Abstract
The pathophysiology of irritable bowel syndrome (IBS) is multifactorial and probably involves genetic predisposition and the effect of environmental factors. Unlike other gastrointestinal diseases with a heritable component, genetic research in IBS has been scarce and mostly characterized by small underpowered studies, leading to inconclusive results. The availability of genomic and health-related data from large international cohorts and population-based biobanks offers unprecedented opportunities for long-awaited, well-powered genetic studies in IBS. This Review focuses on the latest advances that provide compelling evidence for the importance of genes involved in the digestion of carbohydrates, ion channel function, neurotransmitters and their receptors, neuronal pathways and the control of gut motility. These discoveries have generated novel information that might be further refined for the identification of predisposed individuals and selection of management strategies for patients. This Review presents a conceptual framework, the advantages and potential limitations of modern genetic research in IBS, and a summary of available evidence.
Key points
-
The pathophysiological mechanisms associated with irritable bowel syndrome (IBS) are only partially understood; this hampers the development of targeted therapeutic strategies.
-
Genetic research can reveal actionable pathways, but findings have generally been scarce in IBS because genetic investigations have been small scale and based on single-gene approaches that have highlighted only a few putative risk genes related to serotonergic mechanisms, carbohydrate digestion or ion channels.
-
Large-scale, biobank-wide genome-wide association studies (GWAS) that focused on IBS and endophenotypic proxies of gut motility have been reported, and have led to the identification of the first set of unequivocal risk loci.
-
These loci contain several genes relevant to pathways and cell types implicated in the central and enteric nervous system activities, neurotransmitter signalling and enteric motor neuron function; these findings offer avenues for testing novel therapeutic strategies.
-
The genetic architecture of IBS identified to date through GWAS often seems to be shared with comorbid mood disorders and anxiety, which is consistent with the reported efficacy of psychotropic and cognitive therapies in IBS.
-
Polygenic scores derived from GWAS data enabled the identification of individuals at increased risk of IBS in the studied cohorts and could be further refined and validated in independent translational studies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Drossman, D. A. & Tack, J. Rome Foundation clinical diagnostic criteria for disorders of gut-brain interaction. Gastroenterology 162, 675–679 (2022).
Ford, A. C., Sperber, A. D., Corsetti, M. & Camilleri, M. Irritable bowel syndrome. Lancet 396, 1675–1688 (2020).
Drossman, D. A. Functional gastrointestinal disorders: history, pathophysiology, clinical features and Rome IV. Gastroenterology 150, 1262–1279 (2016).
Drossman, D. A. & Hasler, W. L. Rome IV–functional GI disorders: disorders of gut-brain interaction. Gastroenterology 150, 1257–1261 (2016).
Lewis, S. J. & Heaton, K. W. Stool form scale as a useful guide to intestinal transit time. Scand. J. Gastroenterol. 32, 920–924 (1997).
Buono, J. L., Carson, R. T. & Flores, N. M. Health-related quality of life, work productivity, and indirect costs among patients with irritable bowel syndrome with diarrhea. Health Qual. Life Outcomes 15, 35 (2017).
Frändemark, Å., Törnblom, H., Jakobsson, S. & Simrén, M. Work productivity and activity impairment in irritable bowel syndrome (IBS): a multifaceted problem. Am. J. Gastroenterol. 113, 1540–1549 (2018).
Camilleri, M. Diagnosis and treatment of irritable bowel syndrome: a review. JAMA 325, 865–877 (2021).
Drossman, D. A. Do psychosocial factors define symptom severity and patient status in irritable bowel syndrome? Am. J. Med. 107, 41–50 (1999).
Palsson, O. S. & Drossman, D. A. Psychiatric and psychological dysfunction in irritable bowel syndrome and the role of psychological treatments. Gastroenterol. Clin. North. Am. 34, 281–303 (2005).
Camilleri, M. Peripheral mechanisms in irritable bowel syndrome. N. Engl. J. Med. 367, 1626–1635 (2012).
Ohman, L. & Simrén, M. Pathogenesis of IBS: role of inflammation, immunity and neuroimmune interactions. Nat. Rev. Gastroenterol. Hepatol. 7, 163–173 (2010).
Bennet, S. M. P., Ohman, L. & Simren, M. Gut microbiota as potential orchestrators of irritable bowel syndrome. Gut Liver 9, 318–331 (2015).
Moayyedi, P., Simrén, M. & Bercik, P. Evidence-based and mechanistic insights into exclusion diets for IBS. Nat. Rev. Gastroenterol. Hepatol. 17, 406–413 (2020).
Lackner, J. M. et al. Type, rather than number, of mental and physical comorbidities increases the severity of symptoms in patients with irritable bowel syndrome. Clin. Gastroenterol. Hepatol. 11, 1147–1157 (2013).
D’Amato, M. Genes and functional GI disorders: from casual to causal relationship. Neurogastroenterol. Motil. 25, 638–649 (2013).
Waehrens, R., Ohlsson, H., Sundquist, J., Sundquist, K. & Zöller, B. Risk of irritable bowel syndrome in first-degree, second-degree and third-degree relatives of affected individuals: a nationwide family study in Sweden. Gut 64, 215–221 (2015).
Camilleri, M. Genetics and irritable bowel syndrome: from genomics to intermediate phenotype and pharmacogenetics. Dig. Dis. Sci. 54, 2318–2324 (2009).
Heitkemper, M. M., Kohen, R., Jun, S.-E. & Jarrett, M. E. Genetics and gastrointestinal symptoms. Annu. Rev. Nurs. Res. 29, 261–280 (2011).
