Abstract
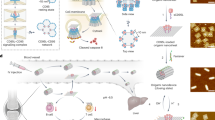
Cell spheroids bridge the discontinuity between in vitro systems and in vivo animal models. However, inducing cell spheroids by nanomaterials remains an inefficient and poorly understood process. Here we use cryogenic electron microscopy to determine the atomic structure of helical nanofibres self-assembled from enzyme-responsive d-peptides and fluorescent imaging to show that the transcytosis of d-peptides induces intercellular nanofibres/gels that potentially interact with fibronectin to enable cell spheroid formation. Specifically, d-phosphopeptides, being protease resistant, undergo endocytosis and endosomal dephosphorylation to generate helical nanofibres. On secretion to the cell surface, these nanofibres form intercellular gels that act as artificial matrices and facilitate the fibrillogenesis of fibronectins to induce cell spheroids. No spheroid formation occurs without endo- or exocytosis, phosphate triggers or shape switching of the peptide assemblies. This study—coupling transcytosis and morphological transformation of peptide assemblies—demonstrates a potential approach for regenerative medicine and tissue engineering.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The cryo-EM models of the structures reported in this study are deposited in the Protein Data Bank (PDB) under deposition ID 7L17 for class 1 NBD–ffsy filaments, 8DST for class 2 NBD–ffsy filaments and 8FOF for BP–ffsy filaments. The data generated in this study are available in the Article and its Supplementary Information. Source data are provided with this paper.
References
Rookmaaker, M. B., Schutgens, F., Verhaar, M. C. & Clevers, H. Development and application of human adult stem or progenitor cell organoids. Nat. Rev. Nephrol. 11, 546–554 (2015).
Baharvand, H., Hashemi, S. M., Kazemi Ashtiani, S. & Farrokhi, A. Differentiation of human embryonic stem cells into hepatocytes in 2D and 3D culture systems in vitro. Int. J. Dev. Biol. 50, 645–652 (2006).
Benya, P. D. & Shaffer, J. D. Dedifferentiated chondrocytes reexpress the differentiated collagen phenotype when cultured in agarose gels. Cell 30, 215–224 (1982).
Nelson, C. M. & Bissell, M. J. Modeling dynamic reciprocity: engineering three-dimensional culture models of breast architecture, function, and neoplastic transformation. Semin. Cancer Biol. 15, 342–352 (2005).
Imamura, Y. et al. Comparison of 2D- and 3D-culture models as drug-testing platforms in breast cancer. Oncol. Rep. 33, 1837–1843 (2015).
Weaver, V. M. et al. Reversion of the malignant phenotype of human breast cells in three-dimensional culture and in vivo by integrin blocking antibodies. J. Cell Biol. 137, 231–245 (1997).
Bhadriraju, K. & Chen, C. S. Engineering cellular microenvironments to improve cell-based drug testing. Drug Discov. Today 7, 612–620 (2002).
Yamada, K. M. & Cukierman, E. Modeling tissue morphogenesis and cancer in 3D. Cell 130, 601–610 (2007).
Cukierman, E., Pankov, R., Stevens, D. R. & Yamada, K. M. Taking cell-matrix adhesions to the third dimension. Science 294, 1708–1712 (2001).
Gao, D. et al. Organoid cultures derived from patients with advanced prostate cancer. Cell 159, 176–187 (2014).
Karthaus, W. R. et al. Identification of multipotent luminal progenitor cells in human prostate organoid cultures. Cell 159, 163–175 (2014).
Drost, J. et al. Organoid culture systems for prostate epithelial and cancer tissue. Nat. Protoc. 11, 347–358 (2016).
Kelm, J. M., Timmins, N. E., Brown, C. J., Fussenegger, M. & Nielsen, L. K. Method for generation of homogeneous multicellular tumor spheroids applicable to a wide variety of cell types. Biotechnol. Bioeng. 83, 173–180 (2003).
Zheng, H. et al. Rotary culture promotes the proliferation of MCF-7 cells encapsulated in three-dimensional collagen–alginate hydrogels via activation of the ERK1/2-MAPK pathway. Biomed. Mater. 7, 015003 (2012).
Raghavan, S. et al. Comparative analysis of tumor spheroid generation techniques for differential in vitro drug toxicity. Oncotarget 7, 16948–16961 (2016).
Marchi, F. & Leblond, C. P. Collagen biogenesis and assembly into fibrils as shown by ultrastructural and 3H-proline radioautographic studies on the fibroblasts of the rat food pad. Am. J. Anat. 168, 167–197 (1983).
