Abstract
Studying the microbial symbionts of eukaryotic hosts has revealed a range of interactions that benefit host biology. Most eukaryotes are also infected by parasites that adversely affect host biology for their own benefit. However, it is largely unclear whether the ability of parasites to develop in hosts also depends on host-associated symbionts, e.g., the gut microbiota. Here, we studied the parasitic wasp Leptopilina boulardi (Lb) and its host Drosophila melanogaster. Results showed that Lb successfully develops in conventional hosts (CN) with a gut microbiota but fails to develop in axenic hosts (AX) without a gut microbiota. We determined that developing Lb larvae consume fat body cells that store lipids. We also determined that much larger amounts of lipid accumulate in fat body cells of parasitized CN hosts than parasitized AX hosts. CN hosts parasitized by Lb exhibited large increases in the abundance of the bacterium Acetobacter pomorum in the gut, but did not affect the abundance of Lactobacillus fructivorans which is another common member of the host gut microbiota. However, AX hosts inoculated with A. pomorum and/or L. fructivorans did not rescue development of Lb. In contrast, AX larvae inoculated with A. pomorum plus other identified gut community members including a Bacillus sp. substantially rescued Lb development. Rescue was further associated with increased lipid accumulation in host fat body cells. Insulin-like peptides increased in brain neurosecretory cells of parasitized CN larvae. Lipid accumulation in the fat body of CN hosts was further associated with reduced Bmm lipase activity mediated by insulin/insulin-like growth factor signaling (IIS). Altogether, our results identify a previously unknown role for the gut microbiota in defining host permissiveness for a parasite. Our findings also identify a new paradigm for parasite manipulation of host metabolism that depends on insulin signaling and the gut microbiota.
Similar content being viewed by others
Introduction
Infectious organisms, including protozoan parasites and pathogenic bacteria, have long been recognized as major human health challenges [1, 2]. It is now appreciated that intimate interactions between hosts and other species are extremely widespread, and sometimes benefit hosts as exemplified by commensal gut microbes [3,4,5]. During the last two decades, the study of the gut microbiota has become one of the most intriguing areas of biological research. Reasons include that gut microbes are widely reported to play crucial roles in host immunity, metabolism and development [6,7,8]. Recently, the gut microbiota has also been found to provide a protective barrier to suppress parasitic infections through either direct competition or indirectly stimulating host immune pathways [9,10,11,12]. As such, understanding the interactions between the host microbiota and parasites could provide significant insights into interactions with hosts and potential therapeutic approaches.
Parasites are well known to use hosts as their primary source of nutrition, often exchanging their ability to synthesize nutrients de novo for efficient salvaging mechanisms [13,14,15]. Nutrients act not only as primary sources of energy but also as regulators of gene expression, metabolism and growth, through various signaling networks that enable parasites to survive [1, 16]. Evolved nutritional strategies can also modify host metabolism to suit specific nutritional requirements of the parasite [17,18,19,20]. Similar to some protozoan parasites, the offspring of parasitic wasps (also known as parasitoids) acquire their nutrients directly from the host throughout their development [21, 22]. Parasitoids also often manipulate host metabolism to meet dietary needs and promote growth and survival [23, 24]. Although alteration of host energy homeostasis by parasites is a widespread phenomenon, underlying mechanisms remain largely unknown.
Lipid stores crucially affect both host and parasite physiology [20, 25, 26]. Neutral lipids or triacylglycerols (TAGs) are primarily stored in cytosolic lipid droplets (LDs) within adipose tissue in mammals or the fat body in insects [27, 28]. Research over the past decade has established LDs as dynamic organelles and as central, active players in lipid homeostasis [29, 30]. The mechanisms underlying LD size and content include regulation of the balance between lipogenesis and lipolysis and are broadly conserved from insects to mammals [31,32,33]. In Drosophila, lipid homeostasis is substantially influenced by a group of neurosecretory cells in the brain that produce insulin-like peptides (Dilps) [34, 35]. These peptides activate the Drosophila insulin receptor to alter the mobilization of TAG stores in a tissue-specific manner. Upregulation of Drosophila insulin signaling specifically in the fat body increases TAG stores [35]. Conversely, a reduction in insulin signaling promotes nuclear translocation and transcriptional activity of Forkhead Box O (FOXO), which in turn leads to lipolysis by upregulation of Brummer (Bmm) lipase, the fly homolog of adipose triglyceride lipase (ATGL) in mammals [33].
Leptopilina boulardi (Lb) is a larval-pupal parasitoid. Adult females lay a single egg per oviposition inside a Drosophila melanogaster 2nd instar larva. After hatching, the Lb larva develops through five instars before pupating and emerging as an adult 19–21 days later [36, 37]. Lb provides an excellent research system for studying host-parasite interactions generally, and is also directly relevant to agricultural pest control since many parasitoids are used as biological control agents. In this study, we report that Lb develop into adults in conventional (CN) D. melanogaster that have a gut microbiota but do not develop in axenic (AX) hosts with no gut microbiota. Parasitism stimulated increased lipid accumulation in the fat body of CN but not AX hosts through increased insulin signaling. The mixed community of microbes in CN hosts substantially rescued Lb development when transferred to AX hosts. Screening assays indicated that transfer of two gut community members in the genus Acetobacter and Bacillus to AX D. melanogaster resulted in gnotobiotic (GN) hosts that also substantially rescued Lb development.
Materials and methods
Drosophila and wasp strains
Drosophila stocks were maintained on standard cornmeal/molasses/agar medium at room temperature. Leptopilina boulardi wasps were bred on a D. melanogaster W1118 stock, and adult wasps were maintained at room temperature in vials with apple juice agar [36]. The following D. melanogaster stains were used: UAS-InR RNAi (THU5741), UAS-AkhR RNAi (TH02253.N) and UAS-Pten RNAi (THU0582) lines were obtained from the TsingHua Fly Center (TH). W1118 (BL5905), Tub-GFP-PH (BL8164) and Dilp2-Gal4 (BL37516) lines were obtained from the Bloomington Drosophila Stock Center (BL). Ppl-Gal4 and UAS-EGFP lines were provided by S. Li (South China Normal University, Guangzhou, China). UAS-Bmm RNAi, UAS-Lipin RNAi, UAS-DGAT RNAi, UAS-Lsd1RNAi, and UAS-Hsl RNAi lines were provided by X. Huang (Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing, China) [30, 38]. Each of the RNAi lines used in the study were validated by comparing relative expression levels of the target gene in fat cells to a positive control by quantitative reverse transcription PCR as described below (Supplementary Fig. 1).
Generation of axenic host animals (AX)
Axenic larvae (AX) were generated as previously described with minor modifications [39]. D. melanogaster embryos were incubated for 2 min in 2.5% sodium hypochlorite, and were subsequently washed twice in 70% ethanol, followed by three washes with sterile water. The embryos were then transferred onto sterile food and routinely checked for bacterial contamination by culturing the homogenates on LB and MRS agar plates. To compensate for the developmental delay observed in axenic larvae, the yeast content was increased to 41.7 g per liter in axenic food comparing to the regular fly food recipe. Similar to the previous report [40], there was no difference in size and development time between AX and conventional (CN) Drosophila hosts when they were fed on the yeast-rich fly food used in this study.
Lb parasitism and developmental stages of offspring
Mated 3-day-old Lb female wasps were allowed to parasitize CN and AX 2nd instar D. melanogaster W1118 at a wasp/host ratio of 1:10 for 3 h, respectively. Ten CN or AX host larvae were randomly selected and dissected under a microscope and the portion of parasitized hosts was calculated as the percentage that contained a wasp egg(s) relative to the total number of hosts that were dissected. Eight replicates were performed for the each group. The remaining parasitized hosts were collected, and the different developing stages of hosts, their fat bodies and wasp offspring were photographed using an Olympus SZX16 microscope. The developmental stages of wasps in CN or AX host larvae (Supplementary Table 1) were identified using previously described morphological characters [41].
