinant phosphorylated Erk2 and the dose-dependent [32P]poly-ADP-ribosylation of recombinant PARP1 bound to DNA with single strand breaks, (nicked DNA, nDNA). [32P]poly-ADP-ribosylation was achieved in a mixture of β-NAD and [32P]NAD at the indicated concentrations (from ref. 19)
Binding to recombinant phosphorylated Erk has been found to induce poly-ADP-ribosylation of recombinant PARP1 at low NAD concentrations (lower than 1 µM), and recombinant PARP1 bound to recombinant phosphorylated Erk demonstrates ~70-fold higher affinity for NAD than recombinant PARP1 bound to nicked DNA (DNA with single-strand breaks)19,38 (Fig. 1c). Since poly-ADP-ribosylation does not interfere with the binding of PARP1 to phosphorylated Erk2, PARP1 that is poly-ADP-ribosylated via other signal transduction mechanisms (e.g., by IP3-induced Ca2+ release into the nucleoplasm18) can bind phosphorylated Erk and retain its activity in the nucleus as effectively as non-poly-ADP-ribosylated PARP1.20,37,44 The DNA-binding domain of PARP1 (Zn1-Zn2) does not possess Erk docking sites.20,44 However, PARP1 binding to DNA interferes with its binding to phosphorylated Erk due to structural rearrangements in DNA-bound PARP1 that occlude its Erk docking sites44 (Fig. 1b). Accordingly, PARP1 fails to bind phosphorylated Erk in the presence of accumulated DNA breaks.19,20
Erk-induced PARP1 activation has been examined by bioinformatics methods, and structural rearrangements in PARP1 bound to phosphorylated Erk2 have been analyzed. A reconstructed phosphorylated Erk2 homodimer (Protein Data Bank (PDB) PubMed ID 9298898) was docked on the helical, catalytic and WGR domains of PARP1 (PDB 4DQY).45,46,47,48,49 Positively charged patches in PARP1 that are predicted to bind phosphorylated Erk2 (aa residues 633–637 and 747–752) (Fig. 1a) were selected for in silico molecular docking by a method that predicts the preferred orientation of two molecules forming a stable complex.47,48,49 The conformational changes in PARP1 and phosphorylated Erk2 following binding were predicted using the anisotropic network model (ANM, http://ignmtest.ccbb.pitt.edu/cgi-bin/anm/anm1.cgi).49 A normal mode analysis plug-in for a molecular graphic viewer47 was used to present the outcome of this analysis. The calculated intramolecular directions of motion in PARP1 bound to phosphorylated Erk2 revealed that the helical domain (HD) and the catalytic (CAT) domain of PARP1 move in opposite directions, thereby exposing the NAD binding site in PARP120 (Fig. 1a and 1S (Movie; Supplemental Information)). Thus, exposure of the NAD binding site in PARP1 bound to phosphorylated Erk through the HD and WGR domains can enhance the frequency of NAD binding to its site in PARP1. Recent findings have shown how the helical domain (HD) of PARP1 can inhibit PARP1 activity by restricting the access of NAD to its binding site and regulating the frequency of NAD binding.50 Additionally, computed structural rearrangements of PARP1 bound to phosphorylated Erk that facilitate NAD binding are compatible with the high NAD affinity of PARP1 activated by binding to phosphorylated Erk19,20,38 (Fig. 1c).
Erk-induced PARP1 activation results in poly-ADP-ribosylation of histone H1
High-frequency electrical stimulation of cultured brain cortical neurons causing synaptic potentiation has been found to induce poly-ADP-ribosylation of PARP1 and its prominent substrate linker histone H1. This poly-ADP-ribosylation was prevented in the presence of specific MEK inhibitors20 (Fig. 2). Additionally, PARP1 was co-immunoprecipitated with phosphorylated Erk in nuclear extracts of the stimulated neurons unless they were treated with MEK inhibitors.20 These findings are in accordance with those of cell-free experiments in which recombinant H1 was poly-ADP-ribosylated in the presence of NAD, recombinant PARP1 and recombinant phosphorylated Erk.19
PARP1 activation, as measured by a shift in the PARP1 isoelectric point (pI) and that of its substrate histone H1, in cultured cortical neurons subjected to high-frequency electrical stimulation (100 Hz; induces synaptic potentiation) was prevented by either MEK or PARP inhibitors (U0126 and PJ-34, respectively) (from ref. 20)
Furthermore, PARP1 and Erk2 were coimmunoprecipitated with segments in the promoters of c-fos and zif268 in cerebral neurons stimulated by high-frequency electrical stimulation.20 Histone H1 was not coimmunoprecipitated with PARP1 and phosphorylated Erk2 in these chromatin coimmunoprecipitation reactions.20 These findings are in accordance with studies demonstrating the eviction of poly-ADP-ribosylated histone H1 from the promoter of c-fos in response to high-frequency electrical stimulation or membrane depolarization of cultured cerebral neurons.51,52
While H1 binding to nucleosomes induces a condensed chromatin structure that represses transcription,53 H1 eviction from nucleosomes evokes chromatin relaxation, rendering the DNA more accessible to proteins and transcription factors and thus facilitating gene expression.51,54,57 Accordingly, PARP1 accumulation accompanied by H1 depletion has been documented in promoters of transcribed genes, and PARP1 and H1 exhibit a reciprocal pattern of binding at promoters across the genome.54,55,56,57,58,59,60 H1 exclusion by PARP1 might not require PARP1 activation.54 However, H1 exclusion associated with transcription of upregulated genes involves poly-ADP-ribosylation.58,59,61 PARP1 activity is dispensable for the expression of genes negatively regulated by PARP1.62,63