Saito, Y. A. The role of genetics in IBS. Gastroenterol. Clin. North. Am. 40, 45–67 (2011).
Mawe, G. M. & Hoffman, J. M. Serotonin signalling in the gut–functions, dysfunctions and therapeutic targets. Nat. Rev. Gastroenterol. Hepatol. 10, 473–486 (2013).
Bellono, N. W. et al. Enterochromaffin cells are gut chemosensors that couple to sensory neural pathways. Cell 170, 185–198 (2017).
Grudell, A. B. M. et al. An exploratory study of the association of adrenergic and serotonergic genotype and gastrointestinal motor functions. Neurogastroenterol. Motil. 20, 213–219 (2008).
Camilleri, M. et al. Alterations in expression of p11 and SERT in mucosal biopsy specimens of patients with irritable bowel syndrome. Gastroenterology 132, 17–25 (2007).
Kim, H. J. et al. Association of distinct α2 adrenoceptor and serotonin transporter polymorphisms with constipation and somatic symptoms in functional gastrointestinal disorders. Gut 53, 829–837 (2004).
Camilleri, M. et al. Serotonin-transporter polymorphism pharmacogenetics in diarrhea-predominant irritable bowel syndrome. Gastroenterology 123, 425–432 (2002).
Mohr, S. et al. The alternative serotonin transporter promoter P2 impacts gene function in females with irritable bowel syndrome. J. Cell. Mol. Med. 25, 8047–8061 (2021).
Gunn, D. et al. Abnormalities of mucosal serotonin metabolism and 5-HT3 receptor subunit 3C polymorphism in irritable bowel syndrome with diarrhoea predict responsiveness to ondansetron. Aliment. Pharmacol. Ther. 50, 538–546 (2019).
Wohlfarth, C. et al. miR-16 and miR-103 impact 5-HT4 receptor signalling and correlate with symptom profile in irritable bowel syndrome. Sci. Rep. 7, 14680 (2017).
Kilpatrick, L. et al. The HTR3A polymorphism c. -42C>T is associated with amygdala responsiveness in patients with irritable bowel syndrome. Gastroenterology 140, 1943–1951 (2011).
Niesler, B. et al. 5-HTTLPR and STin2 polymorphisms in the serotonin transporter gene and irritable bowel syndrome: effect of bowel habit and sex. Eur. J. Gastroenterol. Hepatol. 22, 856–861 (2010).
Kapeller, J. et al. First evidence for an association of a functional variant in the microRNA-510 target site of the serotonin receptor-type 3E gene with diarrhea predominant irritable bowel syndrome. Hum. Mol. Genet. 17, 2967–2977 (2008).
Li, Y. et al. The association of serotonin transporter genetic polymorphisms and irritable bowel syndrome and its influence on tegaserod treatment in Chinese patients. Dig. Dis. Sci. 52, 2942–2949 (2007).
Boesmans, W., Owsianik, G., Tack, J., Voets, T. & Vanden Berghe, P. TRP channels in neurogastroenterology: opportunities for therapeutic intervention. Br. J. Pharmacol. 162, 18–37 (2011).
Fuentes, I. M. & Christianson, J. A. Ion channels, ion channel receptors, and visceral hypersensitivity in irritable bowel syndrome. Neurogastroenterol. Motil. 28, 1613–1618 (2016).
Locke, G. R., Ackerman, M. J., Zinsmeister, A. R., Thapa, P. & Farrugia, G. Gastrointestinal symptoms in families of patients with an SCN5A-encoded cardiac channelopathy: evidence of an intestinal channelopathy. Am. J. Gastroenterol. 101, 1299–1304 (2006).
Beyder, A. et al. Loss-of-function of the voltage-gated sodium channel NaV1.5 (channelopathies) in patients with irritable bowel syndrome. Gastroenterology 146, 1659–1668 (2014).
Strege, P. R. et al. Irritable bowel syndrome patients have SCN5A channelopathies that lead to decreased NaV1.5 current and mechanosensitivity. Am. J. Physiol. Gastrointest. Liver Physiol. 314, G494–G503 (2018).
Sanders, K. M., Ward, S. M. & Koh, S. D. Interstitial cells: regulators of smooth muscle function. Physiol. Rev. 94, 859–907 (2014).
Henström, M. et al. TRPM8 polymorphisms associated with increased risk of IBS-C and IBS-M. Gut 66, 1725–1727 (2017).
Khanna, R., MacDonald, J. K. & Levesque, B. G. Peppermint oil for the treatment of irritable bowel syndrome: a systematic review and meta-analysis. J. Clin. Gastroenterol. 48, 505–512 (2014).
Gericke, B., Amiri, M. & Naim, H. Y. The multiple roles of sucrase-isomaltase in the intestinal physiology. Mol. Cell. Pediatr. 3, 2 (2016).
Gericke, B., Amiri, M., Scott, C. R. & Naim, H. Y. Molecular pathogenicity of novel sucrase-isomaltase mutations found in congenital sucrase-isomaltase deficiency patients. Biochim. Biophys. Acta Mol. Basis Dis. 1863, 817–826 (2017).
Alfalah, M., Keiser, M., Leeb, T., Zimmer, K.-P. & Naim, H. Y. Compound heterozygous mutations affect protein folding and function in patients with congenital sucrase-isomaltase deficiency. Gastroenterology 136, 883–892 (2009).