LD staining and size measurement
At least 20 D. melanogaster larvae were dissected in PBS to collect fat body and Lb larvae that were fixed in 4% paraformaldehyde in PBS for 30 min. After rinsing in PBS, samples were incubated for 30 min in a 1:1000 dilution 1 mg/ml BODIPY 493/503 (Invitrogen) in PBS, and then rinsed twice with PBS containing 0.05% Tween 20, 0.1% Triton (PBST). Stained samples were mounted in ProLong Gold Antifade Mountant with DAPI (Invitrogen). Fluorescence images were captured on a Zeiss LSM 800 confocal microscope. LD size was measured as described [30]. Briefly, the diameters of at least 150 LDs (the five largest LDs within one cell) were measured by Zeiss Zen Software version 2.3.
Lipid measurement
TAG measurements were performed as described [42]. Fat bodies of at least six D. melanogaster larvae were collected and homogenized in PBST. Samples were centrifuged at 12,000 rpm for 10 min and the supernatants were heated at 70 °C for 15 min. Lipid levels were measured by using TAG Determination Kit (Sigma), and were normalized to protein amounts in each homogenate that were determined by Bradford assay (Invitrogen).
Feeding assays
Synchronized 3rd instar Drosophila larvae (parasitized and nonparasitized) were cultured on fresh agar medium followed by fasting for 2 h. Thirty larvae of each group were allowed to feed on yeast diet for 20 min that contained 0.01 g bromophenol blue (FD&C Blue NO.1, TCL)/ml of water. Larvae containing varying amounts of blue color in their abdomens were categorized by visual inspection as fully-fed, partially-fed or non-fed.
Immunohistochemistry
Fat bodies or brains from D. melanogaster 3rd instars were dissected in PBS and fixed in 4% paraformaldehyde in PBS for 30 min. After rinsing in PBST, tissues were blocked with 5% bovine serum albumin (BSA) in PBST, and incubated overnight at 4 °C with rabbit anti-FOXO (1:500; Cosmo Bio, Japan), rabbit anti-Dilp2 (1:500; provided by C. Tong, Zhejiang University, China), rabbit anti-NPF (1:200; provided by Z. Zhao, China Agriculture University, China), or rabbit anti-sNPF (1:200, generated against the peptides DPSLPQMRRTAYDDLLEREL by ABclonal, China). Fat bodies or brains were then washed three times in PBST and incubated with Alexa Fluor 488 or Alexa Fluor 594 secondary antibodies (1:1000; Molecular Probes) for 2 h at room temperature. Samples were washed twice in PBST, once in PBS and mounted in ProLong Gold Antifade Mountant with DAPI (Invitrogen). Fluorescence images were captured on a Zeiss LSM 800 confocal microscope, and were processed using ImageJ (National Institutes of Health) and Photoshop (Adobe).
Lipase activity assay
Fat bodies of 20 D. melanogaster 3rd instars were dissected and homogenized in 100 μl lipase assay buffer. Lipase activity was measured by using Lipase Activity Kit (Sigma) according to manufacturer protocol, and was normalized to protein amounts in each homogenate by using the BCA Protein Assay Kit (Invitrogen). Experiments were repeated at least three times.
Reverse Transcription Quantitative PCR (RT-qPCR)
Total RNA was isolated from wasp larvae as well as host brains, fat bodies, guts and the remainder of the body (carcass) using the RNeasy Mini Kit (Qiagen) according to the manufacturer’s protocol. RT-qPCR assays were performed using the AriaMx real-time PCR system (Agilent Technologies) with the ChamQ SYBR qPCR Master Mix Kit (Vazyme, Nanjing, China). Primers for amplifying 100–200 bp of each PCR product are listed in Supplementary Table 2. Reactions were carried out for 30 min at 50 °C, followed by 10 min at 95 °C, then followed by 40 cycles of two-step PCR for 15 s at 95 °C, 1 min at 60 °C. The RNA levels of target genes were normalized to tubulin RNA and relative concentration was determined using the 2−ΔΔCt Method.
ROS measurement in vivo
Intracellular ROS levels were determined by measuring the oxidative conversion of cell-permeable 2′,7′-dichlorofluorescein diacetate (DCFH-DA) to fluorescent dichlorofluorescein (DCF) through Reactive Oxygen Species Assay Kit (Beyotime Biotechnology, Shanghai, China). Guts of different developmental host larvae were homogenized in 100 μl NP40 buffer, and centrifuged at 12,000 rpm for 10 min. Then, 10 μl of supernatants were incubated with 100 µmol/L DCFH-DA at 37 °C for 1 h in the dark. Fluorescence was read at 488 nm for excitation and 528 nm for emission with the SpectraMax iD5 Multi-Mode Microplate Reader (Molecular Devices). ROS levels were normalized to the total protein of each sample, which was determined by BCA Protein Assay Kit (Invitrogen). At least three repetitions were performed for each group.
Immunoblotting
Total protein from fat bodies of D. melanogaster 3rd instar larvae was isolated using protein extraction buffer (Solarbio, Beijing, China). After centrifugation at 13,000 rpm for 15 min, supernatants were subjected to SDS-PAGE and transferred to PVDF (Millipore). Membranes were incubated in blocking solution (Tris-buffered saline containing 0.1% Tween 20, 5% BSA) for 1 h. Anti-phospho-AKT (Ser505, Cell Signaling Technology, Danvers, MA), anti-AKT (Cell Signaling Technology, Danvers, MA), and anti-β-tubulin (Beyotime Biotechnology, Shanghai, China) primary antibodies and horseradish-peroxidase-conjugated anti-rabbit IgG and anti-mouse IgG secondary antibody (Solarbio, Beijing, China) were used at a dilution of 1:1000. Resulting immunoblots were visualized using a Bio-Rad Universal Hood II Gel Doc System, while densitometric analysis was performed using Bio-Rad’s Image Lab software.
Drosophila gut microbiota analysis
Host guts were surface sterilized with ethanol and dissected in sterile PBS on an ice plate under a stereoscope (Nikon). A total of 150 guts per experiment and replicate were collected from nonparasitized and parasitized 3rd instar hosts. Bacterial DNA was extracted using the QIAmp DNA Mini Kit (Qiagen). A fragment encompassing the V4 and V5 region of the 16S rRNA gene was amplified using the primers 515F and 926R (Supplementary Table 2). Resulting amplicons were then sequenced on the Illumina MiSeq platform (Shanghai Tiny Gene Bio-Tech Company). FASTA files were filtered to a minimum read length of 200 bp and were clustered into operational taxonomic units (OTUs) at 97% sequence identity using Mothur software (http://www.mothur.org). OTU taxonomies were determined based on the SILVA database (v132, http://www.arb-silva.de) and the National Center for Biotechnology Information 16S rRNA bacterial database as of September 10, 2019 with standard blastn algorithm settings. Gut microbiota experiments were conducted with different D. melanogaster host generations: the original generation of both nonparasitized and parasitized larvae (experiment#1) and a second generation (40 generations later) (experiment#2) that included two replicates.
Isolation of Acetobacter pomorum, Lactobacillus fructivorans and the other bacteria
Five D. melanogaster guts from 3rd instar larvae were homogenized in 200 μl of PBS, and spread onto MRS, LB, and 869 agar plates. 869 agar is a reduced nutrient medium comprised of 0.04 g CaCl2–2H2O, 0.1 g glucose, D+, 0.5 g NaCl, 1 g tryptone, 0.5 g yeast extract, and 15 g agar in 1000 ml of diluted water. All plates were incubated for 72 h at 30 °C. A. pomorum, L. fructivorans and the other bacteria were identified by sequencing. Briefly, DNA was isolated from colonies on MRS plates with typical characteristics of Acetobacter and Lactobacillus bacteria [43] and used as template to PCR amplify products generated using universal 16 S rRNA 27 F and 1522 R primers (Supplementary Table 2). The same DNA isolation and PCR procedures were used for randomly selected colonies on LB agar plates and 869 agar plates. Purified PCR products were Sanger sequenced using an ABI Prism 3730 sequencer (Applied Biosystems). Resulting sequences were then referenced to the GenBank database by BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi).