In addition to the fact that histone H1 poly-ADP-ribosylation causes histone H1 eviction from promoters of cfos in depolarized cerebral neurons,51,52 in MCF-7 human breast cancer cells, histone H1 poly-ADP-ribosylation is mediated by poly-ADP-ribosylation of the demethylase KDM5B, which maintains methylation on histone H3 (H3K4me3) adjacent to promoters of transcribed genes.8 In another mechanism, in HeLa cervical cancer cells, PARP1 activation causes local destabilization of chromatin at cfos promoters by facilitating the exchange of the variant histone H2A.Z with histone H2A.64,65
Erk-induced PARP1 activation mediates IEG expression
In a cell-free system, recombinant Elk1 was phosphorylated by recombinant phosphorylated Erk in the presence of recombinant PARP1, ATP and NAD.19 Recombinant PARP1 and Elk1 did not bind directly. They were coimmunoprecipitated only in the presence of recombinant phosphorylated Erk2.19 Elk1 is phosphorylated by stimulation activating the MAP kinase cascade, causing PARP1 binding to phosphorylated Erk2 and PARP1 activation.19,20,38 Additionally, import of active recombinant phosphorylated Erk into the nuclei of permeabilized cortical neurons induces PARP1 activation and acetylation of histone H4, in accordance with the fact that Erk-induced activation of transcription factors is implicated in the activation of HATs (histone acetyltransferases).19,23,66,67,68 Furthermore, high-frequency stimulation of cultured cerebral neurons that induces synaptic potentiation also induces the expression of the IEGs cfos, zif268 and arc, which are implicated in synaptic potentiation and memory acquisition.20,69,70,71,72,73 PARP1 inhibition, silencing or genetic deletion prevents IEG expression.20 The induced expression of cfos and zif268 is consistent with the coimmunoprecipitation of PARP1, phosphorylated Erk and acetylated H4 with DNA segments in the promoters of c-fos and zif268.20 These results are consistent with the finding that PARP1 activation mediates Erk-induced expression of IEGs in stimulated neurons.20,23,74
Stimulation that induces H1 poly-ADP-ribosylation and eviction from chromatin51,52,54,55,58 could render the transcription factor Elk1 in the promoters of cfos and zif268 accessible to phosphorylation by PARP1-bound phosphorylated Erk.23 Elk1 phosphorylation-mediated activation of the HAT activity of CBP/p300 induces acetylation of core histone-promoting transcription.23
High-frequency electrical stimulation or treatment with nerve growth factors could induce the expression of cfos, zif268 and arc following Erk phosphorylation and the binding of phosphorylated Erk translocated into the nucleus to PARP1.20,21,69,70,71,72,73,74 In accordance with this finding, MEK inhibition, PARP1 inhibition, PARP1 silencing and PARP1 genetic deletion prevent both the expression of these IEGs and synaptic potentiation20 (Fig. 3). These findings may outline a rapid signal transduction mechanism mediating IEG expression in cerebral neurons in response to electrical stimulation75 (Fig. 4). Findings indicating that RNApolII positioned on IEG promoters is rapidly activated in response to stimulation25 are consistent with the rapid responses implicated in IEG expression in cerebral neurons. Measurements in cellular model systems of stimulated cerebral neurons could reflect physiological responses.20,75 In vivo experiments with rodents and PARP1-KO mice have supported the pivotal role of PARP1 activity in memory acquisition. PARP1 inhibition in rodents (and also in the marine mollusk Aplysia) or PARP1 genetic deletion in PARP1-KO mice prevents long-term memory acquisition during learning.76,77,78
The relative expression rates of the IEGs c-fos, zif268 and arc were measured by RT-PCR at the indicated time intervals after the indicated electrical stimulation (1 s, 3 repeats) of cultured brain cortical neurons with three different frequencies (100 Hz, 10 Hz or 1 Hz). Enhanced expression rates of these genes were measured only in response to high-frequency stimulation (100 Hz; black line), which induces synaptic potentiation. The expression of these genes was prevented in stimulated neurons treated with the PARP inhibitors PJ-34 and Tiq-A (gray lines) (from ref. 20)
A schematic diagram (in red) indicating the regulation of IEG expression by PARP1-Erk synergism as part of a signal transduction network (in gray and black) that mediates synaptic plasticity, MEF proliferation, and newborn cardiomyocyte development. (poly-ADP-ribosylated PARP1 and histone H1: pADPr-PARP1 and pADPr-H1, respectively; phosphorylated Erk and Elk1: pErk and pElk1, respectively; poly-ADP-ribosylated PARP1 bound to phosphorylated Erk; pADPr-PARP1  pErk)
pErk)
DNA damage prevents PARP1-Erk binding in cerebral neurons
In a cell-free system, recombinant PARP1 was found not to bind or be activated by phosphorylated Erk in the presence of nicked DNA (DNA single strand breaks).19 In stimulated cultured cerebral neurons, IEG expression was prevented in the presence of accumulated DNA single-strand breaks, similar to the effect of PARP1 inhibition, silencing or genetic deletion. Preventing the binding of PARP1 to DNA restored the expression of IEGs.20 These results are consistent with the recently indicated structural modifications in DNA-bound PARP1 that occlude Erk docking sites in its HD and WGR domains20,44 (Fig. 1a, b).