Foley, A. et al. Adult sucrase-isomaltase deficiency masquerading as IBS. Gut 71, 1237–1238 (2021).
Muldoon, C., Maguire, P. & Gleeson, F. Onset of sucrase-isomaltase deficiency in late adulthood. Am. J. Gastroenterol. 94, 2298–2299 (1999).
Ringrose, R. E., Preiser, H. & Welsh, J. D. Sucrase-isomaltase (palatinase) deficiency diagnosed during adulthood. Dig. Dis. Sci. 25, 384–387 (1980).
Henström, M. et al. Functional variants in the sucrase-isomaltase gene associate with increased risk of irritable bowel syndrome. Gut 67, 263–270 (2018).
Thingholm, L. et al. Sucrase-isomaltase 15Phe IBS risk variant in relation to dietary carbohydrates and faecal microbiota composition. Gut 68, 177–178 (2019).
Garcia-Etxebarria, K. et al. Increased prevalence of rare sucrase-isomaltase pathogenic variants in irritable bowel syndrome patients. Clin. Gastroenterol. Hepatol. 16, 1673–1676 (2018).
Chumpitazi, B. P. et al. Hypomorphic SI genetic variants are associated with childhood chronic loose stools. PLoS ONE 15, e0231891 (2020).
Zheng, T. et al. Rare hypomorphic sucrase isomaltase variants in relation to irritable bowel syndrome risk in UK biobank. Gastroenterology 161, 1712–1714 (2021).
Zheng, T. et al. Reduced efficacy of low FODMAPs diet in patients with IBS-D carrying sucrase-isomaltase (SI) hypomorphic variants. Gut 69, 397–398 (2020).
Nilholm, C., Roth, B. & Ohlsson, B. A dietary intervention with reduction of starch and sucrose leads to reduced gastrointestinal and extra-intestinal symptoms in IBS patients. Nutrients 11, E1662 (2019).
Klein, R. J. et al. Complement factor H polymorphism in age-related macular degeneration. Science 308, 385–389 (2005).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
de Lange, K. M. et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017).
Dubois, P. C. A. et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 42, 295–302 (2010).
Trynka, G. et al. Dense genotyping identifies and localizes multiple common and rare variant association signals in celiac disease. Nat. Genet. 43, 1193–1201 (2011).
Coleman, C. et al. Common polygenic variation in coeliac disease and confirmation of ZNF335 and NIFA as disease susceptibility loci. Eur. J. Hum. Genet. 24, 291–297 (2016).
Smyth, D. J. et al. Shared and distinct genetic variants in type 1 diabetes and celiac disease. N. Engl. J. Med. 359, 2767–2777 (2008).
Jefremow, A. & Neurath, M. F. All are equal, some are more equal: targeting IL 12 and 23 in IBD - a clinical perspective. Immunotargets Ther. 9, 289–297 (2020).
Seyed Tabib, N. S. et al. Big data in IBD: big progress for clinical practice. Gut 69, 1520–1532 (2020).
Gettler, K. et al. Common and rare variant prediction and penetrance of IBD in a large, multi-ethnic, health system-based biobank cohort. Gastroenterology 160, 1546–1557 (2021).
Uffelmann, E. et al. Genome-wide association studies. Nat. Rev. Methods Prim. 1, 59 (2021).
Wijmenga, C. & Zhernakova, A. The importance of cohort studies in the post-GWAS era. Nat. Genet. 50, 322–328 (2018).
Bycroft, C. et al. The UK biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Eijsbouts, C. et al. Genome-wide analysis of 53,400 people with irritable bowel syndrome highlights shared genetic pathways with mood and anxiety disorders. Nat. Genet. 53, 1543–1552 (2021).
Ek, W. E. et al. Exploring the genetics of irritable bowel syndrome: a GWA study in the general population and replication in multinational case-control cohorts. Gut 64, 1774–1782 (2015).
Holliday, E. G. et al. Genome-wide association study identifies two novel genomic regions in irritable bowel syndrome. Am. J. Gastroenterol. 109, 770–772 (2014).
Bonfiglio, F. et al. A GWAS meta-analysis from 5 population-based cohorts implicates ion channel genes in the pathogenesis of irritable bowel syndrome. Neurogastroenterol. Motil. 30, e13358 (2018).
Bonfiglio, F. et al. Female-specific association between variants on chromosome 9 and self-reported diagnosis of irritable bowel syndrome. Gastroenterology 155, 168–179 (2018).
Day, F. R. et al. Genomic analyses identify hundreds of variants associated with age at menarche and support a role for puberty timing in cancer risk. Nat. Genet. 49, 834–841 (2017).
Elks, C. E. et al. Thirty new loci for age at menarche identified by a meta-analysis of genome-wide association studies. Nat. Genet. 42, 1077–1085 (2010).
Heitkemper, M., Jarrett, M., Bond, E. F. & Chang, L. Impact of sex and gender on irritable bowel syndrome. Biol. Res. Nurs. 5, 56–65 (2003).
Kim, Y. S. & Kim, N. Sex-gender differences in irritable bowel syndrome. J. Neurogastroenterol. Motil. 24, 544–558 (2018).
Whitehead, W. E. et al. Evidence for exacerbation of irritable bowel syndrome during menses. Gastroenterology 98, 1485–1489 (1990).
Norcliffe-Kaufmann, L., Slaugenhaupt, S. A. & Kaufmann, H. Familial dysautonomia: history, genotype, phenotype and translational research. Prog. Neurobiol. 152, 131–148 (2017).
