Abstract
Disease-modifying management aims to prevent deterioration and progression of the disease, and not just to relieve symptoms. We present a solution for the management by a methodology that allows the prediction of progression risk and morphology in individuals using a latent extrapolation approach. To this end, we combined a regularized generative adversarial network and a latent nearest neighbour algorithm for joint optimization to generate plausible images of future time points. We evaluated our method on osteoarthritis data from a multicenter longitudinal study (the Osteoarthritis Initiative). With presymptomatic baseline data, our model is generative and considerably outperforms the end-to-end learning model in discriminating the progressive cohort. Two experiments were performed with seven radiologists. When no synthetic follow-up radiographs were provided, our model performed better than all seven radiologists. In cases in which the synthetic follow-ups generated by our model were made available to the radiologist for diagnosis support, the specificity and sensitivity of all readers in discriminating progressors increased from 72.3% to 88.6% and from 42.1% to 51.6%, respectively. Our results open up a new possibility of using model-based morphology and risk prediction to make predictions about disease occurrence, as demonstrated by the example of osteoarthritis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All datasets used in this study are publicly available: the dataset of knee radiographs of the OAI and MOST datasets can be requested from https://nda.nih.gov/oai/ and https://agingresearchbiobank.nia.nih.gov/studies/most/.
Code availability
The code used in this study is made fully publicly available under https://github.com/peterhan91/disease_progression (ref. 43).
References
Grigorescu, S., Trasnea, B., Cocias, T. & Macesanu, G. A survey of deep learning techniques for autonomous driving. J. Field Robot. 37, 362–386 (2020).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. In Advances in Neural Information Processing Systems 25 (NIPS, 2012).
Senior, A. W. et al. Improved protein structure prediction using potentials from deep learning. Nature 577, 706–710 (2020).
Jumper, J. et al. Highly accurate protein structure prediction with alphafold. Nature 596, 583–589 (2021).
Lundberg, S. M. et al. Explainable machine-learning predictions for the prevention of hypoxaemia during surgery. Nat. Biomed. Eng. 2, 749–760 (2018).
Recht, B., Roelofs, R., Schmidt, L. & Shankar, V. Do CIFAR-10 classifiers generalize to CIFAR-10? Preprint at https://arxiv.org/abs/1806.00451 (2018).
Deng, J. et al. Imagenet: a large-scale hierarchical image database. In 2009 IEEE Conference on Computer Vision and Pattern Recognition 248–255 (IEEE, 2009).
Petersen, R. C. et al. Alzheimer’s disease neuroimaging initiative (ADNI): clinical characterization. Neurology 74, 201–209 (2010).
Menze, B. H. et al. The multimodal brain tumor image segmentation benchmark (brats). IEEE Trans. Med. Imaging 34, 1993–2024 (2014).
Casey, B. et al. The adolescent brain cognitive development (ABCD) study: imaging acquisition across 21 sites. Dev. Cogn. Neurosci. 32, 43–54 (2018).
Eckstein, F., Wirth, W. & Nevitt, M. C. Recent advances in osteoarthritis imaging—the osteoarthritis initiative. Nat. Rev. Rheumatol. 8, 622–630 (2012).
Yim, J. et al. Predicting conversion to wet age-related macular degeneration using deep learning. Nat. Med. 26, 892–899 (2020).
Nebelung, S. et al. Functional MR imaging mapping of human articular cartilage response to loading. Radiology 282, 464–474 (2017).
Goodfellow, I. et al. Generative adversarial nets. In Advances in Neural Information Processing Systems 27 (NIPS, 2014).
Karras, T., Laine, S. & Aila, T. A style-based generator architecture for generative adversarial networks. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 4401–4410 (IEEE, 2019).
Karras, T. et al. Analyzing and improving the image quality of stylegan. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 8110–8119 (IEEE, 2020).