Accumulation of single-strand DNA breaks is most common in aged cerebral neurons, which cannot be replaced during an organism’s lifetime. These breaks are caused by oxidative damage due to high energy demands in the central nervous system, and due to declines in antioxidant defensive mechanisms during senescence.74,79,80,81,82 Thus, gene expression might be suppressed in aged neurons by mechanisms preventing the transcription of damaged DNA.83 However, the expression of cfos and zif268, which is suppressed in stimulated neurons carrying accumulated DNA breaks, has been found to be restored by preventing the binding of PARP1 to DNA breaks. This restoration was demonstrated by IEG expression when recombinant PARP1 lacking the DNA-binding domain was expressed in PARP1-KO cortical neurons treated with a DNA damaging agent or when poly-ADP-ribose glycohydrolase (PARG) was inhibited84 and the recurrent binding of PARP1 to DNA was prevented.20,84
Since cfos, zif268 and arc expression have been implicated in synaptic potentiation,67,68,69,70,71,72,73 DNA damage that suppresses their expression by preventing the binding of PARP1 to phosphorylated Erk might affect synaptic potentiation.20,74 In support of this possibility, DNA damage, PARP1 inhibition and PARP1 genetic deletion were found to prevent long-term synaptic potentiation in a hippocampal cell model and to prevent long-term memory in rodents.20,76,77,78 Additionally, PARG inhibitors have been found to improve learning ability in aged rats.85 Recent findings have associated failure of synaptic potentiation, or synaptic silencing with the initiation of Alzheimer’s disease.86,87
PARP1-Erk synergism in newborn cardiomyocytes
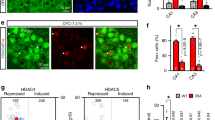
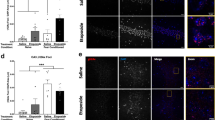
Cardiomyocytes cannot be replaced during an organism’s lifetime; thus, stress conditions causing persistent DNA damage and cell death may cause permanent damage to the myocardium (heart muscle).88 Under ischemia caused by myocardial infarction (MI), cell death could be induced in cardiomyocytes due to the transportation of poly-ADP-ribose polymers of highly activated PARP1 to the mitochondria, causing the release of AIF (apoptotic inducing factor) that activates DNA-dependent caspases.88,89,90,91 In support of this finding, PARP1-KO mice have better cardiac function under ischemia imposed by MI than wild-type mice,90 and PARP1 inhibitors reduce cardiac cell death caused by MI in normal mice.90,91
In contrast, PARP1 inhibition might not be beneficial in newborn cardiomyocytes. PARP-Erk synergism has been documented in newborn cardiomyocytes treated with the hormone/growth factor angiotensin-II (AngII).43 Intracellular Ca2+ release and activation of the MAP kinase phosphorylation cascade mediate the AngII-induced high contraction rates of newborn cardiomyocytes in cell cultures.43 In these cells, PARP1 is activated and coimmunoprecipitated with phosphorylated Erk in response to AngII-induced stimulation, and PARP1 is coimmunoprecipitated with segments in the cfos promoter. Additionally, cfos expression is suppressed by both PARP1 and MEK inhibitors.43,90 These findings implicate PARP1 in the expression of cfos in newborn cardiomyocytes exposed to AngII. Phosphorylated cFos protein bound to GATA4 acts as a transcription factor of atrial natriuretic factor (ANF),92 which is implicated in the growth and development of newborn cardiomyocytes.92,93 In cultured newborn cardiomyocytes, Erk-induced PARP1 poly-ADP-ribosylation mediates the assembly of cFos bound to GATA4 in the ANF promoter, inducing ANF expression.43 Accordingly, PARP1 inhibition, or silencing prevents both c-fos and ANF expression in these cells,43 leading to a negative influence of PARP1 inhibition on the growth and development of newborn cardiomyocytes91,92,93 (Fig. 4). This mechanism might be of interest when PARP1 inhibitors, currently offered for cancer treatments, are administered during pregnancy or early childhood.