Quantification of A. pomorum
A series of tenfold dilutions of A. pomorum were used to construct a standard curve. Briefly, CT values in each dilution were measured using qPCR with A. pomorum-specific primers. The CT values were then plotted against the logarithm of their colony-forming units. Next, the genomic DNA from ten nonparasitized or parasitized Drosophila larval guts were extracted using the DNeasy Kit (Qiagen) according to the manufacture protocol. Absolute colony numbers per larval gut were determined from the above standard curves, using the resulting CT values. qPCR analysis was performed with the AriaMx real-time PCR system (Agilent Technologies) using ChamQ SYBR qPCR Master Mix Kit (Vazyme) and specific primers (Supplementary Table 2).
Inoculation of AX larvae
Fifteen host guts from 3rd instar nonparasitized CN Drosophila larvae were dissected and homogenized in 200 μl PBS. Approximately 50 μl of this gut mixture was added to each axenic food vial containing germ-free embryos. To generate GN D. melanogaster, AX larvae were fed A. pomorum alone, L. fructivorans alone, A. pomorum plus L. fructivorans, or different combinations of 6 other bacteria identified as gut community members (M. sp., R. sp., C. sp., B. sp., P. sp., and Stenotrophomonas maltophilia) (approximately 1 × 108 CFUs for each bacteria). Ten guts for each group were dissected from larvae after inoculation and compared to an AX control by isolating genomic DNA from dissected guts. A. pomorum and L. fructivorans were then detected by PCR using specific primers (Supplementary Table. 2). Five μl of each PCR reaction was then subjected to agarose gel electrophoresis and visualized. Amplification of the D. melanogaster tubulin gene using specific primers (Supplementary Table. 2) served as an endogenous control.
Statistical analyses
Statistical analyses were performed in GraphPad Prism version 9.0 (GraphPad Software) and SPSS version 19.0 (IBM).
Results
Conventional D. melanogaster larvae with a gut microbiota support the development of Lb into adults but axenic larvae do not
Under nutrient-limiting conditions, the gut microbiota promotes the growth of D. melanogaster larvae, whereas AX and CN larvae usually exhibit no growth differences when fed nutrient-rich diets [40]. We similarly detected no differences between AX and CN larvae from our laboratory culture in growth to the adult stage when fed a nutrient-rich yeast-based diet (See methods). In contrast, adult Lb emerged from 85% of parasitized CN hosts while no adults emerged from parasitized AX hosts (Fig. 1A). Dissection of host larvae indicated that Lb females parasitized the same proportion of CN and AX hosts as measured by the presence of a wasp egg after oviposition (Supplementary Fig. 2). Lb eggs similarly hatched and developed to the 3rd instar in CN and AX host larvae, which coincided with when hosts ceased feeding as late third instars and initiated metamorphosis to the pupal stage (Supplementary Fig. 3). The fat body in D. melanogaster dissociates into individual adipocytes after pupation that are refractive to autophagic cell death, which eliminates most other larval cell types [44]. We observed that the fat body in CN and AX hosts similarly dissociated into individual adipocytes if parasitized by Lb (Supplementary Fig. 3). We also noted that Lb larvae rapidly consumed these dissociated adipocytes, which were visualized by BODIPY which stains neutral lipids in LDs (Supplementary Fig. 3). However, while most Lb larvae in CN host pupae molted to the 5th instar, pupated and emerged as adult wasps, nearly all Lb larvae in AX host pupae remained 4th instars that ultimately died (Fig. 1A, Supplementary Figs. 3 and 4). These observations overall indicated that Lb larvae in both CN and AX hosts consumed adipocytes, but while the former successfully developed into adult wasps the latter did not.
A The percentage of CN and AX hosts that produce adult Lb. Bars show the mean percentage of hosts ± SD that produced adult Lb in three replicates of 200 hosts for each treatment. Significance was determined by two-sided Mann–Whitney U test (***p < 0.001). B Fluorescent images of fat body cells from late 3rd instar (3LL) CN or AX larvae that were nonparasitized or parasitized. LDs in fat body cells were labeled by BODIPY (green) while nuclei were labeled by DAPI (blue). Scale bars: 20 μm. C LD diameters in (B). Each data point is a single LD. A minimum of 200 LDs were measured in fat body cells from 30 individual larvae for each treatment. Error bars show the mean ± SD for each treatment. Significance was determined by two-sided unpaired Student’s t test (ns non-significant; ***p < 0.001). D Relative lipid levels in fat bodies from 3LL CN and AX larvae that were nonparasitized or parasitized. Fat bodies from 20 individual larvae were measured per replicate. Bars show mean lipid level ± SD from three independent samples. Significance was determined by two-sided unpaired Student’s t test (ns: non-significant; *p < 0.05).
Given the preceding results, we BODIPY stained adipocytes in the fat body of Drosophila hosts from the 2nd until the end of the 3rd instar when feeding ceased (Fig. 1B, Supplementary Fig. 5A). This showed that LD diameters increased more in parasitized than nonparasitized CN hosts, which correlated with higher TAG levels (Fig. 1C, D, Supplementary Fig. 5A–C). In contrast, no differences were detected in LD diameters or TAG levels between parasitized and nonparasitized AX hosts, which indicated increased TAG levels after parasitism depended on the presence of a gut microbiota (Fig. 1B–D). We initially thought parasitized CN larvae might have larger lipid stores because they fed more, but bioassays using dye-marked food indicated they overall fed less than nonparasitized CN larvae (Supplementary Fig. 6A, B). We also immunocytochemically visualized neuropeptide F (NPF) and short neuropeptide F (sNPF) in the brains of 3rd instars since both have functions in regulating feeding by Drosophila larvae [45, 46]. We detected less NPF in the brains of parasitized than nonparasitized CN larvae (Supplementary Fig. 6C, D), which suggested reduced feeding could be related to downregulation of this neuropeptide. In contrast, we detected no differences in sNPF between parasitized and nonparasitized larvae (Supplementary Fig. 6E, F).
Lb infection changes the composition of the gut microbiota in host larvae
Recent studies show that lipid metabolic homeostasis in many organisms relies on a healthy commensal community [47, 48]. We thus generated 16S rRNA sequencing libraries from two generations of CN larvae (experiment#1 and experiment#2) to characterize the bacteria present in the guts of D. melanogaster third instars (nonparasitized and parasitized) from our laboratory culture (See “Methods”). For the first generation (experiment#1), more than 99% of resulting reads after quality filtering were bacteria in four Phyla (Proteobacteria, Firmicutes, Bacteroidetes, and Actinobacteria) (Supplementary Fig. 7A, Supplementary Table 3). Reads corresponding to Proteobacteria were much lower in the second generation (experiment#2), whereas the abundance of Firmicutes, Fusobacteria, and Epsilonbacteraeota were higher (Supplementary Fig. 7A, Supplementary Table 4). Overall, 58 genera were shared across both generations and replicates, which accounted for 38.80%, 40.75%, and 44.76% of the total reads from experiment#1, experiment#2 replicate 1, and experiment#2 replicate 2 (Supplementary Fig. 7B, Supplementary Tables 3 and 4). At the generic level, only Lactobacillus and Acetobacter were among the top nine most abundant taxa in each generation and replicate, which was consistent with detection of these genera in many D. melanogaster laboratory cultures [49, 50]. High variation in gut microbiota composition has also been reported in other studies where larvae were maintained in the same laboratory and fed the same diet [51, 52]. Our results thus supported a growing consensus that gut microbiota composition is variable due to colonization by a number of opportunistic bacteria, while only a few genera are commonly shared [49, 53]. Gut homogenates plated on MRS agar from experiment#1 yielded a large number of colonies with characteristics of Acetobacter and Lactobacillus spp. 16 S ribosomal gene sequencing of 30 Lactobacillus colonies indicated each was a >99% nucleotide match to L. fructivorans (Supplementary Fig. 8), while sequencing of 50 Acetobacter colonies indicated each was a > 99% nucleotide match to Acetobacter pomorum (Supplementary Fig. 9). Comparing the gut microbiota present in parasitized and nonparasitized CN larvae at the level of genus indicated the most abundant taxa in each treatment was qualitatively similar for experiment #1 and experiment #2 (Supplementary Fig. 7A–C, Supplementary Tables 3–6) although some quantitative differences were also noted including a more than 20-fold increase in A. pomorum in parasitized larvae from experiment #1, and a more than 10-fold increase across the two replicates in experiment #2 (Supplementary Fig. 7C). qPCR assays indicated that A. pomorum also progressively increased in abundance from the early to late third instar in both nonparasitized and parasitized larvae (Supplementary Fig. 7D).