Camilleri, M. Gastrointestinal motility disorders in neurologic disease. J. Clin. Invest. 131, 143771 (2021).
Tillisch, K. et al. Sex specific alterations in autonomic function among patients with irritable bowel syndrome. Gut 54, 1396–1401 (2005).
Viramontes, B. E. et al. Gender-related differences in slowing colonic transit by a 5-HT3 antagonist in subjects with diarrhea-predominant irritable bowel syndrome. Am. J. Gastroenterol. 96, 2671–2676 (2001).
Hogan, A. M., Collins, D., Baird, A. W. & Winter, D. C. Estrogen and its role in gastrointestinal health and disease. Int. J. Colorectal Dis. 24, 1367–1375 (2009).
Taché, Y. & Bonaz, B. Corticotropin-releasing factor receptors and stress-related alterations of gut motor function. J. Clin. Invest. 117, 33–40 (2007).
Bernstein, M. T. et al. Gastrointestinal symptoms before and during menses in healthy women. BMC Women’s Health 14, 14 (2014).
Meleine, M. & Matricon, J. Gender-related differences in irritable bowel syndrome: potential mechanisms of sex hormones. World J. Gastroenterol. 20, 6725–6743 (2014).
Houghton, L. A., Lea, R., Jackson, N. & Whorwell, P. J. The menstrual cycle affects rectal sensitivity in patients with irritable bowel syndrome but not healthy volunteers. Gut 50, 471–474 (2002).
Wu, Y. et al. GWAS of peptic ulcer disease implicates Helicobacter pylori infection, other gastrointestinal disorders and depression. Nat. Commun. 12, 1146 (2021).
Boutwell, B. et al. Replication and characterization of CADM2 and MSRA genes on human behavior. Heliyon 3, e00349 (2017).
Pasman, J. A. et al. GWAS of lifetime cannabis use reveals new risk loci, genetic overlap with psychiatric traits, and a causal influence of schizophrenia. Nat. Neurosci. 21, 1161–1170 (2018).
Frei, J. A., Andermatt, I., Gesemann, M. & Stoeckli, E. T. The SynCAM synaptic cell adhesion molecules are involved in sensory axon pathfinding by regulating axon-axon contacts. J. Cell. Sci. 127, 5288–5302 (2014).
Ibrahim-Verbaas, C. A. et al. GWAS for executive function and processing speed suggests involvement of the CADM2 gene. Mol. Psychiatry 21, 189–197 (2016).
Petrovska, J. et al. The NCAM1 gene set is linked to depressive symptoms and their brain structural correlates in healthy individuals. J. Psychiatr. Res. 91, 116–123 (2017).
Schmid, R. S. & Maness, P. F. L1 and NCAM adhesion molecules as signaling coreceptors in neuronal migration and process outgrowth. Curr. Opin. Neurobiol. 18, 245–250 (2008).
Pappa, S. et al. PHF2 histone demethylase prevents DNA damage and genome instability by controlling cell cycle progression of neural progenitors. Proc. Natl Acad. Sci. USA 116, 19464–19473 (2019).
Shi, L. Dock protein family in brain development and neurological disease. Commun. Integr. Biol. 6, e26839 (2013).
Detera-Wadleigh, S. D. et al. Sequence variation in DOCK9 and heterogeneity in bipolar disorder. Psychiatr. Genet. 17, 274–286 (2007).
Kuramoto, K., Negishi, M. & Katoh, H. Regulation of dendrite growth by the Cdc42 activator Zizimin1/Dock9 in hippocampal neurons. J. Neurosci. Res. 87, 1794–1805 (2009).
Nagel, M., Watanabe, K., Stringer, S., Posthuma, D. & van der Sluis, S. Item-level analyses reveal genetic heterogeneity in neuroticism. Nat. Commun. 9, 905 (2018).
Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
Whitehead, W. E., Palsson, O. & Jones, K. R. Systematic review of the comorbidity of irritable bowel syndrome with other disorders: what are the causes and implications? Gastroenterology 122, 1140–1156 (2002).
Gottesman, I. I. & Gould, T. D. The endophenotype concept in psychiatry: etymology and strategic intentions. Am. J. Psychiatry 160, 636–645 (2003).
Visscher, P. M., Brown, M. A., McCarthy, M. I. & Yang, J. Five years of GWAS discovery. Am. J. Hum. Genet. 90, 7–24 (2012).
Camilleri, M. & Katzka, D. A. Irritable bowel syndrome: methods, mechanisms, and pathophysiology. Genetic epidemiology and pharmacogenetics in irritable bowel syndrome. Am. J. Physiol. Gastrointest. Liver Physiol. 302, G1075–G1084 (2012).
Camilleri, M. et al. Neuropeptide S receptor induces neuropeptide expression and associates with intermediate phenotypes of functional gastrointestinal disorders. Gastroenterology 138, 98–107.e4 (2010).
Wong, B. S. et al. A Klothoβ variant mediates protein stability and associates with colon transit in irritable bowel syndrome with diarrhea. Gastroenterology 140, 1934–1942 (2011).
D’Amato, M. et al. Neuropeptide s receptor 1 gene polymorphism is associated with susceptibility to inflammatory bowel disease. Gastroenterology 133, 808–817 (2007).
Camilleri, M. & Nurko, S. Bile acid diarrhea in adults and adolescents. Neurogastroenterol. Motil. 34, e14287 (2021).
Henström, M. et al. NPSR1 polymorphisms influence recurrent abdominal pain in children: a population-based study. Neurogastroenterol. Motil. 26, 1417–1425 (2014).