PARP1-Erk synergism in proliferating cells and targeted therapy
PARP1 is coimmunoprecipitated with phosphorylated Erk in nuclear protein extracts prepared from mouse embryonic fibroblasts (MEFs) treated with PMA (phorbol 12-myristate 13 acetate).37 PMA activates the MAP kinase cascade via PKC activation.94 Similar to the case in cerebral neurons and newborn cardiomyocytes, PARP1 is required to maintain long-lasting activity of phosphorylated Erk in the nuclei of MEFs, and both PARP1 and Erk remain activated for more than an hour after stimulation.37
In proliferating cells, activation of the transcription factor AP1, which is an heterodimer frequently composed of phosphorylated c-Fos protein bound to c-Jun,95 eventually leads to cyclin D expression, and initiates mitosis95,96 (Fig. 4). Similar to the case in neuronal cells, PARP1 silencing and PARP1 genetic deletion downregulate the presence of phosphorylated Erk in the nuclei of MEFs, leading to PARP1-dependent downregulation of their Erk-induced proliferation.27,28,29 However, unlike in cerebral neurons, PARP1 inhibition does not suppress cfos expression. Delayed elevations in cFos have been measured in the nuclei of MEFs pre-treated with PARP1 inhibitors.37 This might indicate a parallel alternative PARP1-independent pathway promoting cfos expression in MEFs treated with PMA. Phosphorylation of transcription factors by phosphorylated RSK, one of the substrates of phosphorylated Erk acting in both the cytoplasm and nuclei mainly in proliferating cells,94 could mediate the expression of cfos in MEFs after PARP1 inhibition.
Blocking the activation of the MAP kinase phosphorylation cascade to downregulate Erk-induced oncogene expression and proliferation in malignant cells has been thoroughly examined.95,96,97,98,99 Erk is constantly phosphorylated in RAS mutant cancer cells that are mostly resistant to therapy.98,99 However, treatments that inhibit the MAP kinase phosphorylation cascade by blocking the activity of MEK or by blocking receptors of growth factors that activate the MAP kinase cascade94,95,96,97 have failed to prevent the consequences of sustained uncontrolled Erk activity in RAS mutant cancer cells.98,99 Recently, a treatment combining PARP1 and MEK inhibitors yielded positive results in patients with RAS mutant cancer tumors.99 These findings are consistent with the idea that PARP1 activity preserves the long-lasting activity of phosphorylated Erk in the nuclei of these malignant cells, although PARP1-Erk synergism has not been reported in RAS mutant cancer cells.
PARP1 inhibitors also efficiently eradicate MCF-7 breast cancer cells,100,101,102 and PARP1 silencing downregulates the activity of phosphorylated Erk in the nuclei of these cells.37 In HeLa human cervical cancer cells, MAP kinase phosphorylation-mediated the binding of PARP1 to the promoter of cfos. The activation of the transcription factor NF1 downstream of MAPK activation mediates the binding of PARP1 to the cfos promoter in these cells.64 An additional mechanism controlling oncogene expression in malignant cells is mediated by MAP kinase activation. In human malignant cells, activation of the MAP kinase phosphorylation cascade has been implicated in the regulation of a group of miRNAs that downregulate the expression of immediate early oncogenes.103,104
Despite evidence indicating that PARP1 inhibition interferes with oncogene expression in malignant cells,31 PARP1 inhibitors have been mainly examined for their role in reinforcing the activity of DNA-damaging agents or in BRCA mutant cancer cells105 in which double-strand DNA break repair is impaired.106,107 PARP1 inhibition preventing DNA repair also interferes with the repair of damaged DNA in p53 mutant cancer cells,108,109 and promotes cell death in PTEN phosphatase mutant cells with an uncontrolled Akt kinase activity.110
Recent findings have identified molecules that have been tagged as PARP1 inhibitors but that act through a PARP1-independent mechanism. A group of phenanthrenes (PJ34, Phen and Tiq-A) acting as potent PARP1 inhibitors that share high affinity for the NAD binding site in PARP1,111 target the activity of NuMA (nuclear mitotic apparatus protein-1)112 that stabilizes the spindle poles during mitosis, by inhibiting the serine-threonine kinase Pim1 and tankyrase 1 (a PARP family member), both of which are scarcely expressed in normal somatic cells.113 This activity prevents the binding of NuMA to α-tubulin and interferes with its sliding towards the spindle poles. Unstable spindle poles prevent chromosomes segregation and causes G2/M phase arrest followed by cell death through mitotic catastrophe death. These molecules have been shown to efficiently eradicate a variety of resistant human cancer cells without impairing normal cells.113
Conclusion
A rapid signal transduction mechanism that mediates stimulation-induced IEG expression is based on Erk-induced PARP1 activation that renders transcription factors accessible to phosphorylated Erk. Binding to PARP1 results in long-lasting activity of phosphorylated Erk in the nucleus and PARP1 activation with a high affinity for NAD, which lasts as long as PARP1 is bound to phosphorylated Erk. This mechanism could be involved in rapid responses to signals that induce memory acquisition during learning as well as in long-lasting stimulation-induced development or proliferation. Jeopardizing this mechanism could impair synaptic potentiation and memory but could be beneficial in targeted cancer therapy.
References
Krishnakumar, R. & Kraus, W. L. The PARP side of the nucleus: Molecular actions, physiological outcomes, and clinical targets. Mol. Cell 39, 8–24 (2010).
Gibson, B. A. & Kraus, W. L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat. Rev. Mol. Cell Biol. 13, 411–424 (2012).
Zhang, Q. & Wang, Y. High mobility group proteins and their post-translational modifications. Biochem. Biophys. Acta 1784, 1159–1166 (2008).
Ciccarone, F., Zampieri, M. & Caifa, P. PARP1 orchestrates epigenetic events setting-up chromatin domains. Semin. Cell Dev. Biol. 63, 123–134 (2017).