Schiratti, J.-B., Allassonniere, S., Colliot, O. & Durrleman, S. Learning spatiotemporal trajectories from manifold-valued longitudinal data. In Advances in Neural Information Processing Systems 28 (NIPS, 2015).
Ravi, D. et al. Degenerative adversarial NeuroImage nets: generating images that mimic disease progression. In International Conference on Medical Image Computing and Computer-Assisted Intervention 164–172 (Springer, 2019).
Isola, P., Zhu, J.-Y., Zhou, T. & Efros, A. A.Image-to-image translation with conditional adversarial networks. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 1125–1134 (IEEE, 2017).
Kingma, D. P. & Welling, M. Auto-encoding variational Bayes. Preprint at https://arxiv.org/abs/1312.6114 (2013).
Couronné, R., Vernhet, P. & Durrleman, S. Longitudinal self-supervision to disentangle inter-patient variability from disease progression. In International Conference on Medical Image Computing and Computer-Assisted Intervention 231–241 (Springer, 2021).
Venkatraghavan, V. et al. Event-based modeling with high-dimensional imaging biomarkers for estimating spatial progression of dementia. In International Conference on Information Processing in Medical Imaging 169–180 (Springer, 2019).
Han, T. et al. Advancing diagnostic performance and clinical usability of neural networks via adversarial training and dual batch normalization. Nat. Commun. 12, 1–11 (2021).
Tsipras, D., Santurkar, S., Engstrom, L., Turner, A. & Madry, A. Robustness may be at odds with accuracy. Preprint at https://arxiv.org/abs/1805.12152 (2018).
Hsu, D. & A Marshall, G. Primary and secondary prevention trials in Alzheimer disease: looking back, moving forward. Curr. Alzheimer Res. 14, 426–440 (2017).
Pimpin, L. et al. Burden of liver disease in europe: epidemiology and analysis of risk factors to identify prevention policies. J. Hepatol. 69, 718–735 (2018).
Compston, J. et al. UK clinical guideline for the prevention and treatment of osteoporosis. Arch. Osteoporos. 12, 43 (2017).
Gorenek, B. et al. European Heart Rhythm Association (EHRA)/European Association of Cardiovascular Prevention and Rehabilitation (EACPR) position paper on how to prevent atrial fibrillation endorsed by the Heart Rhythm Society (HRS) and Asia Pacific Heart Rhythm Society (APHRS). Eur. J. Preventive Cardiol. 24, 4–40 (2017).
Alkhatib, A. et al. Functional foods and lifestyle approaches for diabetes prevention and management. Nutrients 9, 1310 (2017).
Donahue, J., Krähenbühl, P. & Darrell, T. Adversarial feature learning. Preprint at https://arxiv.org/abs/1605.09782 (2016).
Donahue, J. & Simonyan, K. Large scale adversarial representation learning. Preprint at https://arxiv.org/abs/1907.02544 (2019).
Xia, W. et al. GAN inversion: a survey. In IEEE Transactions on Pattern Analysis and Machine Intelligence (IEEE, 2022).
Tiulpin, A., Melekhov, I. & Saarakkala, S. Kneel: knee anatomical landmark localization using hourglass networks. In Proc. IEEE/CVF International Conference on Computer Vision Workshops (IEEE, 2019).
Segal, N. A. et al. The multicenter osteoarthritis study (most): opportunities for rehabilitation research. PM R 5, 647–654 (2013).
Heusel, M., Ramsauer, H., Unterthiner, T., Nessler, B. & Hochreiter, S. GANs trained by a two time-scale update rule converge to a local Nash equilibrium. In Advances in Neural Information Processing Systems 6626–6637 (NIPS, 2017).
Sajjadi, M. S., Bachem, O., Lucic, M., Bousquet, O. & Gelly, S. Assessing generative models via precision and recall. Preprint at https://arxiv.org/abs/1806.00035 (2018).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2014).