Degen, L. P. & Phillips, S. F. How well does stool form reflect colonic transit? Gut 39, 109–113 (1996).
Zinsmeister, A. R., Burton, D. & Camilleri, M. Pharmacodynamic and clinical endpoints for functional colonic disorders: statistical considerations. Dig. Dis. Sci. 58, 509–518 (2013).
Jankipersadsing, S. A. et al. A GWAS meta-analysis suggests roles for xenobiotic metabolism and ion channel activity in the biology of stool frequency. Gut 66, 756–758 (2017).
Bonfiglio, F. et al. GWAS of stool frequency provides insights into gastrointestinal motility and irritable bowel syndrome. Cell Genom. https://doi.org/10.1016/j.xgen.2021.100069 (2021).
Smillie, C. S. et al. Intra- and inter-cellular rewiring of the human colon during ulcerative colitis. Cell 178, 714–730.e22 (2019).
Drokhlyansky, E. et al. The human and mouse enteric nervous system at single-cell resolution. Cell 182, 1606–1622.e23 (2020).
Browning, K. N. & Travagli, R. A. Central nervous system control of gastrointestinal motility and secretion and modulation of gastrointestinal functions. Compr. Physiol. 4, 1339–1368 (2014).
Modarresi, F. et al. Inhibition of natural antisense transcripts in vivo results in gene-specific transcriptional upregulation. Nat. Biotechnol. 30, 453–459 (2012).
Hing, B. et al. A polymorphism associated with depressive disorders differentially regulates brain derived neurotrophic factor promoter IV activity. Biol. Psychiatry 71, 618–626 (2012).
Maisonpierre, P. C. et al. NT-3, BDNF, and NGF in the developing rat nervous system: parallel as well as reciprocal patterns of expression. Neuron 5, 501–509 (1990).
Di Carlo, P., Punzi, G. & Ursini, G. Brain-derived neurotrophic factor and schizophrenia. Psychiatr. Genet. 29, 200–210 (2019).
Liu, S. Neurotrophic factors in enteric physiology and pathophysiology. Neurogastroenterol. Motil. 30, e13446 (2018).
Grider, J. R., Piland, B. E., Gulick, M. A. & Qiao, L. Y. Brain-derived neurotrophic factor augments peristalsis by augmenting 5-HT and calcitonin gene-related peptide release. Gastroenterology 130, 771–780 (2006).
Chai, N.-L. et al. Effects of neurotrophins on gastrointestinal myoelectric activities of rats. World J. Gastroenterol. 9, 1874–1877 (2003).
Chen, F. et al. Brain-derived neurotrophic factor accelerates gut motility in slow-transit constipation. Acta Physiol. 212, 226–238 (2014).
Coulie, B. et al. Recombinant human neurotrophic factors accelerate colonic transit and relieve constipation in humans. Gastroenterology 119, 41–50 (2000).
Taché, Y., Garrick, T. & Raybould, H. Central nervous system action of peptides to influence gastrointestinal motor function. Gastroenterology 98, 517–528 (1990).
Taché, Y., Kolve, E., Maeda-Hagiwara, M. & Kauffman, G. L. Central nervous system action of calcitonin to alter experimental gastric ulcers in rats. Gastroenterology 94, 145–150 (1988).
Fukudo, S., Nomura, T. & Hongo, M. Impact of corticotropin-releasing hormone on gastrointestinal motility and adrenocorticotropic hormone in normal controls and patients with irritable bowel syndrome. Gut 42, 845–849 (1998).
Stengel, A. & Taché, Y. Neuroendocrine control of the gut during stress: corticotropin-releasing factor signaling pathways in the spotlight. Annu. Rev. Physiol. 71, 219–239 (2009).
Kaji, I. et al. Free fatty acid receptor 3 activation suppresses neurogenic motility in rat proximal colon. Neurogastroenterol. Motil. 30, 13157 (2018).
Nøhr, M. K. et al. GPR41/FFAR3 and GPR43/FFAR2 as cosensors for short-chain fatty acids in enteroendocrine cells vs FFAR3 in enteric neurons and FFAR2 in enteric leukocytes. Endocrinology 154, 3552–3564 (2013).
Soret, R. et al. Short-chain fatty acids regulate the enteric neurons and control gastrointestinal motility in rats. Gastroenterology 138, 1772–1782 (2010).
Sinha, T. et al. Analysis of 1135 gut metagenomes identifies sex-specific resistome profiles. Gut Microbes 10, 358–366 (2019).
Shastri, P., McCarville, J., Kalmokoff, M., Brooks, S. P. J. & Green-Johnson, J. M. Sex differences in gut fermentation and immune parameters in rats fed an oligofructose-supplemented diet. Biol. Sex. Differ. 6, 13 (2015).
Pittayanon, R. et al. Gut microbiota in patients with irritable bowel syndrome–a systematic review. Gastroenterology 157, 97–108 (2019).
Agnello, M. et al. Gut microbiome composition and risk factors in a large cross-sectional IBS cohort. BMJ Open Gastroenterol. 7, e000345 (2020).
Vich Vila, A. et al. Gut microbiota composition and functional changes in inflammatory bowel disease and irritable bowel syndrome. Sci. Transl Med. 10, eaap8914 (2018).
Botschuijver, S. et al. Intestinal fungal dysbiosis is associated with visceral hypersensitivity in patients with irritable bowel syndrome and rats. Gastroenterology 153, 1026–1039 (2017).