A major modulator of gut immunity and microbial populations in Drosophila is the NADPH oxidase Duox, which can generate reactive oxygen species (ROS) to defend against pathogenic bacteria [39, 54]. To test whether changes in Duox expression might underlie the microbiome changes induced by parasitoid infection, we used RT-qPCR to measure relative Duox RNA. Expression of Duox was significantly reduced in the guts of parasitized Drosophila larvae, while Duox expression in carcasses lacking the guts remained unaltered (Supplementary Fig. 7E). We also measured ROS levels using 2′,7′-dichlorofluorescein diacetate (DCFH-DA) and found a significant reduction in the guts in response to parasitization (Supplementary Fig. 10). Bacteria in the guts can also be controlled by antimicrobial peptides (AMPs) [55]. We tested the expression profiles of six AMP genes, chosen based on their roles in Drosophila gut immunity. Interestingly, three AMP genes encoding Diptericin (Dpt), Attacin (Att), and Metch were significantly up-regulated in the guts after parasitization (Supplementary Fig. 7F), and all six AMP genes were up-regulated in carcasses lacking the guts (Supplementary Fig. 7G). Thus, parasitism strongly affected the expression of Duox and AMPs, two major classes of bactericidal agents in the guts, and substantially altered the host gut microbiota population including a large increase in the proportion of A. pomorum.
GN larvae inoculated with certain gut community members support Lb development
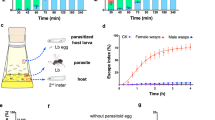
To investigate whether particular members of the host gut microbiota affect development of Lb, we made gut homogenates from nonparasitized CN hosts. We referred to this homogenate as a “gut mixture” that we used to inoculate AX larvae (Fig. 2A). We then allowed Lb females to parasitize these hosts, which resulted in 43.5% yielding adult wasps (Fig. 2B). This outcome indicated a mixed community of microbes from the guts of CN larvae significantly increased wasp survival when compared to AX hosts from which no adult wasps emerge. However, a successful parasitism rate of 43.5% was also lower than ~85% of CN hosts producing Lb adults when the gut microbiota was naturally acquired from the environment (see Fig. 1A). Since A. pomorum and L. fructivorans were consistently abundant gut community members, we next assessed whether AX larvae inoculated with either or both of these species produced GN larvae that supported Lb development. Both successfully colonized AX larvae (Supplementary Fig. 11) but only 7.5% of GN larvae inoculated with A. pomorum and 8.0% of GN larvae inoculated with A. pomorum and L. fructivorans produced adult wasps while no GN larvae inoculated with only L. fructivorans did so (Fig. 2C). We thus isolated other bacteria present in the guts of CN larvae by plating gut homogenates on LB and 869 agar plates. Randomly picking and sequencing 95 colonies from LB plates indicated 40 were a Mesorhizobium sp. (Supplementary Fig. 12), 25 were a Rhodococcus sp. (Supplementary Fig. 13), 15 were a Chitinophaga sp. (Supplementary Fig. 14), 10 were a Bacillus sp. (Supplementary Fig. 15), and 5 were a Pseudomonas sp. (Supplementary Fig. 16). Doing the same for 152 colonies from 869 agar plates indicated 92 were A. pomorum, 26 were the Mesorhizobium sp., 20 were S. maltophilia (Supplementary Fig. 17), and the remaining 14 were the Pseudomonas sp. Inoculating AX larvae with a mixture of S. maltophilia, Mesorhizobium sp., Rhodococcus sp., Chitinophaga sp., Bacillus sp. and Pseudomonas sp. resulted in only 3.1% of hosts producing adult Lb (Fig. 2C). However, inoculating larvae with these six species plus A. pomorum or A. pomorum and L. fructivorans resulted in 38-40% of hosts producing Lb which was comparable to inoculating larvae with our gut mixture from CN larvae (Fig. 2C). Inoculating AX larvae with A. pomorum plus Bacillus sp. and Rhodococcus sp. resulted in 29.2% of GN larvae producing adult Lb, while AX larvae inoculated with A. pomorum plus Bacillus sp. resulted in 20.2% of GN larvae producing adult Lb (Fig. 2C). Comparing LD sizes among treatments further indicated that AX larvae inoculated with A. pomorum plus Bacillus sp. or A. pomorum plus Bacillus sp. and Rhodococcus sp. were larger than AX larvae inoculated with A. pomorum alone (Fig. 2D, E). Thus, A. pomorum together with the Bacillus sp. alone or in combination with the Rhodococcus sp. significantly increased the proportion of GN larvae that produced Lb adults which was associated with increased LD size in fat body.
A Schematic showing how AX hosts were inoculated with bacteria to produce GN hosts that were parasitized. The percentage of hosts for each treatment producing adult wasps (eclosion) were then determined. B Results showing the percentage of AX and GN hosts that produced adult wasps from the experimental design shown in (A). Treatments were parasitized AX hosts and GN hosts inoculated with gut mixture from CN larvae. From left to right: n = 672, 368. Bars show the mean percentage of hosts ± SD that produced adults in at least six replicates per treatment. Significance was determined by two-sided Mann–Whitney U test (***p < 0.001). C Results showing the percentage of AX and GN hosts that produced adult wasps from the experimental design shown in (A). Treatments were parasitized AX hosts and GN hosts inoculated with A. pomorum only, L. fructivorans only, A. pomorum plus L. fructivorans, A. pomorum plus combinations of six other bacteria identified as gut community members. From left to right: n = 672, 641, 581, 655, 415, 782, 496, 634, 547, 383, 483, 781, 616, 422. A. po Acetobacter pomorum, L. fru Lactobacillus fructivorans, P. sp. Pseudomonas sp., M. sp. Mesorhizobium sp., S. ma Stenotrophomonas maltophilia, B. sp. Bacillus sp., R. sp. Rhodococcus sp., C. sp. Chitinophaga sp. Bars show the mean percentage of hosts ± SD that produced adult in at least six replicates per treatment. Significance was determined by Kruskal–Wallis test with Dunn’s multiple comparisons test. (ns non-significant; ***p < 0.001). D BODIPY and DAPI labeled images of fat body cells from 3LL GN D. melanogaster larvae inoculated with A. pomorum only, A. pomorum and Bacillus. sp., or A. pomorum, Bacillus. sp. and Rhodococcus sp. Scale bars: 20 μm. E Quantification of LD diameters in (D). Each data point is a single LD. A minimum of 150 LDs were measured in fat body cells from 20 individual larvae for each treatment. Error bars show the mean ± SD for each treatment. Significance was determined by Kruskal–Wallis test with Dunn’s multiple comparisons test. (ns: non-significant; ***p < 0.001).