Zhang, R., Isola, P., Efros, A. A., Shechtman, E. & Wang, O. The unreasonable effectiveness of deep features as a perceptual metric. In Proc. IEEE Conference on Computer Vision and Pattern Recognition 586–595 (IEEE, 2018).
Tiulpin, A. et al. Multimodal machine learning-based knee osteoarthritis progression prediction from plain radiographs and clinical data. Sci. Rep. 9, 1–11 (2019).
Ardila, D. et al. End-to-end lung cancer screening with three-dimensional deep learning on low-dose chest computed tomography. Nat. Med. 25, 954–961 (2019).
Kniep, H. C. et al. Radiomics of brain MRI: utility in prediction of metastatic tumor type. Radiology 290, 479–487 (2019).
Paszke, A et al. PyTorch: an imperative style, high-performance deep learning library. In Advances in Neural Information Processing Systems 8026–8037 (NIPS, 2019).
Han, T. Peterhan91/Disease_Progression: Initial Release (Zenodo, 2022); https://doi.org/10.5281/zenodo.6983073
Foreman, S. et al. Accelerated knee osteoarthritis is nearly always associated with meniscal breakdown: data from the osteoarthritis initiative. Osteoarthr. Cartil. 26, S443–S444 (2018).
Driban, J. B. et al. Risk factors and the natural history of accelerated knee osteoarthritis: a narrative review. BMC Musculoskelet. Disord. 21, 1–11 (2020).
Acknowledgements
This work is supported in part by the German Federal Ministry of Health (DEEP LIVER, grant no. ZMVI1-2520DAT111) and the Max-Eder-Programme of the German Cancer Aid (grant no. 70113864 received by J.N.K).
Author information
Authors and Affiliations
Contributions
T.H and D.T devised the concept of the study, D.T, S.N, M.S.-H, M.Z, F.P, S.K and M.T performed the reader tests. T.H wrote the code and completed the performance studies. T.H and D.T performed the statistical analysis. T.H and D.T wrote the first draft of the manuscript. All authors contributed to correcting the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.N.K declares consulting services for Owkin, France, and Panakeia, UK. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Machine Intelligence thanks Nico van den Berg, Xinxing Xu and Florian Dubost for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Schematic Overview of Model-Based OA Predictions on Radiographs.
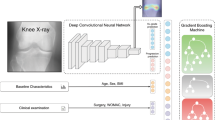
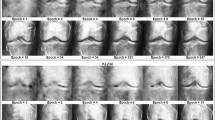
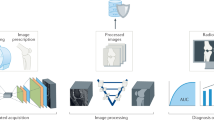
(a) On the original knee radiographs (bilateral, posteroanterior projections, and fixed flexion), the image region containing the knee joints is identified using an Hourglass network. Knee joints are cropped accordingly, and a generative model is trained to generate synthetic radiographs from a vector in a low-dimensional latent space. Corresponding original knee X-rays are then back mapped to the point in latent space best matching the synthetic image. (b) In the latent inference module (right panel), scans of the same patient but at different time points are used to generate a vector field in latent space. A knee radiograph with unknown OA onset and progression can be mapped to its latent nearest neighbor at a follow-up visit using the vector field. (c) Predicting multiple follow-up scans via iterative latent inference. In six follow-up visits, rapid progression of OA on the left knee of a 72-year-old male participant was correctly predicted by iteratively applying the latent inference module. Given an input knee X-ray, our latent inference performs a nearest neighbor search and makes an image prediction based on its nearest neighbor trajectory. When considering multiple follow-ups, we recursively apply latent inference on previously predicted knee images to get the next time point prediction. Note, images in red frames are generated sequentially by the proposed method and the given baseline image.
Extended Data Fig. 2 The architecture of the generator model.
(a) An architectural layout for the style-based G employed in our work. On the right, the latent code w and random Gaussian noise ni×i, for example, n4×4and n8×8, are injected into every resolution block of G. (b)-(c) Details about resolution blocks. The architecture of a modulated 4 × 4 resolution block and higher resolution blocks are shown in b and c, respectively.