Mihindukulasuriya, K. A. et al. Multi-omics analyses show disease, diet, and transcriptome interactions with the virome. Gastroenterology 161, 1194–1207.e8 (2021).
Hadizadeh, F. et al. Stool frequency is associated with gut microbiota composition. Gut 66, 559–560 (2017).
Tigchelaar, E. F. et al. Gut microbiota composition associated with stool consistency. Gut 65, 540–542 (2016).
Zhernakova, A. et al. Population-based metagenomics analysis reveals markers for gut microbiome composition and diversity. Science 352, 565–569 (2016).
Sanna, S. et al. Causal relationships among the gut microbiome, short-chain fatty acids and metabolic diseases. Nat. Genet. 51, 600–605 (2019).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Gacesa, R. et al. Environmental factors shaping the gut microbiome in a Dutch population. Nature 604, 732–739 (2022).
Sanna, S., Kurilshikov, A., van der Graaf, A., Fu, J. & Zhernakova, A. Challenges and future directions for studying effects of host genetics on the gut microbiome. Nat. Genet. 54, 100–106 (2022).
Kurilshikov, A. et al. Large-scale association analyses identify host factors influencing human gut microbiome composition. Nat. Genet. 53, 156–165 (2021).
Hughes, D. A. et al. Genome-wide associations of human gut microbiome variation and implications for causal inference analyses. Nat. Microbiol. 5, 1079–1087 (2020).
Rühlemann, M. C. et al. Genome-wide association study in 8,956 German individuals identifies influence of ABO histo-blood groups on gut microbiome. Nat. Genet. 53, 147–155 (2021).
Lopera-Maya, E. A. et al. Effect of host genetics on the gut microbiome in 7,738 participants of the Dutch Microbiome Project. Nat. Genet. 54, 143–151 (2022).
Qin, Y. et al. Combined effects of host genetics and diet on human gut microbiota and incident disease in a single population cohort. Nat. Genet. 54, 134–142 (2022).
Goodrich, J. K. et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 19, 731–743 (2016).
Turpin, W. et al. Association of host genome with intestinal microbial composition in a large healthy cohort. Nat. Genet. 48, 1413–1417 (2016).
Watanabe, K. et al. A global overview of pleiotropy and genetic architecture in complex traits. Nat. Genet. 51, 1339–1348 (2019).
Olds, L. C. & Sibley, E. Lactase persistence DNA variant enhances lactase promoter activity in vitro: functional role as a cis regulatory element. Hum. Mol. Genet. 12, 2333–2340 (2003).
Morales, E. et al. The European lactase persistence genotype determines the lactase persistence state and correlates with gastrointestinal symptoms in the Hispanic and Amerindian Chilean population: a case–control and population-based study. BMJ Open 1, e000125 (2011).
Brandao Gois, M. F. et al. Role of the gut microbiome in mediating lactose intolerance symptoms. Gut 71, 215–217 (2022).
Poole, A. C. et al. Human salivary amylase gene copy number impacts oral and gut microbiomes. Cell Host Microbe 25, 553–564.e7 (2019).
Zhou, W. et al. Global Biobank Meta-analysis Initiative: powering genetic discovery across human diseases. Preprint at medRxiv https://doi.org/10.1101/2021.11.19.21266436 (2021).
Camilleri, M. & Chedid, V. Actionable biomarkers: the key to resolving disorders of gastrointestinal function. Gut 69, 1730–1737 (2020).
Scholtens, S. et al. Cohort Profile: LifeLines, a three-generation cohort study and biobank. Int. J. Epidemiol. 44, 1172–1180 (2015).
Polygenic Risk Score Task Force of the International Common Disease Alliance. Responsible use of polygenic risk scores in the clinic: potential benefits, risks and gaps. Nat. Med. 27, 1876–1884 (2021).
Sirugo, G., Williams, S. M. & Tishkoff, S. A. The missing diversity in human genetic studies. Cell 177, 26–31 (2019).
DeBoever, C. et al. Assessing digital phenotyping to enhance genetic studies of human diseases. Am. J. Hum. Genet. 106, 611–622 (2020).
Biomarkers Definitions Working Group. Biomarkers and surrogate endpoints: preferred definitions and conceptual framework. Clin. Pharmacol. Ther. 69, 89–95 (2001).
Kawahara, H., Minami, R. & Yokota, N. BAG6/BAT3: emerging roles in quality control for nascent polypeptides. J. Biochem. 153, 147–160 (2013).
Binici, J. & Koch, J. BAG-6, a jack of all trades in health and disease. Cell. Mol. Life Sci. 71, 1829–1837 (2014).
Cheishvili, D. et al. IKAP/Elp1 involvement in cytoskeleton regulation and implication for familial dysautonomia. Hum. Mol. Genet. 20, 1585–1594 (2011).
Jackson, M. Z., Gruner, K. A., Qin, C. & Tourtellotte, W. G. A neuron autonomous role for the familial dysautonomia gene ELP1 in sympathetic and sensory target tissue innervation. Development 141, 2452–2461 (2014).
Malumbres, M. et al. Cyclin-dependent kinases: a family portrait. Nat. Cell Biol. 11, 1275–1276 (2009).
Cole, A. R. PCTK proteins: the forgotten brain kinases? Neurosignals 17, 288–297 (2009).
Vieira, N., Rito, T., Correia-Neves, M. & Sousa, N. Sorting out sorting nexins functions in the nervous system in health and disease. Mol. Neurobiol. 58, 4070–4106 (2021).