McCaffrey, G. et al. Tight junctions contain oligomeric protein assembly critical for maintaining blood–brain barrier integrity in vivo. J. Neurochem. 103, 2540–2555 (2007).
Seger, D., Seger, R. & Shaltiel, S. The CK2 phosphorylation of vitronectin. Promotion of cell adhesion via the αvβ3-phosphatidylinositol 3-kinase pathway. J. Biol. Chem. 276, 16998–17006 (2001).
Weber, G. F. et al. Phosphorylation-dependent interaction of osteopontin with its receptors regulates macrophage migration and activation. J. Leukoc. Biol. 72, 752–761 (2002).
Yalak, G. & Vogel, V. Extracellular phosphorylation and phosphorylated proteins: not just curiosities but physiologically important. Sci. Signal. 5, re7 (2012).
Wu, D. et al. Polymers with controlled assembly and rigidity made with click-functional peptide bundles. Nature 574, 658–662 (2019).
Du, X., Zhou, J., Shi, J. & Xu, B. Supramolecular hydrogelators and hydrogels: from soft matter to molecular biomaterials. Chem. Rev. 115, 13165–13307 (2015).
Cheetham, A. G., Zhang, P., Lin, Y. A., Lock, L. L. & Cui, H. Supramolecular nanostructures formed by anticancer drug assembly. J. Am. Chem. Soc. 135, 2907–2910 (2013).
Tibbitt, M. W. & Anseth, K. S. Hydrogels as extracellular matrix mimics for 3D cell culture. Biotechnol. Bioeng. 103, 655–663 (2009).
Jayawarna, V. et al. Nanostructured hydrogels for three‐dimensional cell culture through self‐assembly of fluorenylmethoxycarbonyl–dipeptides. Adv. Mater. 18, 611–614 (2006).
Smith, D. J. et al. A multiphase transitioning peptide hydrogel for suturing ultrasmall vessels. Nat. Nanotechnol. 11, 95–102 (2016).
Alvarez, Z. et al. Bioactive scaffolds with enhanced supramolecular motion promote recovery from spinal cord injury. Science 374, 848–856 (2021).
Winkler, S. M., Harrison, M. R. & Messersmith, P. B. Biomaterials in fetal surgery. Biomater. Sci. 7, 3092–3109 (2019).
Wang, H., Feng, Z. & Xu, B. Intercellular instructed-assembly mimics protein dynamics to induce cell spheroids. J. Am. Chem. Soc. 141, 7271–7274 (2019).
Wang, H. et al. An in situ dynamic continuum of supramolecular phosphoglycopeptides enables formation of 3D cell spheroids. Angew. Chem. Int. Ed. 56, 16297–16301 (2017).
He, H. et al. Enzymatic noncovalent synthesis. Chem. Rev. 120, 9994–10078 (2020).
Zhang, Y., Kuang, Y., Gao, Y. & Xu, B. Versatile small-molecule motifs for self-assembly in water and the formation of biofunctional supramolecular hydrogels. Langmuir 27, 529–537 (2011).
Reches, M. & Gazit, E. Casting metal nanowires within discrete self-assembled peptide nanotubes. Science 300, 625–627 (2003).
Gao, Y., Shi, J., Yuan, D. & Xu, B. Imaging enzyme-triggered self-assembly of small molecules inside live cells. Nat. Commun. 3, 1033 (2012).
Van Itallie, C. M. & Anderson, J. M. Occludin confers adhesiveness when expressed in fibroblasts. J. Cell Sci. 110, 1113–1121 (1997).
Mrsny, R. J. et al. A key claudin extracellular loop domain is critical for epithelial barrier integrity. Am. J. Pathol. 172, 905–915 (2008).
Lee, M., Ghosh, U., Thurber, K. R., Kato, M. & Tycko, R. Molecular structure and interactions within amyloid-like fibrils formed by a low-complexity protein sequence from FUS. Nat. Commun. 11, 5735 (2020).
Roder, C. et al. Atomic structure of PI3-kinase SH3 amyloid fibrils by cryo-electron microscopy. Nat. Commun. 10, 3754 (2019).
Cao, Q., Boyer, D. R., Sawaya, M. R., Ge, P. & Eisenberg, D. S. Cryo-EM structures of four polymorphic TDP-43 amyloid cores. Nat. Struct. Mol. Biol. 26, 619–627 (2019).