Lipid stores in the fat body of parasitized CN hosts are affected by reduction of Bmm lipase activity
Four enzymatic steps regulate the conversion of glycerol-3-phosphate to TAG by esterification with fatty acids [32]. In Drosophila, Lipin converts phosphatidic acid into diacylglycerol (DAG) in the third step, while diacylglycerol acyltransferase (DGAT) converts DAG to TAG in the fourth step (Supplementary Fig. 18) [56, 57]. In turn, lipolysis involves two lipases, Bmm and hormone-sensitive Hsl, plus lipid storage droplet protein 1 (Lsd1) that recruits Hsl but not Bmm to the surface of LDs (Supplementary Fig. 18) [58]. To investigate the direct effectors of parasitization-induced lipid accumulation, we used the ppl-gal4 driver to knock down lipogenesis and lipolysis genes in the fat body. We found that fat bodies with reduced expression of the lipogenesis genes Lipin or DGAT had smaller LDs and neutral lipids stores in nonparasitized CN larvae (Fig. 3A, B, D), confirming an important role of Lipin and DGAT in TAG synthesis in vivo. Nevertheless, parasitism by Lb still increased LD size and TAG levels in the hosts with reduced Lipin or DGAT activity in CN larvae (Fig. 3A, B, D). Thus, Lb greatly alters LD size under conditions of severely reduced lipogenic capacity, suggesting that parasitism primarily alters host lipolysis rather than lipogenesis. As expected, knockdown of Bmm or Hsl or Lsd1 increased lipid stores in nonparasitized CN larvae (Fig. 3A, C, E). LD size and TAG level were further increased upon parasitization in Hsl-deficient CN larvae, Lsd1-deficient CN larvae but not in Bmm-deficient CN larvae, suggesting that parasitism increases lipid stores in host fat body principally by reducing Bmm lipase activity (Fig. 3A, C, E). Consistent with this hypothesis and a key role for regulating lipolysis, lipase activity in the fat body was also overall lower in parasitized versus nonparasitized CN larvae which correlated with lower transcript levels for Bmm but not Hsl or Lsd1 (Fig. 3F, G). Transcription of Bmm depends on FOXO activity in the nucleus, which functions as a regulatory intermediate of PI3K-AKT signaling through the IIS pathway [33, 59]. As such, increased insulin signaling in parasitized CN larvae is thought to reduce Bmm gene expression through phosphorylation and associated inhibition of FOXO entry into the nucleus [32, 60], which was consistent with our results showing that nuclear FOXO was lower in parasitized than nonparasitized CN larvae (Fig. 3H). Collectively then, the preceding results indicated parasitism enhanced lipid accumulation by down-regulating FOXO nuclear activity and which in turn reduced Bmm lipase activity in the fat body of CN hosts.
A Fluorescent images of fat bodies from nonparasitized and parasitized hosts. Genotypes are as follows: ppl/+ (ppl-Gal4/+), ppl > DGAT RNAi (ppl-Gal4/+; UAS-DGAT RNAi/+), ppl > Lipin RNAi (ppl-Gal4/+; UAS-Lipin RNAi/+), ppl > Bmm RNAi (ppl-Gal4/+; UAS-bmm RNAi/+), ppl > Hsl RNAi (ppl-Gal4/+; UAS-Hsl RNAi/+) and ppl > Lsd1 RNAi (ppl-Gal4/+; UAS-lsd1 RNAi/+). LDs were labeled by BODIPY (green) and nuclei were labeled by DAPI (blue). Scale bars: 20 μm. B, C LD diameters in (A). Each data point is a single LD. A minimum of 200 LDs were measured in fat body cells from 30 individual larvae for each treatment. Error bars show the mean ± SD for each treatment. Significance was determined by two-sided unpaired Student’s t test (ns: non-significant; ***p < 0.001). D Relative lipid levels in fat bodies from nonparasitized and parasitized larvae ppl > DGAT or ppl > Lipin RNAi hosts. Fat bodies from 20 larvae were measured per replicate. Bars show mean lipid level ± SD from three independent samples. Significance was determined by two-sided unpaired Student’s t test (*p < 0.05; **p < 0.01; ***p < 0.001). E Relative lipid levels in fat bodies from nonparasitized and parasitized ppl > Bmm, ppl > Hsl, or ppl > Lsd1 RNAi hosts. Fat bodies from 20 larvae were measured per replicate. Bars show mean lipid level ± SD from three independent samples. Significance was determined by two-sided unpaired Student’s t test (ns non-significant; *p < 0.05; **p < 0.01; ***p < 0.001). F Relative lipase activity in fat bodies from 3rd instar nonparasitized and parasitized hosts (n = 20 pooled larvae per sample). Bars for each treatment show the mean ± SD from three independent samples. Significance was determined by two-sided unpaired Student’s t test (*p < 0.05). G Relative mRNA levels of Bmm, Hsl, and Lsd1 in fat bodies from 3rd instar nonparasitized and parasitized hosts (n = 10 pooled larvae per sample). Bars for each treatment show the mean ± SD from three independent experiments. Significance was determined by two-sided unpaired Student’s t test (ns: non-significant; **p < 0.001). H Immunocytochemical localization of FOXO by an anti-FOXO antibody (red) in larval fat body cells. FOXO is detected in the cytoplasm and nuclei of cells from nonparasitized hosts, whereas FOXO localization was reduced in nuclei from parasitized hosts (arrowheads). Nuclei were labeled by DAPI (blue). Scale bar: 50 μm.
Since knockdown of DGAT reduced lipogenesis as measured by smaller lipid stores, we asked if Lb development was adversely affected in parasitized hosts where DGAT was knocked down. Results showed that Lb usually developed to the adult stage in DGAT knockdown hosts but males showed a 1-day delay in emergence while females showed a 2-day delay (Supplementary Fig. 19A, B). Adult female wasps were also smaller than female wasps from control hosts (Supplementary Fig. 19C). Since parasitism depends on adult females injecting both eggs and venom secretions into hosts, we compared ovary and venom gland sizes between females that emerged from DGAT knockdown and control hosts. Venom gland but not ovary size was significantly smaller in Lb females from DGAT knockdown hosts (Supplementary Fig. 19D-G). The reduction in venom gland size led us to hypothesize that females from DGAT knockdown hosts could also suffer a fitness penalty in parasitizing new hosts. Indeed, rates of successful parasitism were lower for females that emerged from DGAT knockdown than control hosts (Supplementary Fig. 19H). Taken together, knockdown of DGAT resulted in lipid stores that were lower than in control parasitized hosts but were higher than in parasitized AX hosts. Correspondingly, Lb fitness was reduced in parasitized CN hosts where DGAT was knocked down when compared to control CN hosts, but was also much higher than in AX hosts where Lb failed to develop into adults.
