Abstract
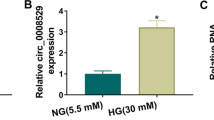
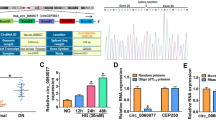
Circular RNA (circRNA) is key regulator of diabetic nephropathy (DN) progression. However, the role of circ_0008529 in DN progression remains to be better deciphered. Cell viability, cell cycle, apoptosis and inflammation were measured by MTS assay, flow cytometry and corresponding assay kits. RT-qPCR was used to assess the expression of circ_0008529, miR-185-5p and SMAD family member 2 (SMAD2). Also, western blotting was performed to measure protein expression. Target relationship was validated by RNA pull-down assay, dual-luciferase reporter assay and RNA immunoprecipitation assay. Urinary exosome was isolated using ultracentrifugation method and identified by transmission electron microscopy. Receiver operating characteristic curve was used to analyze the diagnostic value of circ_0008529 in DN patients. Circ_0008529 and SMAD2 were upregulated, while miR-185-5p was downregulated in high glucose (HG)-induced renal tubular HK-2 cells. Under HG treatment, cell viability and cell cycle process were suppressed, while apoptosis, inflammation and extracellular matrix accumulation were enhanced. However, interfering circ_0008529 could attenuate HG-induced effects, and this protection was abated by miR-185 inhibition or SMAD2 re-expression. Mechanically, circ_0008529 and SMAD2 were competing endogenous RNAs for miR-185-5p via target binding, and circ_0008529 regulated SMAD2 expression via miR-185-5p. Notably, circ_0008529 expression was upregulated in urinary exosomes of DN patients, and showed diagnostic value (Sensitivity: 70.21%; Specificity: 86.67%). Circ_0008529 might be a potential target for DN, which regulated DN progression via miR-185-5p/SMAD2 pathway.







Similar content being viewed by others
Abbreviations
- circRNA:
-
Circular RNA
- miRNA:
-
MicroRNA
- SMAD2:
-
SMAD family member 2
- DN:
-
Diabetic nephropathy
- RT-qPCR:
-
Reverse transcription-quantitative PCR
- C-casp3:
-
Cleaved-caspase-3
- α-SMA:
-
α- Smooth muscle actin
- GAPDH:
-
Glyceraldehyde-phosphate dehydrogenase
- ROC:
-
Receiver operating characteristic
- AUC:
-
Area under the curve
- CI:
-
Confidence intervals
- TEM:
-
Transmission electron microscopy
- ELISA:
-
Enzyme-linked immunosorbent assay
- TNF-α:
-
Tumor necrosis factor-α
- IL-6:
-
Interleukin IL-6
- WT:
-
Wild type
- MUT:
-
Mutant type
- RIP:
-
RNA immunoprecipitation
- ANOVA:
-
Analysis of variance
References
Akash MSH, Rehman K, Liaqat A (2018) Tumor necrosis factor-alpha: role in development of insulin resistance and pathogenesis of type 2 diabetes mellitus. J Cell Biochem 119:105–110. https://doi.org/10.1002/jcb.26174
Gewin LS (2018) Renal fibrosis: primacy of the proximal tubule. Matrix Biol 68–69:248–262. https://doi.org/10.1016/j.matbio.2018.02.006
Hamid Akash MS, Rehman K, Jabeen K, Fiayyaz F, Sabir S, Ejaz Ul Haq M (2021) Assessment of knowledge, attitude and practice of Pakistani population about the risk factors, causes, complications and management of diabetes mellitus. J Pak Med Assoc 71:286–296. https://doi.org/10.47391/JPMA.434
Hu W, Han Q, Zhao L, Wang L (2019) Circular RNA circRNA_15698 aggravates the extracellular matrix of diabetic nephropathy mesangial cells via miR-185/TGF-beta1. J Cell Physiol 234:1469–1476. https://doi.org/10.1002/jcp.26959
Jia Y, Guan M, Zheng Z, Zhang Q, Tang C, Xu W, Xiao Z, Wang L, Xue Y (2016) miRNAs in urine extracellular vesicles as predictors of early-stage diabetic nephropathy. J Diabetes Res 2016:7932765. https://doi.org/10.1155/2016/7932765
Jin J, Sun H, Shi C, Yang H, Wu Y, Li W, Dong YH, Cai L, Meng XM (2020) Circular RNA in renal diseases. J Cell Mol Med 24:6523–6533. https://doi.org/10.1111/jcmm.15295
Kato M, Natarajan R (2019) Epigenetics and epigenomics in diabetic kidney disease and metabolic memory. Nat Rev Nephrol 15:327–345. https://doi.org/10.1038/s41581-019-0135-6
Kinget L, Roussel E, Verbiest A, Albersen M, Rodriguez-Antona C, Grana-Castro O, Inglada-Perez L, Zucman-Rossi J, Couchy G, Job S, de Reynies A, Laenen A, Baldewijns M, Beuselinck B (2021) MicroRNAs targeting HIF-2alpha, VEGFR1 and/or VEGFR2 as potential predictive biomarkers for VEGFR tyrosine kinase and HIF-2alpha inhibitors in metastatic clear-cell renal cell carcinoma. Cancers. https://doi.org/10.3390/cancers13123099