Jin, Q. et al. Novel function of FAXDC2 in megakaryopoiesis. Blood Cancer J. 6, e478 (2016).
Rooman, I. et al. Expression of the Notch signaling pathway and effect on exocrine cell proliferation in adult rat pancreas. Am. J. Pathol. 169, 1206–1214 (2006).
Kadur Lakshminarasimha Murthy, P. et al. Radical and lunatic fringes modulate notch ligands to support mammalian intestinal homeostasis. eLife 7, e35710 (2018).
Katoh, M. & Katoh, M. Notch signaling in gastrointestinal tract (review). Int. J. Oncol. 30, 247–251 (2007).
Ratanasirintrawoot, S. & Israsena, N. Stem cells in the intestine: possible roles in pathogenesis of irritable bowel syndrome. J. Neurogastroenterol. Motil. 22, 367–382 (2016).
W, C. et al. SCF-FBXO24 regulates cell proliferation by mediating ubiquitination and degradation of PRMT6. Biochem. Biophys. Res. Commun. 530, 75–81 (2020).
Moore, S. W. & Johnson, G. Acetylcholinesterase in Hirschsprung’s disease. Pediatr. Surg. Int. 21, 255–263 (2005).
Soreq, H. & Seidman, S. Acetylcholinesterase–new roles for an old actor. Nat. Rev. Neurosci. 2, 294–302 (2001).
Demir, I. E., Schäfer, K.-H., Tieftrunk, E., Friess, H. & Ceyhan, G. O. Neural plasticity in the gastrointestinal tract: chronic inflammation, neurotrophic signals, and hypersensitivity. Acta Neuropathol. 125, 491–509 (2013).
Kano, M. et al. Altered brain and gut responses to corticotropin-releasing hormone (CRH) in patients with irritable bowel syndrome. Sci. Rep. 7, 12425 (2017).
Sagami, Y. et al. Effect of a corticotropin releasing hormone receptor antagonist on colonic sensory and motor function in patients with irritable bowel syndrome. Gut 53, 958–964 (2004).
Taché, Y. & Million, M. Role of corticotropin-releasing factor signaling in stress-related alterations of colonic motility and hyperalgesia. J. Neurogastroenterol. Motil. 21, 8–24 (2015).
Samuel, B. S. et al. Effects of the gut microbiota on host adiposity are modulated by the short-chain fatty-acid binding G protein-coupled receptor, Gpr41. Proc. Natl Acad. Sci. USA 105, 16767–16772 (2008).
Tazoe, H. et al. Expression of short-chain fatty acid receptor GPR41 in the human colon. Biomed. Res. 30, 149–156 (2009).
Gee, H. Y., Tang, B. L., Kim, K. H. & Lee, M. G. Syntaxin 16 binds to cystic fibrosis transmembrane conductance regulator and regulates its membrane trafficking in epithelial cells. J. Biol. Chem. 285, 35519–35527 (2010).
Tang, B. L. Syntaxin 16’s newly deciphered roles in autophagy. Cells 8, E1655 (2019).
Longstreth, G. F. et al. Functional bowel disorders. Gastroenterology 130, 1480–1491 (2006).
Acknowledgements
The research time of M.C. for IBS studies is funded by NIH RO1 DK115950. A.Z. is supported by ERC Starting Grant 715772, Netherlands Organization for Scientific Research NWO-VIDI grant 016.178.056, Netherlands Heart Foundation CVON grant 2018-27, and NWO Gravitation grant ExposomeNL 024.004.017. The research of M.D’A. in IBS is supported by the Spanish Ministry of Science and Innovation (MICINN, PID2020-113625RB) and the Spanish Agency for Investigation (AEI, PCI2021-122064-2A) under the umbrella of the European Joint Programming Initiative “A Healthy Diet for a Healthy Life” (JPI HDHL) and of the ERA-NET Cofund ERA-HDHL (GA no. 696295 of the EU Horizon 2020 Research and Innovation Programme).
Author information
Authors and Affiliations
Contributions
All authors researched data for the article. All authors contributed substantially to discussion of the content. M.D’A., M.C. and A.Z. wrote the article. All authors reviewed and/or edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
M.C. has received funding for single-centre pharmacodynamic studies from VANDA Pharmaceuticals and NGM Pharmaceuticals. M.D’A. has received financial support from QOL Medical, in the form of unrestricted research grants. A.Z. and I.B. declare no competing interests.
Peer review
Peer review information
Nature Reviews Gastroenterology & Hepatology thanks Hans Tornblom and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
GWAS Catalog: https://www.ebi.ac.uk/gwas/
International Classification of Diseases, Ninth Revision (ICD9): https://www.cdc.gov/nchs/icd/icd9.htm
International Classification of Diseases, Tenth Revision (ICD10): https://www.cdc.gov/nchs/icd/icd10.htm
National Institute for Health and Care Excellence: https://www.nice.org.uk/guidance/cg61
Rome criteria: https://theromefoundation.org/rome-iv/rome-iv-criteria/
Glossary
- Dysbiosis
-
Disruption of the intestinal microbiota homeostasis, determining a reduction in microbial diversity, alterations in the composition and distribution of the autochthon commensals, and changes in metabolic activities.
- Genetic architecture
-
Broadly, the genetic landscape that underlies a phenotypic trait and its variation, characterized by the number of genetic variants that contribute to that phenotype, their effect size and the patterns of interactions. It summarizes the ‘genetics’ behind a trait.