Elevated insulin signaling promotes lipid accumulation in parasitized hosts with a gut microbiota
Since Drosophila adipokinetic hormone (AKH) and insulin-like peptides (Dilps) are key regulators of lipid homeostasis [61], we used the ppl-Gal4 driver to knock down the AKH (UAS-AkhR RNAi) and insulin-like receptor (UAS-InR RNAi) in the fat body [38]. Depletion of the AkhR in nonparasitized CN third instars increased LD diameter when compared to nonparasitized control larvae (Fig. 4A, B). This outcome was consistent with AkhR null mutants promoting obesity [61] and overexpression of AKH or AkhR reducing fat stores [35]. However, AkhR depletion in parasitized CN larvae increased LD diameters more than in nonparasitized CN larvae, which suggested the larger lipid stores in parasitized hosts did not require the AkhR. In contrast, InR depletion reduced LD diameters in both nonparasitized and parasitized CN larvae (Fig. 4A, B), while activation of the insulin pathway via reduction of Pten (phosphatase and tensin homolog) increased LD diameters in nonparasitized CN larvae to similar levels as parasitized CN larvae (Supplementary Fig. 20A, B). Tub-PH-GFP encodes the pleckstrin homology (PH) domain of the Drosophila Steppke (Grp1) protein which is fused to GFP under control of the tubulin gene promoter. This results in global expression at a modest level and recruitment of the GFP-tagged PH domain to the surface of cells via binding to phosphatidylinositol 3, 4, 5-trisphosphate (PIP3), which is the second messenger generated by phosphoinositide 3-kinase (PI3K). The GFP signal thus provides a measure of insulin pathway signaling via PI3 kinase activity [62]. PH-GFP levels in fat body adipocytes were higher in parasitized than nonparasitized CN larvae (third instars) (Fig. 4C, D). Increased PIP3 levels further result in activation of protein kinase AKT, a central target of the InR-PI3K pathway, via phosphorylation at residue Ser505 [63]. Immunoblotting assays showed that Ser505 phosphorylation in fat body extracts was also higher in parasitized than nonparasitized CN larvae (Fig. 4E, F).
A Fat body cells from nonparasitized and parasitized CN hosts with LDs labeled with BODIPY (green) and nuclei labeled with DAPI (blue). Genotypes are: ppl/+ (ppl-Gal4/+), ppl > AkhR RNAi (ppl-Gal4/+; UAS-AkhR RNAi/+), and ppl > InR RNAi (ppl-Gal4/+; UAS-InR RNAi/+). Scale bars: 20 μm. B LD diameters in (A). Each data point is a single LD. A minimum of 200 LDs were measured in fat body cells from 30 individual larvae for each treatment. Error bars show the mean ± SD for each treatment. Significance was determined by two-sided unpaired Student’s t test (ns: non-significant; ***p < 0.001). C PI3K activity in nonparasitized and parasitized fat body cells as marked by PH-GFP (green). Scale bars: 50 μm. D Fluorescence intensity of PH-GFP on the fat body membrane of nonparasitized and parasitized hosts. At least 100 fat body cells from 30 individual larvae were measured per treatment. Bars show mean fluorescence intensity per cell ± SD. Significance was determined by two-sided unpaired Student’s t test (**p < 0.01). E, F Immunoblot analysis of phosphorylated (pS505) AKT, total AKT, and β-tubulin (loading control) in the fat body of nonparasitized and parasitized host larvae. p-AKT and AKT levels were normalized to the loading control in comparing relative protein levels. Bars show mean ± SD from three immunoblots prepared from independent samples. Significance was determined by two-sided unpaired Student’s t test (ns: non-significant; *p < 0.05).
The preceding results indicated that parasitism of CN hosts increases insulin signaling above what normally occurs in nonparasitized CN hosts. Three (Dilp2, Dilp3 and Dilp5) of the eight Dilp genes are selectively expressed in ilp-producing medial neurosecretory cells (IPCs) and are known regulators of metabolic homeostasis in the fat body [64]. Dilp2 was more strongly detected in IPCs from parasitized than nonparasitized CN larvae by anti-Dilp2 (Fig. 5A, B), as was a Dilp2-Gal4 transgene that drove UAS-GFP expression (Supplementary Fig. 21). RT-qPCR assays also detected higher expression of Dilp2, Dilp3 and Dilp5 in the brains of parasitized than nonparasitized CN larvae (Fig. 5C). In contrast, no differences were detected in the expression of Dilp 2, 3 and 5 between parasitized and nonparasitized AX larvae, which strongly suggested elevated expression required the presence of a gut microbiota (Fig. 5C).
A Immunohistochemical analysis of Dilp2 in IPCs from nonparasitized and parasitized hosts. IPCs were labeled with anti-Dilp2 (green) while nuclei were labeled with DAPI (blue). Scale bars: 100 μm. B Fluorescence intensity of Dilp2 in IPCs from nonparasitized and parasitized hosts (n = 20 for each group). Bars show mean fluorescence per cell ± SD. Significance was determined by two-sided unpaired Student’s t test (***p < 0.001). C Relative mRNA levels of Dilp2, Dilp3, and Dilp5 in brains from CN and AX 3rd instar hosts that were nonparasitized or parasitized (n = 20 for each group). Bars show mean ± SD from three independent experiments. Significance was determined by two-sided unpaired Student’s t test (ns: non-significant; **p < 0.01).
Discussion
Host nutrients are a key currency that parasites often manipulate [1, 5, 18], while the gut microbiota of many host organisms influences metabolism or susceptibility to infection [2, 47]. Here, we present results where these inter-species interactions intersect: a parasite that increases host lipid stores for nutritional needs and survival but also requires the host gut microbiota to do so (Fig. 6).
Lb successfully develops into adults in conventional hosts (CN) but fails to develop in axenic hosts (AX). In addition, increased abundance of A. pomorum plus other identified gut community members including a Bacillus sp. are necessary for wasp survival, which is associated with increased insulin signaling in brain neurosecretory cells of parasitized CN larvae and increased lipid accumulation in host fat body cells.
Lipid profiles are probably the best-characterized parasitism-associated metabolic alteration [20, 25, 26]. Host lipid profiles are also known to affect the metabolism, survival and/or transmission of some parasites [1, 16]. For example, Toxoplasma gondii secures cellular fatty acids essential for its proliferation [20]. While parasitoid wasps are unable to de novo synthesize lipids [15] and thus rely on host lipid metabolism to meet dietary needs for development into adults [21, 23, 65]. In this study, we identify a developmental strategy whereby Lb allows the host to grow to the pupal stage, but its development into adults also depends on increased lipid accumulation in fat body cells of hosts that wasp larvae consume. Our results indicate increased lipid accumulation involves increased expression of Dilps in IPCs and insulin signaling in the host fat body. How Lb stimulates this increase in insulin signaling remains unclear but our results do reveal the novel finding that manipulation of host metabolism by the wasp depends on the presence of a gut microbiota in the host. Key evidence supporting this conclusion is our finding that AX larvae inoculated with gut microbes from CN larvae substantially increases the proportion of parasitized hosts that produce adult Lb. While A. pomorum has previously been reported to positively modulate Drosophila energy metabolism by producing modulators like acetic acid that promote insulin signaling [47], inoculating AX larvae with this gut community member alone or L. frutivorans does not substantially increase the proportion of parasitized GN larvae that produce adult Lb. In contrast, inoculating AX larvae with A. pomorum plus six other gut community members increases the proportion of Lb that develop into adults to similar levels as our positive control (gut mixture). A. pomorum plus a Bacillus sp. present in CN larvae also substantially increased the proportion of GN larvae that produce adult Lb. We thus conclude that successful development of Lb depends on particular members of the gut microbiota present in CN larvae. A key goal going forward is to identify how the gut microbiota promotes insulin signaling and nutrient storage in the fat body even though parasitized host larvae feed less than nonparasitized larvae. Another is to determine how gut community members interact to promote Lb fitness given that A. pomorum plus six other gut community members or A. pomorum plus Bacillus sp. in GN larvae substantially increase the proportion of Lb larvae that develop into adults but do not restore survival to levels in CN larvae that acquire a gut microbiota by feeding. Our results overall suggest LD quantity in host adipocytes does affect Lb development or adult fitness. However, our finding that hosts where DGAT was knocked down produced adult wasps with reduced fitness while AX hosts produced no adult wasps also suggests the host gut microbiota potentially affects the specific types of lipids that are stored in the fat body which in turn influence Lb development. Thus, profiling lipid composition in parasitized versus nonparasitized hosts and how lipid composition is potentially affected by specific gut community members is another important goal for future investigation.