Caifa, P., Guastafierro, T. & Zampieri, M. Epigenetics: polyADP-ribosylation of PARP1 regulates genomic methylation patterns. FASEB J. 23, 672–678 (2009).
Ohlsson, R., Lobanenkov, V. & Klenova, E. Does CTCF mediate between nuclear organization and gene expression? Bioessays 32, 37–50 (2010).
Ji, Y. & Tulin, A. V. The roles of PARP1 in gene control and cell differentiation. Curr. Opin. Genet. Dev. 20, 512–518 (2010).
Krishnakumar, R. & Kraus, W. L. PARP1-regulated chromatin structure and transcription through a KDM5B-dependent pathway. Mol. Cell 39, 736–749 (2010).
Althaus, F. R. Poly(ADP-ribose): a coregulatory of DNA methylation? Oncogene 24, 11–12 (2005).
Cholewa-Waclaw, J. et al. The role of epigenetic mechanisms in the regulation of gene expression in the nervous system. J. Neurosci. 36, 11427–11434 (2016).
Day, J. J. & Sweatt, J. D. DNA methylation and memory formation. Nat. Neurosci. 13, 1319–1323 (2010).
Day, J. J. et al. DNA methylation regulates associative reward learning. Nat. Neurosci. 16, 1445–1452 (2013).
Lax, E. et al. PARP-1 is required for retrieval of cocaine-associated memory by binding to the promoter of a novel gene encoding a putative transposase inhibitor. Mol. Psychiatry 22, 570–579 (2017).
Wang, M. et al. PARP1 and Ku compete for repair of DNA double strand breaks by distinct NHEJ pathways. Nuc Acids Res. 34, 6170–6182 (2006).
Chaudhuri, A. R. & Nussenzweig, A. The multifaceted roles of PARP1 in DNA repair and chromatin remodeling. Nat. Rev. Mol. Cell Biol. 18, 610–621 (2017).
Li Oei, S. & Shi, Y. Transcription factor Yin Yang1 stimulates polyADP-ribosylation and DNA repair. Biochem. Biophys. Res. Comm. 284, 450–454 (2001).
Ju, B. G. et al. Activating the PARP1 sensor component of the Groucho/TLE1 corepressor complex mediates a CAMkinase Iid-dependent neurogenic gene activation pathway. Cell 119, 815–829 (2004).
Homburg, S. et al. A fast signal- induced activation of poly(ADP-ribose) polymerase: A novel downstream target of phospholipase C. J. Cell. Biol. 150, 293–308 (2000).
Cohen-Armon, M. et al. DNA-independent PARP-1 activation by phosphorylated ERK2 increases Elk1 activity:a link to histone acetylation. Mol. Cell 25, 297–308 (2007).
Visochek, L. et al. A PARP1-ERK2 synergism is required for the induction of LTP. Sci. Rep. 6, 24950 (2016).
Visochek, L. et al. PolyADP-ribosylation is involved in neurotrophic activity. J. Neurosci. 25, 7420–7428 (2005).
Baharami, S. & Drables, F. Gene regulation in the immediate early response process. Adv. Biol. Reg. 62, 37–49 (2016).
Buchwalter, G., Gross, C. & Wasylyk, B. Ets ternary complex transcription factors. Gene 24, 1–14 (2004).
Esnault, C. et al. 2017 Erk induced activation of TCF family of SRF cofactors initiates a chromatin modification cascade associated with transcription. Mol. Cell 65, 1081–1095.e5 (2017).
Saha, R. N. et al. Rapid activity-induced transcription of Arc and other IEGs relies on poised RNA polymerase-II. Nat. Neurosci. 14, 848–856 (2011).
Yang, S.-H., Shaaocks, A. D. & Whitmarsh, A. J. MAP kinase signaling cascades and transcriptional regulation. Gene 513, 1–13 (2013).
Zhang, W. & Liu, H. T. 2002. MAPK signal pathways in the regulation of cell proliferation in mammalian cells. Cell Res. 12, 9–18 (2002).
Whitmarsh, A. J. & Davis, R. J. Transcription factor AP-1 regulation by mitogen-activated protein kinase signal transduction pathways. J. Mol. Med. 74, 589–607 (1996).
Chambard, J.-C., Lefloch, R., Pouyssegur, J. & Lenormand, P. Erk implication in cell cycle regulation. Biochem. Biophys. Acta 1773, 1299–1310 (2007).
Wang, Z., Ge, L., Wang, M. & Carr, B. I. Phosphorylation regulates Myc expressionvia prolonged activation of the mitogen-activated protein kinase pathway. J. Cell. Physiol. 208, 133–140 (2006).
Carbone, M., Rossi, M. N., Cavaklesi, M., Amati, P. & Maione, R. PolyADP-ribosylation is implicated in the G0-G1 transision of resting cells. Oncogene 27, 6083–6092 (2008).
Jones, M. W. et al. A requirement for the immediate early gene Zif268 in the expression of late LTP and long-term memories. Nat. Neurosci. 4, 289–296 (2001).
Plotnikov, A., Zehorai, E., Procaccia, S. & Seger, R. The MAPK cascade: signaling components, nuclear roles and mechanism of nuclear translocation. Biochim. Biophys. Acta 1813, 1619–1633 (2011).