- Biobanks
-
Collections of extensive data from biomedical research linked to study participants, including biospecimens, electronic health-related records, genetic data, imaging, biomarkers and information regarding environment and lifestyle.
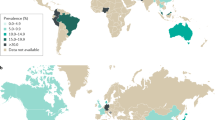
- Prevalence
-
The proportion of individuals in a population who have a specific characteristic (typically a disease) at a specific time.
- GWAS Catalog
-
A free online curated collection of human genome-wide association study (GWAS) results representing a collaboration between the European Bioinformatics Institute of the European Molecular Biology Laboratory and the National Human Genome Research Institute; it summarizes significant single-nucleotide polymorphism–trait associations, and sample metadata from each publication (as of 9 July 2022, the catalogue contains 5,848 publications and 398,342 associations).
- Polygenic scores
-
(PGS). Values summarizing the estimated (cumulative) effects of multiple DNA variants in determining the probability of an individual manifesting specific phenotypes or traits. The score is useful for evaluating an individual’s genetic predisposition for a particular trait and can also be used as a predictor relative to the general population (increased or decreased risk of disease, expressed as a polygenic risk score for that disease).
- Biomarkers
-
Characteristics that can be objectively measured and evaluated as indicators of normal biological processes, pathogenic processes or pharmacological responses to a therapeutic intervention (adapted from the definition of the Biomarkers Definitions Working Group).
- Omics
-
A set of scientific disciplines (such as genomics, metagenomics, metabolomics, proteomics and transcriptomics) that exploit large-scale data to characterize and quantify molecular interactions at a global level, whether biochemical, molecular or cellular data, or data from organs or other systems.
- Genome-wide significance level
-
The consensus threshold defining the statistical significance of a reported genome-wide association between a single-nucleotide polymorphism and a given trait. Currently defined as P < 5 × 10−8 (originally based on a Bonferroni correction).
- Gene set enrichment analysis
-
A computational method that, based on existing knowledge, allows the identification of biological pathways, functions and/or disease-related categories that are significantly (P < 0.05) enriched in a specific set of genes compared with a reference panel (most often the whole transcriptome).
- Suggestive association
-
A threshold that, although not reaching genome-wide significance, could be used as evidence of association warranting further investigation and validation in independent datasets (often set at P ≤ 5 × 10−6).
- Minor allele
-
The less common allele for a specific single-nucleotide polymorphism in a given population.
- Locus
-
The specific location of a DNA sequence, gene or genetic marker on a chromosome.
- Genetic correlations
-
Parameters that quantify the proportion of variance that two traits share owing to genetic causes, estimating the degree of pleiotropy (the phenomenon by which a single gene or locus influences the phenotypic appearance of multiple traits, including different diseases) or causal overlap.
- Mendelian randomization
-
An analytical method that uses random segregation of genetic variants as an instrument to emulate a randomized controlled trial, to investigate the putative causal effect of an exposure (using genetic variations strongly associated with it) on an outcome of interest, allowing confounders and reverse causation to be reduced; it can be used to test whether genetic predisposition to a disease (outcome) is mediated via DNA variants (instruments) predisposing to another disease (exposure), and vice versa.
- Primary and secondary ICD10 diagnoses
-
According to the ICD-10-CM Official Guidelines for Coding and Reporting, FY 2021, the principal diagnosis is defined as the condition established to be chiefly responsible for occasioning the admission of the patient to the hospital, whereas other diagnoses are all conditions that coexist at the time of admission, develop subsequently, or affect the treatment received and/or the length of stay.
- Linkage disequilibrium
-
In a given population, the non-random segregation of alleles from different loci, resulting in their combinations (haplotypes) occurring more or less often than expected based merely on their individual allele frequencies (random, independent segregation).
- Lead SNP
-
The marker that gives rise to the strongest association signal at a given genome-wide association study locus.
- SNP heritability
-
The proportion of phenotypic variance of a given trait, which is causally explained by a specific set of single-nucleotide polymorphisms (SNPs).
- Functional annotation
-
The process of collecting and assigning functional information to genomic regions, such as genome-wide association study loci (including gene content, regulatory elements, expression, molecular function, subcellular location, interactions, etc).
- Major allele
-
The more common allele for a specific single-nucleotide polymorphism in a given population.
- Promoter
-
A regulatory element; a sequence of DNA that allows the binding of RNA polymerase and transcription factors responsible for the transcription of the downstream gene, therefore fundamentally contributing to its expression.
- Alternative splicing
-
The post-transcriptional process resulting in different combinations of exons from the same precursor mRNA producing alternative mRNA transcripts; this allows a single gene to code for multiple proteins, often associated with developmental and tissue-specific expression.
- Post-translational modifications
-
All the chemical modifications that take place after the translation of a polypeptide chain (such as the addition of functional groups and/or polypeptides, proteolytic cleavage, glycosylation and phosphorylation), expanding the diversity in structures and functions of proteins.
- Exposome
-
The totality of environmental factors an individual is exposed to over a lifetime, usually considered in relation to health measures.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Camilleri, M., Zhernakova, A., Bozzarelli, I. et al. Genetics of irritable bowel syndrome: shifting gear via biobank-scale studies. Nat Rev Gastroenterol Hepatol 19, 689–702 (2022). https://doi.org/10.1038/s41575-022-00662-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41575-022-00662-2
This article is cited by
-
Shared genetic architecture between irritable bowel syndrome and psychiatric disorders reveals molecular pathways of the gut-brain axis
Genome Medicine (2023)
-
The neurobiology of irritable bowel syndrome
Molecular Psychiatry (2023)