In summary, our results demonstrate reliance of a parasite on the gut microbiota of a host for nutritional needs and survival. Our results also provide new insights into the mechanisms underlying how Lb manipulates the metabolism of hosts with a gut microbiota. Parasitoid wasps are extremely diverse with more than 150,000 described species and estimates that suggest ~20% of all insects are parasitoids [66, 67]. Given the enormous number of wasp species with life histories that are similar to Lb, our findings point to the possible common strategy by which many parasitoids interact with the host gut microbiota to meet nutritional requirements that enhance their fitness. Our findings further suggest the possibility that other parasites including species that cause disease in humans and other vertebrates are also potentially affected by the gut microbiota of their hosts.
Data availability
16S rRNA gene sequence data have been deposited in the National Center for Biotechnology Information Sequence Read Archive (accession no. PRJNA550325 for experiment#1 and PRJNA679648 for experiment#2). All other data are available in the main text or Supplementary materials.
References
Zuzarte-Luís V, Mota MM. Parasite sensing of host nutrients and environmental cues. Cell Host Microbe. 2018;23:749–58.
Gilbert JA, Quinn RA, Debelius J, Xu ZZ, Morton J, Garg N, et al. Microbiome-wide association studies link dynamic microbial consortia to disease. Nature. 2016;535:94–103.
Valzania LK, Coon L, Vogel KJ, Brown MR, Strand MR. Hypoxia-induced transcription factor signaling is essential for larval growth of the mosquito Aedes aegypti. Proc Natl Acad Sci USA. 2018;115:457–65.
Vadder FD, Grasset E, Holm LM, Karsenty G, Macpherson AJ, Olofsson LE, et al. Gut microbiota regulates maturation of the adult enteric nervous system via enteric serotonin networks. Proc Natl Acad Sci USA. 2018;115:6458–63.
Motta EVS, Raymann K, Moran NA. Glyphosate perturbs the gut microbiota of honey bees. Proc Natl Acad Sci USA. 2018;115:10305–10.
Hooper LV, Littman DR, Macpherson AJ. Interactions between the microbiota and the immune system. Science. 2012;336:1268–73.
Clemente JC, Ursell LK, Parfrey LW, Knight R. The impact of the gut microbiota on human health: an integrative view. Cell. 2012;148:1258–70.
Mao K, Baptista AP, Tamoutounour S, Zhuang L, Bouladoux N, Martins AJ, et al. Innate and adaptive lymphocytes sequentially shape the gut microbiota and lipid metabolism. Nature. 2018;554:255–9.
Villarino NF, LeCleir GR, Denny JE, Dearth SP, Harding CL, Sloan SS, et al. Composition of the gut microbiota modulates the severity of malaria. Proc Natl Acad Sci USA. 2016;113:2235–40.
Narasimhan S, Rajeevan N, Liu L, Zhao YO, Heisig J, Pan J, et al. Gut microbiota of the tick vector Ixodes scapularis modulate colonization of the Lyme disease spirochete. Cell Host Microbe. 2014;15:58–71.
Dong Y, Manfredini F, Dimopoulos G. Implication of the mosquito midgut microbiota in the defense against malaria parasites. PLoS Pathog. 2009;5:e1000423.
Ling F, Steninel N, Weber J, Ma L, Smith C, Correa D, et al. The gut microbiota response to helminth infection depends on host sex and genotype. ISME J. 2020;14:1141–53.
Morita YS, Paul KS, Englund PT. Specialized fatty acid synthesis in African trypanosomes: myristate for GPI anchors. Science. 2000;288:140–3.
Luginbuehl LH, Menard GN, Kurup S, Erp HV, Radhakrishnan GV, Breakspear A, et al. Fatty acids in arbuscular mycorrhizal fungi are synthesized by the host plant. Science. 2017;356:117–78.
Visser B, Lann CL, Den Blanken FJ, Harvey JA, Van Alphen JJM. Loss of lipid synthesis as an evolutionary consequence of a parasitic lifestyle. Proc Natl Acad Sci USA. 2010;107:8677–82.
Rajendran E, Rajendran E, Hapuarachchi SV, Miller CM, Fairweather SJ, Cai Y, et al. Cationic amino acid transporters play key roles in the survival and transmission of apicomplexan parasites. Nat Commun. 2017;8:14455.
Nabity PD, Haus MJ, Berenbaum MR, DeLucia EH. Leaf-galling phylloxera on grapes reprograms host metabolism and morphology. Proc Natl Acad Sci USA. 2013;110:16663–8.
Mancio-Silva L, Slavic K, Grilo Ruivo MT, Grosso AR, Modrzynska KK, Vera IM, et al. Nutrient sensing modulates malaria parasite virulence. Nature. 2017;547:213–6.
Daily JP, Scanfeld D, Pochet N, Le Roch K, Plouffe D, Kamal M, et al. Distinct physiological states of Plasmodium falciparum in malaria-infected patients. Nature. 2007;450:1091–5.
Pernas L, Bean C, Boothroyd JC, Scorrano L. Mitochondria restrict growth of the intracellular parasite Toxoplasma gondii by limiting its uptake of fatty acids. Cell Metab. 2018;27:886–97.
Beckage NE, Gelman DB. Wasp parasitoid disruption of host development: Implications for new biologically based strategies for insect control. Annu Rev Entomol. 2004;49:299–330.
Whitfield JB, Austin AD, Fernandez-Triana JL. Systematics, biology, and evolution of microgastrine parasitoid wasps. Annu Rev Entomol. 2018;63:389–406.
Pennacchio F, Caccia S, Digilio MC. Host regulation and nutritional exploitation by parasitic wasps. Curr Opin Insect Sci. 2014;6:74–79.
Fisher RC, Ganesalingam VK. Changes in the composition of host haemolymph after attack by an insect parasitoid. Nature. 1970;227:191–2.
Vallochi AL, Teixeira L, da Silva Oliveira K, Maya-Monteiro CM, Bozza PT. Lipid droplet, a key player in host-parasite interactions. Front Immunol. 2018;9:1022.
Charron AJ, Sibley LD. Host cells: mobilizable lipid resources for the intracellular parasite Toxoplasma gondii. J Cell Sci. 2002;115:3049–59.
Walther TC, Farese RV. Lipid droplets and cellular lipid metabolism. Annu Rev Biochem. 2012;81:687–714.
Kühnlein RP. Lipid droplet-based storage fat metabolism in Drosophila. J Lipid Res. 2012;53:1430–6.
Hashemi HF, Goodman JM. The life cycle of lipid droplets. Curr Opin Cell Biol. 2015;33:119–24.
Fan W, Lam SM, Xin J, Yang X, Liu Z, Liu Y, et al. Drosophila TRF2 and TAF9 regulate lipid droplet size and phospholipid fatty acid composition. PLoS Genet. 2017;13:e1006664.
Lehmann M. Endocrine and physiological regulation of neutral fat storage in Drosophila. Mol Cell Endocrinol. 2018;461:165–77.
Wang B, Moya N, Niessen S, Hoover H, Mihaylova MM, Shaw RJ, et al. A hormone-dependent module regulating energy balance. Cell. 2011;145:596–606.
Grönke S, Mildner A, Fellert S, Tennagels N, Petry S, Müller G, et al. Brummer lipase is an evolutionary conserved fat storage regulator in Drosophila. Cell Metab. 2005;1:323–30.
Grönke S, Müller G, Hirsch J, Fellert S, Andreou A, Haase T, et al. Dual lipolytic control of body fat storage and mobilization in Drosophila. PLoS Biol. 2007;5:e137.
DiAngelo JR, Birnbaum MJ. Regulation of fat cell mass by insulin in Drosophila melanogaster. Mol Cell Biol 2009;29:6341–52.
Anderl I, Vesala L, Ihalainen TO, Vanha-Aho LM, Andó I, Rämet M, et al. Transdifferentiation and proliferation in two distinct hemocyte lineages in Drosophila melanogaster larvae after wasp infection. PLoS Pathog. 2016;12:e1005746.