Extended Data Fig. 3 A participant shows sudden OA progression within two years.
(a) This OAI participant (male, 51 years old) exhibited accelerated knee OA between six- and eight-year follow-ups. This sudden OA onset can be seen in the final 8-year visit radiograph indicated by the red arrows in the 1st row of (a). The 2nd row visualizes the images as predicted by our model. Progression towards OA was correctly predicted between 4 to 8 years of synthesized follow-ups generated by our method, yet not at the same rate as the true progression. (b) The proposed OA risk curves between real visits and generated predictions. The real curve in blue colour precisely captured the rapid OA onset within this participant. A quantitative evaluation of our predictions (red curve) shows a moderate OA increase that is slower than the true course.
Extended Data Fig. 4 Confusion matrices for each radiologist and experiment.
Seven radiologists were tasked to judge whether a given knee radiograph is likely to undergo OA progression in the future (prog) or not (non-prog), based on the baseline radiograph alone (a) or the baseline radiograph and the synthesized predicted radiograph (b). For both experiments, we randomly selected 486 knee radiographs from the OAI test set.
Extended Data Fig. 5
Performance metrics of individual radiologists in predicting OA onset and progression as a function of model assistance.
Extended Data Fig. 6
List of Symbols and Notations.
Extended Data Fig. 7 Details of the training process and performance metrics of GAN.
(a) Scores for real (blue) and synthetic radiographs along the training process of the GAN. Scores are stabilized after 10 million images. (b) Values of G (PL penalty) and D (R1 penalty) penalties along the training process. (c) Fréchet Inception Distance (FID) scores along the training process to measure convergence to realistic radiographs. The convergence of the GAN is indicated by the FID scores plateauing after 10 million images are shown to the Discriminator. (d) Precision (F1/8) and recall (F8) metrics for different GAN models. The model with the highest F1/8 and F8 scores (as indicated by the red arrow) was selected. The inset tiled plot is a close-up of the right upper corner. (e) Further characterization of the best-performing GAN model via the corresponding distributional precision and recall curve. (f) Examples of GAN-generated knee radiographs with a spatial resolution of 256 × 256 pixels.
Extended Data Fig. 8 Workflow pipeline to visualize the embedding procedure of a representative knee radiograph into the latent space.
(a) After computing the learned perceptual image patch similarity (LPIPS) loss, we freeze the parameters in G and the pre-trained VGG net to exclusively optimize the intermediate latent w via backpropagation. Representative original and synthetic radiographs are framed in grey and red, respectively. (b) In both OAI (n=52,981 radiographs) and MOST (n=19,340 radiographs) datasets, we characterize the embedding procedure by computing the Structural Similarity Index (SSIM) between original and synthetic radiographs. Lines and boxes indicate medians and upper or lower quartiles, while whiskers detail the range of the data. In the box plot, minima are the minimum value, maxima are the maximum value. The lower and upper lines correspond to the first and third quartiles; the upper and lower whiskers extend to values no farther than the 1.5 interquartile range (IQR). (c) Selected reconstructions from the OAI StyleGAN model. As shown, anatomical regions such as the tibia, femur, and knee joint are consistent between original and reconstructed radiographs. Differences between real and synthesized images were assessed to be minor by the radiologists.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12, Discussion and Tables 1–5.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Han, T., Kather, J.N., Pedersoli, F. et al. Image prediction of disease progression for osteoarthritis by style-based manifold extrapolation. Nat Mach Intell 4, 1029–1039 (2022). https://doi.org/10.1038/s42256-022-00560-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42256-022-00560-x
This article is cited by
-
Enhancing domain generalization in the AI-based analysis of chest radiographs with federated learning
Scientific Reports (2023)
-
Collaborative training of medical artificial intelligence models with non-uniform labels
Scientific Reports (2023)