Fitzpatrick, A. W. P. et al. Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 547, 185–190 (2017).
Falcon, B. et al. Novel tau filament fold in chronic traumatic encephalopathy encloses hydrophobic molecules. Nature 568, 420–423 (2019).
Wang, F. et al. Deterministic chaos in the self-assembly of β sheet nanotubes from an amphipathic oligopeptide. Matter 4, 3217–3231 (2021).
To, W. S. & Midwood, K. S. Plasma and cellular fibronectin: distinct and independent functions during tissue repair. Fibrogenesis Tissue Repair 4, 21 (2011).
Lu, Y. et al. Vacuolin-1 potently and reversibly inhibits autophagosome-lysosome fusion by activating RAB5A. Autophagy 10, 1895–1905 (2014).
Du, X. et al. In situ generated d‐peptidic nanofibrils as multifaceted apoptotic inducers to target cancer cells. Cell Death Dis. 8, e2614–e2614 (2017).
Feng, Z., Wang, H., Chen, X. & Xu, B. Self-assembling ability determines the activity of enzyme-instructed self-assembly for inhibiting cancer cells. J. Am. Chem. Soc. 139, 15377–15384 (2017).
Shigemitsu, H. et al. An adaptive supramolecular hydrogel comprising self-sorting double nanofibre networks. Nat. Nanotechnol. 13, 165–172 (2018).
Epstein, I. R. & Xu, B. Reaction–diffusion processes at the nano- and microscales. Nat. Nanotechnol. 11, 312–319 (2016).
Liang, G., Ren, H. & Rao, J. A biocompatible condensation reaction for controlled assembly of nanostructures in living cells. Nat. Chem. 2, 54–60 (2010).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Ottinger, E. A., Shekels, L. L., Bernlohr, D. A. & Barany, G. Synthesis of phosphotyrosine-containing peptides and their use as substrates for protein tyrosine phosphatases. Biochemistry 32, 4354–4361 (1993).
Liu, S. et al. Enzymatically forming intranuclear peptide assemblies for selectively killing human induced pluripotent stem cells. J. Am. Chem. Soc. 143, 15852–15862 (2021).
Basu Ray, G., Chakraborty, I. & Moulik, S. P. Pyrene absorption can be a convenient method for probing critical micellar concentration (cmc) and indexing micellar polarity. J. Colloid Interface Sci. 294, 248–254 (2006).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Acknowledgements
This work was partially supported by National Institutes of Health (NIH) grants CA142746 (B.X.), GM122510 (E.H.E.) and GM138756 (F.W.), as well as NSF grant DMR-2011846 (B.X.). This research was, in part, supported by the National Cancer Institute’s National Cryo-EM Facility at the Frederick National Laboratory for Cancer Research under contract 75N91019D00024. The cryo-EM imaging was, in part, done at the Molecular Electron Microscopy Core Facility at the University of Virginia. The cryo-EM screening process was, in part, supported by the O’Neal Comprehensive Cancer Center at the University of Alabama at Birmingham. We thank P. D. Camilli for providing the TKO cell line. We thank J. T. Hsien for providing HeLa–GFP, PC-3–DsRed and Saos-2–GFP.
Author information
Authors and Affiliations
Contributions
B.X. and J.G. conceived the study. J.G., under the supervision of B.X., designed and performed the chemical synthesis, generated the images in cell-free and cell-based assays and analysed the results. Y.H. and M.Y. assisted in the chemical synthesis. H.H. and W.T. performed the liquid chromatography–mass spectrometry analysis. F.W. and E.H.E. performed the cryo-EM reconstructions and model building. J.G., F.W., E.H.E. and B.X. wrote the manuscript with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Nanotechnology thanks Deling Kong and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–43, Tables 1 and 2, materials and instruments and references.
Supplementary Video 1
Suspended HS-5 cells incubated with NBD–ffspy (500 µM) over the first 6 h.
Supplementary Data 1
Full scans of the western blot results in Supplementary Fig. 22d.
Supplementary Data 2
Full scans of the western blot results in Supplementary Fig. 34a.
Supplementary Data 3
Full scans of the western blot results in Supplementary Fig. 40c.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Guo, J., Wang, F., Huang, Y. et al. Cell spheroid creation by transcytotic intercellular gelation. Nat. Nanotechnol. 18, 1094–1104 (2023). https://doi.org/10.1038/s41565-023-01401-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-023-01401-7