Schlenke TA, Morales J, Govind S, Clark AG. Contrasting infection strategies in generalist and specialist wasp parasitoids of Drosophila melanogaster. PLoS Pathog. 2007;3:e158.
Liu Y, Wang W, Shui G, Huang X. CDP-diacylglycerol synthetase coordinates cell growth and fat storage through phosphatidylinositol metabolism and the insulin pathway. PLoS Genet. 2014;10:e1004172.
Ha EM, Lee KA, Seo YY, Kim SH, Lim JH, Oh BH, et al. Coordination of multiple dual oxidase-regulatory pathways in responses to commensal and infectious microbes in Drosophila gut. Nat Immunol. 2009;10:949–57.
Leitão-Gonçalves R, Carvalho-Santos Z, Francisco AP, Fioreze GT, Anjos M, Baltazar C, et al. Commensal bacteria and essential amino acids control food choice behavior and reproduction. PLoS Biol. 2017;15:e2000862.
Carton Y, Bouletreau M, van Alphen JJM, Van Lenteren JC. The Drosophila parasitic wasps. In: Ashburner M, Carson HL, Thompson JN, editors. The genetics and biology of Drosophila. London: Academic Press Inc. Ltd; 1986. pp 347–94.
Palanker L, Tennessen JM, Lam G, Thummel CS. Drosophila HNF4 regulates lipid mobilization and β-oxidation. Cell Metab. 2009;9:228–39.
Koyle ML, Veloz M, Judd AM, Wong AC, Newell PD, Douglas AE, et al. Rearing the fruit fly Drosophila melanogaster under axenic and gnotobiotic conditions. J Vis Exp. 2016. https://doi.org/10.3791/54219.
Aguila JR, Suszko J, Gibbs AG, Hoshizaki DK. The role of larval fat cells in adult Drosophila melanogaster. J Exp Biol. 2007;210:956–63.
Wang Y, Pu Y, Shen P. Neuropeptide-gated perception of appetitive olfactory inputs in Drosophila larvae. Cell Rep. 2013;3:820–30.
Lee KS, You KH, Choo JK, Han YM, Yu K. Drosophila short neuropeptide F regulates food intake and body size. J Biol Chem. 2004;279:50781–9.
Shin SC, Kim SH, You H, Kim B, Kim AC, Lee KA, et al. Drosophila microbiome modulates host developmental and metabolic homeostasis via insulin signaling. Science. 2011;334:670–4.
Lee KA, Cho C, Kim B, Jang IH, Nam K, Kwon YE, et al. Inflammation-modulated metabolic reprogramming is required for DUOX-dependent gut immunity in Drosophila. Cell Host Microbe. 2018;23:338–52.
Douglas AE. The Drosophila model for microbiome research. Lab Anim. 2018;47:157–64.
Broderick NA, Lemaitre B. Gut-associated microbes of Drosophila melanogaster. Gut Microbes. 2012;3:307–21.
Wong ACN, Chaston JM, Douglas AE. The inconstant gut microbiota of Drosophila species revealed by 16S rRNA gene analysis. ISME J. 2013;7:1922–32.
Chandler JA, Lang JM, Bhatnagar S, Eisen JA, Kopp A. Bacterial communities of diverse Drosophila species: Ecological context of a host-microbe model system. PLoS Genet. 2011;7:e1002272.
Obadia B, Güvener ZT, Zhang V, Ceja-Navarro JA, Brodie EL, Ja WW, et al. Probabilistic invasion underlies natural gut microbiome stability. Curr Biol. 2017;27:1999–2006.e8.
Ha EM, Oh CT, Bae YS, Lee WJ. A direct role for dual oxidase in Drosophila gut immunity. Science 2005;310:847–50.
Ryu JH, Kim SH, Lee HY, Bai JY, Nam YD, Bae JW, et al. Innate immune homeostasis by the homeobox gene caudal and commensal-gut mutualism in Drosophila. Science. 2008;319:777–82.
Ugrankar R, Liu Y, Provaznik J, Schmitt S, Lehmann M. Lipin is a central regulator of adipose tissue development and function in Drosophila melanogaster. Mol Cell Biol. 2011;1:1646–56.
Buszczak M, Lu X, Segraves WA, Chang TY, Cooley L. Mutations in the midway gene disrupt a Drosophila acyl coenzyme A: diacylglycerol acyltransferase. Genetics. 2002;160:1511–8.
Beller M, Bulankina AV, Hsiao HH, Urlaub H, Jäckle H, Kühnlein RP. Perilipin-dependent control of lipid droplet structure and fat storage in Drosophila. Cell Metab. 2010;12:521–32.
Molaei M, Vandehoef C, Karpac J. NF-κB shapes metabolic adaptation by attenuating Foxo-mediated lipolysis in Drosophila. Dev Cell. 2019;49:802–10. e6.
Kang P, Chang K, Liu Y, Bouska M, Birnbaum A, Karashchuk G, et al. Drosophila Kruppel homolog 1 represses lipolysis through interaction with dFOXO. Sci Rep. 2017;7:16369.
Gáliková M, Diesner M, Klepsatel P, Hehlert P, Xu Y, Bickmeyer I, et al. Energy homeostasis control in Drosophila adipokinetic hormone mutants. Genetics. 2015;201:665–83.
Britton JS, Lockwood WK, Li L, Cohen SM, Edgar BA. Drosophila’s insulin/PI3-kinase pathway coordinates cellular metabolism with nutritional conditions. Dev Cell. 2002;2:239–49.
Roth SW, Bitterman M D, Birnbaum MJ, Bland ML. Innate immune signaling in Drosophila blocks insulin signaling by uncoupling PI(3,4,5)P3 production and akt activation. Cell Rep. 2018;22:2550–6.
Brogiolo W, Stocker H, Ikeya T, Rintelen F, Fernandez R, Hafen E. An evolutionarily conserved function of the Drosophila insulin receptor and insulin-like peptides in growth control. Curr Biol. 2001;11:213–21.
Wang YP, Wu XT, Wang ZH, Chen T, Zhou SC, Chen JJ, et al. Symbiotic bracovirus of a parasite manipulates host lipid metabolism via tachykinin signaling. PLoS Pathog. 2021;17:e1009365.
Pennacchio F, Strand MR. Evolution of developmental strategies in parasitic hymenoptera. Annu Rev Entomol. 2006;51:233–58.
Zang LS, Wang S, Zhang F, Desneux N. Biological control with Trichogramma in China: history, present status, and perspectives. Annu Rev Entomol. 2021;66:463–84.
Acknowledgements
We thank Drs. Dan Hultmark, Sheng Li, Xun Huang, Chao Tong, Zhangwu Zhao, and Bloomington Drosophila Stock Center and TsingHua Fly Center for the wasp and fly stocks and antibodies; we also thank Drs. Daniel Kalderon and Shusheng Liu for their careful reading of the manuscript. This work was supported by the National Science Foundation of China (32172467 and 31622048 to JH; 31630060 to XXC), the Zhejiang Provincial Natural Science Foundation of China (LR18C140001 to JH), the National Key R&D Program of China (2017YFD0200400 to JH; 2019YFD0300100 to XXC), the Provincial Key R&D Program of Zhejiang (2021C02045 to XXC), the US National Institutes of Health, US Department of Agriculture, and Pulliam Endowment (R01 AI106892, GEO0772 to MRS).
Author information
Authors and Affiliations
Contributions
JH conceived and supervised the project. JH, MRS, and XXC designed the experiments. SZ, YL, JC, ZP, LP, YW, and QZ performed the experiments. SZ, YL, MRS, and JH analyzed data. JH, MRS, and SZ wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhou, S., Lu, Y., Chen, J. et al. Parasite reliance on its host gut microbiota for nutrition and survival. ISME J 16, 2574–2586 (2022). https://doi.org/10.1038/s41396-022-01301-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-022-01301-z