Sasagawa, S., Ozaki, Y. I., Fujita, K. & Kuroda, S. Prediction and validation of the distinct dynamics of transient and sustained ERK activation. Nat. Cell Biol. 7, 365–373 (2005).
Brondello, J. M., Brunet, A., Pouysse´gur, J. & McKenzie, F. R. The Dual Specificity Mitogen-activated Protein Kinase Phosphatase1 and -2 Are Induced by the p42/p44MAPK Cascade. J. Biol. Chem. 272, 1368–1376 (1997).
Moreno-Layseca, P. & Streuli, C. H. Signaling pathways linking integrins with cell cycle progression. Matrix Biol. 34, 144–153 (2014).
Visochek, L. & Cohen-Armon, M. PARP-Erk synergism in proliferating cells. Oncotarget 9, 29140–29145 (2018).
Cohen-Armon, M. PARP1 activation in the ERK signaling pathway. Trends Pharmacol. Sci. 28, 556–260 (2007).
Tanoue, T., Adachi, M., Moriguchi, T. & Nishida, E. A conserved docking motif in MAP kinases common to substrates activators and regulators. Nat. Cell Biol. 2, 110–116 (2000).
Jacobs, D., Glossip, D., Xing, H., Muslin, A. J. & Kornfeld, K. Multiple docking sites on substrate proteins form modular system that mediates recognition by Erk MAP kinase. Gene Dev. 13, 163–175 (1999).
Fantz, D. A., Jacobs, D., Glossip, D. & Kornfeld, K. Docking sites on substrate proteins direct extra-cellular signal regulated kinase tophophorylate specific residues. J. Biol. Chem. 276, 27256–27265 (2001).
Tanoue, T. & Nishida, E. Docking interactions in the mitogen-activated protein kinase cascades. Pharmacol. Ther. 2–3, 193–202 (2002).
Geistrikh, I. et al. Ca2+ -induced PARP-1 activation and ANF expression are coupled events in cardiomyocytes. Biochem. J. 438, 337–347 (2011).
Langelier, M. F., Planck, J. L., Roy, S. & Pascal, J. M. Structural Basis for DNA Damage–Dependent Poly(ADP-ribosyl)ation by Human PARP-1. Science 336, 728–732 (2012).
Bakan, A., Meireles, L. M. & Bahar, I. ProDy: Protein dynamics inferred from theory and experiments. Bioinformatics 27, 1575–1577 (2011).
Jimenez-Garcia, B., Pons, C. & Fernandez-Recio, J. pyDockWEB: a web server for rigidbody protein-protein docking using electrostatics and desolvation scoring. Bioinformatics 29, 1698–1699 (2013).
Humphrey, W., Dalke, A. & Chulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Sanner, M. F. Python: A Programming Language for Software Integration and Development. J. Mol. Graphics Mod. 17, 57–61 (1999).
Atilgan, A. R. et al. Anisotropy of fluctuation dynamics of proteins with an elastic network model. Biophys. J. 80, 505–515 (2001).
Langelier, M.-F., Zandarashvili, L., Aguiar, P. M., Black, B. E. & Pascal, J. M. NAD+ analog reveals PARP-1 substrate-blocking mechanism and allosteric communication from catalytic center to DNA-binding domains. Nat. Commun. 9, 844–857 (2018).
Á. Fontán-Lozano, A. et al. Histone H1 Poly[ADP]-Ribosylation Regulates the Chromatin Alterations Required for learning consolidation. J. Neurosci. 30, 13305–13313 (2010).
Azad, G. K. et al. PARP1-dependent eviction of the linker histone H1mediates immediate early gene expression during neuronal activation. J. Cell. Biol. 217, 1–9 (2017).
Happel, N. & Doenecke, D. Histone H1 and its isoforms:contribution to chromatin structure and function. Gene 43, 1–12 (2009).
Kim, N. Y., Mauro, S., Gevry, N., Lis, J. T. & Kraus, W. L. NAD+ dependent modulation of chromatin structure and transcription by nucleosome binding properties of PARP1. Cell 119, 803–814 (2004).
Krishnakumar, R. & Kraus, W. L. The PARP side of the nucleus: molecular actions, physiological outcomes and clinical targets. Mol. Cell 39, 8–24 (2010).
Tulin, A. & spradling, A. Chromatin loosening by polyADP-ribose polymerase (PARP) at Drosophila puff loci. Science 299, 560–562 (2003).
Wacker, D. A. et al. The DNA binding and catalytic domains of poly(ADP-ribose) polymerase 1 cooperate in the regulation of chromatin structure and transcription. Mol. Cell. Biol. 27, 7475–7485 (2007).
Fan, Y. et al. Histone H1 depletion in mammals alters global chromatin structure but causes specific changes in gene regulation. Cell 123, 1199–1212 (2005).
Lu, X. et al. Linker histone H1 essential for Drosophila development, the establishment of pericentric heterochromatin, and a normal polytene chromosome structure. Genes Dev. 23, 452–465 (2009).
Petesch, S. J. & Lis, J. T. Rapid transcription-independent loss of nucleosomes over a large chromatin domain at Hsp70 loci. Cell 134, 74–84 (2008).