Lan HY (2012) Transforming growth factor-beta/Smad signalling in diabetic nephropathy. Clin Exp Pharmacol Physiol 39:731–738. https://doi.org/10.1111/j.1440-1681.2011.05663.x
Lei GS, Kline HL, Lee CH, Wilkes DS, Zhang C (2016) Regulation of collagen V expression and epithelial-mesenchymal transition by miR-185 and miR-186 during idiopathic pulmonary fibrosis. Am J Pathol 186:2310–2316. https://doi.org/10.1016/j.ajpath.2016.04.015
Li G, Qin Y, Qin S, Zhou X, Zhao W, Zhang D (2020) Circ_WBSCR17 aggravates inflammatory responses and fibrosis by targeting miR-185-5p/SOX6 regulatory axis in high glucose-induced human kidney tubular cells. Life Sci 259:118269. https://doi.org/10.1016/j.lfs.2020.118269
Liu LD, Pang YX, Zhao XR, Li R, Jin CJ, Xue J, Dong RY, Liu PS (2019) Curcumin induces apoptotic cell death and protective autophagy by inhibiting AKT/mTOR/p70S6K pathway in human ovarian cancer cells. Arch Gynecol Obstet 299:1627–1639. https://doi.org/10.1007/s00404-019-05058-3
Liu R, Zhang M, Ge Y (2021) Circular RNA HIPK3 exacerbates diabetic nephropathy and promotes proliferation by sponging miR-185. Gene 765:145065. https://doi.org/10.1016/j.gene.2020.145065
Loganathan TS, Sulaiman SA, Abdul Murad NA, Shah SA, Abdul Gafor AH, Jamal R, Abdullah N (2020) Interactions among non-coding RNAs in diabetic nephropathy. Front Pharmacol 11:191. https://doi.org/10.3389/fphar.2020.00191
Meng XM, Huang XR, Chung AC, Qin W, Shao X, Igarashi P, Ju W, Bottinger EP, Lan HY (2010) Smad2 protects against TGF-beta/Smad3-mediated renal fibrosis. J Am Soc Nephrol 21:1477–1487. https://doi.org/10.1681/ASN.2009121244
Peng F, Gong W, Li S, Yin B, Zhao C, Liu W, Chen X, Luo C, Huang Q, Chen T, Sun L, Fang S, Zhou W, Li Z, Long H (2020) circRNA_010383 acts as a sponge for miR-135a and its downregulated expression contributes to renal fibrosis in diabetic nephropathy. Diabetes. https://doi.org/10.2337/db200203
Rehman K, Haider K, Jabeen K, Akash MSH (2020a) Current perspectives of oleic acid: regulation of molecular pathways in mitochondrial and endothelial functioning against insulin resistance and diabetes. Rev Endocr Metab Disord 21:631–643. https://doi.org/10.1007/s11154-020-09549-6
Rehman K, Jabeen K, Awan FR, Hussain M, Saddique MA, Akash MSH (2020b) Biochemical investigation of rs1801282 variations in PPAR-gamma gene and its correlation with risk factors of diabetes mellitus in coronary artery disease. Clin Exp Pharmacol Physiol 47:1517–1529. https://doi.org/10.1111/1440-1681.13339
Ren H, Wang Q (2021) Non-Coding RNA and diabetic kidney disease. DNA Cell Biol 40:553–567. https://doi.org/10.1089/dna.2020.5973
Wen S, Li S, Li L, Fan Q (2020) circACTR2: a novel mechanism regulating high glucose-induced fibrosis in renal tubular cells via pyroptosis. Biol Pharm Bull 43:558–564. https://doi.org/10.1248/bpb.b19-00901
Yuan Q, Xu T, Chen Y, Qu W, Sun D, Liu X, Sun L (2020) MiR-185-5p ameliorates endoplasmic reticulum stress and renal fibrosis by downregulation of ATF6. Lab Invest 100:1436–1446. https://doi.org/10.1038/s41374-020-0447-y
Zhao D, Guo J, Liu L, Huang Y (2021) Rosiglitazone attenuates high glucose-induced proliferation, inflammation, oxidative stress and extracellular matrix accumulation in mouse mesangial cells through the Gm26917/miR-185-5p pathway. Endocr J. https://doi.org/10.1507/endocrj.EJ20-0783
Zhou L, Liu S, Han M, Ma Y, Feng S, Zhao J, Lu H, Yuan X, Cheng J (2018) miR-185 Inhibits fibrogenic activation of hepatic stellate cells and prevents liver fibrosis. Mol Ther Nucleic Acids 10:91–102. https://doi.org/10.1016/j.omtn.2017.11.010
Zhou J, Zhou H, Liu Y, Liu C (2021a) Inhibition of CTCF-regulated miRNA-185-5p mitigates renal interstitial fibrosis of chronic kidney disease. Epigenomics 13:859–873. https://doi.org/10.2217/epi-2020-0243
Zhou S, Fang J, Hu M, Pan S, Liu D, Xing G, Liu Z (2021b) Determining the influence of high glucose on exosomal lncRNAs, mRNAs, circRNAs and miRNAs derived from human renal tubular epithelial cells. Aging (albany NY) 13:8467–8480. https://doi.org/10.18632/aging.202656
Acknowledgements
None.
Funding
None.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Niu, Z., Ren, G., Huang, L. et al. Circ_0008529 Contributes to Renal Tubular Cell Dysfunction in High Glucose Stress via miR-185-5p/SMAD2 Pathway in Diabetic Nephropathy. Biochem Genet 61, 963–978 (2023). https://doi.org/10.1007/s10528-022-10296-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-022-10296-3