Frizzell, K. M. et al. Global analysis of transcriptional regulation by polyADP-ribose polymerase-1 and polyADP-ribose glycohydrolase in MCF-7 human breast cancer cells. J. Biol. Chem. 284, 33926–33938 (2009).
Hassa, P. O. & Hottiger, M. O. The functional role of polyadp-ribose polymerase 1 as novel coactivator of NF-kappaB in inflammatory disorders. Cell. Mol. Life Sci. 59, 1534–1553 (2002).
Pavri, R. et al. PARP1 determines specificity in a retinoid signaling pathway via direct modulation of mediator. Mol. Cell 18, 83–96 (2005).
O’Donnell, A., Yang, S.-H. & Sharrocks, A. D. PARP1 orchestrates variant histone exchange in signal-mediated transcriptional activation. EMBO Rep. 14, 1084–1091 (2013).
Martinez-Zamudio, R. & Ha., H. C. Histone polyADP-ribosylation facilitates gene transcription by directly remodeling nucleosomes. Mol Cell. Bio. 32, 2490–2502 (2012).
Li, Q. J. et al. MAP kinase phosphorylation-dependent activation of Elk-1 leads to activation of the co-activator p300. EMBO J. 15, 281–291 (2003).
Korzus, E., Rosenfeld, M. G. & Mayford, M. CBP histone acetyltransferase activity is a critical component of memory consolidation. Neuron 42, 961–97248 (2004).
Besnard, A., Gala-Rodriguez, B., Vanhoutte, P. & Caboche, J. Elk1 a transcription factor with multiple facets in the brain. Front Neurosci. 5, 35 (2011).
Sng, J. C. G., Taniura, H. & Yoneda, Y. A tale of early response genes. Biol. Pharm. Bull. 27, 606–612 (2004).
Wang, S.-H. et al. NGF promotes long-term memory formation by activating poly(ADPribose) polymerase-1. Neuropharmacology 63, 1085–1092 (2012).
Impey, S. K., Obrietan, K. & Storm, D. R. Making new connections: Role of ERK-MAP kinase signaling in neuronal plasticity. Neuron 23, 11–14 (1999).
Jones, M. W. et al. A requirement for the immediate early gene Zif268 in the expression of late LTP and long-term memories. Nat. Neurosci. 4, 289–296 (2001).
Flavell, S. W. & Greenberg, M. E. Signaling mechanisms linking neuronal activity to gene expression and plasticity of the nervous system. Ann Rev. Neurosci. 31, 563–590 (2008).
Thomas, G. M. & Huganir, R. L. MAPK cascade signaling and synaptic plasticity. Nature 5, 173–183 (2004).
Bliss, T. V. P. & Collingridge, G. L. A synaptic model of memory: long-term potentiation in the hippocampus. Nature 361, 31–39 (1993).
Goldberg, S., Visochek, L., Giladi, E., Gozes, I. & Cohen-Armon, M. PolyADP-ribosylation is required for long-term memory formation in mammals. J. Neurochem. 111, 72–79 (2009).
Piskunova, T. S. et al. Deficiancy in ADP-ribose polymerase in mice. Curr. Gerontol. Geriatr. Res. 754, 190–197 (2008).
Cohen-Armon, M. et al. Long-term memory requires polyADP-ribosylation. Science 304, 1820–1822 (2004).
Lu, T. et al. Gene regulation and DNA damage in the ageing human brain. Nature 429, 883–891 (2004).
Kumar, A. Long-term potentiation at CA3-CA1 hippocampal synapses with special emphasis on aging, disease and stress. Front Aging Neurosci. 3, 2–20 (2011).
Kann, O. R. Mitochondria and neuronal activity. Am. J. Physiol. Cell. Physiol. 292, C641–C657 (2007).
Mattson, M. P. & Magnus, T. Ageing and neuronal vulnerability. Nat. Rev. Neurosci. 7, 278–294 (2006).
Gregersen, L. H. & Jesper, J. Q. The cellular response to transcription blocking DNA damage. Trend Biochem. Sci. 43, 341 (2018).
Finch, K. E., Knezevik, C. E., Nottbohm, A. C., Partlow, K. C. & Hergenrother, P. J. Selective small molecule inhibition of Poly(ADP-ribose)glycohydrolase (PARG). ACS Chem. Biol. 7, 563–570 (2012).
Tian, Y. et al. High molecular weight persimmon tannin ameliorates cognition deficits and attenuates oxidative damage in senescent mice induced by D-galactose. Food. Chem. Toxicol. 49, 1728–1736 (2011).
Wang, Z. et al. Human brain-derived Aβ oligomers bind to synapses and disrupt synaptic activity in a manner that requires APP. J. Neurosci. 37, 11947–11966 (2017).
Styr, B. & Slutsky, I. Imbalance between firing homeostasis and synaptic plasticity drives early phase Alzheimer’s disease. Nat. Neurosci. 21, 463–673 (2018).
Pacher, P. & Szabo, C. Role of polyADP-ribosepolymerase1 (PARP1) in cardiovascular diseases: the therapeutic potential of PARP inhibitors. Cardiovasc. Drug. Rev. 25, 235–260 (2007).
Bano, D. & Prehn, J. H. M. Apoptosis-inducing factor (AIF) in physiology and disease: the tale of arepented natural born killer. EBioMedicine 30, 29–37 (2018).
Pillai, J. B. et al. PolyADP-ribose polymerase-1-deficient mice are protected from angiotensin-II-induced cardiac hypertrophy. Am. J. Heart Circ. Physiol. 291, H1545–H1553 (2006).
Palfi, A. et al. PARP inhibition prevents postinfarction myocardial remodeling and heart failure via the protein kinase C/glycogen synthase kinase-3β pathway. J. Mol. Cell. Cardiol. 41, 149–159 (2006).
McBride, K., Charron, F., Lefebvre, C. & Nemer, M. Interaction with GATA transcription factors provides a mechanism for cell-specific effects of c-Fos. Oncogene 22, 8403–8412 (2003).
Small, E. M. & Krieg, A. P. Transgenic analysis of atrial naturetic factor (ANF) promoter: Nikx2-5 and GATA4 binding sites are required for atrial specific expression of ANF. Dev. Biol. 261, 116–131 (2003).
Yoon, S. & Seger, R. The extracellular signal-regulated kinase: Multiple substrates regulate diverse cellular functions. Growth Factors 24, 21–44 (2006).
Milde-Langosch, K. The Fos family of transcription factors and their role in tumourigenesis. Eur. J. Canc. 41, 2449–2461 (2005).
Herdegen, T. & Leah, J. D. Inducible and constitutive transcription factors in the mammalian nervous system: control of gene expression by Jun, Fos and Krox and CREB/ATF proteins. Brain Res. Rev. 28, 370–490 (1998).
Reinhart, J. et al. Multicenter phase II study of the oral MEK inhibitor, CI-1040 in patients with advanced non-small cell lung, breast, colon, and pancreatic cancer. J. Clin. Oncol. 22, 4456–4462 (2004).
Ryan, B. M., Der, C. J., Wang-Gillam, A. & Cox, A. D. Targeting RAS mutant cancers: Is Erk the key? Trends Canc. 1, 183–198 (2015).
Sun, C. et al. Rational combination therapy with PARP and MEK inhibitors capitalizes on therapeutic liabilities in RAS mutant cancers. Sci. Transl. Med. 9, eaal5148 (2017).
Inbar-Rozensal, D. et al. A selective eradication of human nonhereditary breast cancer cellsby phenanthridine-derived polyADP-ribose polymerase inhibitors. Breast Cancer Res. 11, R78 (2009).
Duan, R., Xie, W. & Burghardt, R. C. Safe Estrogen receptor mediated activation of the serum response element in MCF-7 cells through MAPK-dependent phosphorylation of Elk1. J. Biol. Chem. 276, 11590–11598 (2001).
Lu, C. et al. cFos is critical for MCF-7 breast cancer cell growth. Oncogene 24, 6516–6524 (2005).
Avraham, R. et al. EGF decreases the abundance of micro RNA that retain oncogenic transcription factors. Sci. Signaling 3, ra43 (2010).
O’Donnell, A. & Odrowaz, Za Sharrocks. Immediate early gene activation by the MAPK pathways: What do and don’t we know? Biochem. Soc. Trans. 40, 58–66 (2012).
Bitler, B. G., Watson, Z. L., Wheeler, L. J. & Behbakht, K. PARP inhibitors: clinical utility and possibilities of overcoming resistance. Genecol. Oncol. 147, 695–704 (2017).
Bryant, H. E. et al. Helleday. Specific killing of BRCA2-deficient tumours with inhibitors of polyADP-ribose polymerase. Nature 434, 913–917 (2005).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Hollstein, M., Sidransky, D., Vogelstein, B. & Harris, C. p53 mutations in human cancers. Science 253, 49–53 (1991).
Vance, S. et al. Selective radiosensitization of p53 mutant pancreatic cancer cells by combined inhibition of Chk1 and PARP1. Cell Cycl. 10, 4321–4329 (2011).
Mendes-Pereira, A. M. et al. A. Ashworth. Synthetic lethal targeting of PTEN mutant cells with PARP inhibitors. EMBO Mol. Med. 1, 315–322 (2009).
Passeri, D. et al. Concepts and molecular aspects in the polypharmacology of PARP1 inhibitors. ChemMedChem. 11, 1219–1226 (2016).
Silk, A. D., Holland, A. J. & Cleveland, D. W. Requirement for NuMA in maintenance and establishment of mammalian spindle poles. J. Cell. Biol. 184, 677–690 (2009).
Visochek, L. et al. Exclusive destruction of mitotic spindles in human cancer cells. Oncotarget 8, 20813–20824 (2017).
Acknowledgements
Our research was supported by the Israel Science Foundation, the Israeli Ministry of Health and NIH grant 1R21DA027776 (M.C.-A.). The Pascal lab is supported by CIHR grant BMA342854 (J.M.P).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cohen-Armon, M., Yeheskel, A. & Pascal, J.M. Signal-induced PARP1-Erk synergism mediates IEG expression. Sig Transduct Target Ther 4, 8 (2019). https://doi.org/10.1038/s41392-019-0042-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-019-0042-0