Abstract
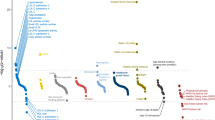
We report a genome-wide association study (GWAS) of coronary artery disease (CAD) incorporating nearly a quarter of a million cases, in which existing studies are integrated with data from cohorts of white, Black and Hispanic individuals from the Million Veteran Program. We document near equivalent heritability of CAD across multiple ancestral groups, identify 95 novel loci, including nine on the X chromosome, detect eight loci of genome-wide significance in Black and Hispanic individuals, and demonstrate that two common haplotypes at the 9p21 locus are responsible for risk stratification in all populations except those of African origin, in which these haplotypes are virtually absent. Moreover, in the largest GWAS for angiographically derived coronary atherosclerosis performed to date, we find 15 loci of genome-wide significance that robustly overlap with established loci for clinical CAD. Phenome-wide association analyses of novel loci and polygenic risk scores (PRSs) augment signals related to insulin resistance, extend pleiotropic associations of these loci to include smoking and family history, and precisely document the markedly reduced transferability of existing PRSs to Black individuals. Downstream integrative analyses reinforce the critical roles of vascular endothelial, fibroblast, and smooth muscle cells in CAD susceptibility, but also point to a shared biology between atherosclerosis and oncogenesis. This study highlights the value of diverse populations in further characterizing the genetic architecture of CAD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Summary statistics for the Biobank Japan study were obtained from http://jenger.riken.jp/en/result. Summary statistics for the CARDIoGRAMplusC4D study were obtained from http://www.cardiogramplusc4d.org. Summary statistics for the UK Biobank study for CAD were obtained from https://www.cardiomics.net/download-data. The full summary level association data from the individual population association analyses in MVP as well as the multi-population meta-analysis from this report will be available via the dbGaP Study accession number phs001672. This research has been conducted using the UK Biobank Resource under Application Numbers 13721 and 19416.
References
Roth, G. A. et al. Global burden of cardiovascular diseases and risk factors, 1990–2019: update from the GBD 2019 Study. J. Am. Coll. Cardiol. 76, 2982–3021 (2020).
National Center for Health Statistics. Health, United States Spotlight: Racial and Ethnic Disparities in Heart Disease (Centers for Disease Control and Prevention, 2019).
Churchwell, K. et al. Call to action: structural racism as a fundamental driver of health disparities: a Presidential advisory from the American Heart Association. Circulation 142, e454–e468 (2020).
Popejoy, A. B. & Fullerton, S. M. Genomics is failing on diversity. Nature 538, 161–164 (2016).
Martin, A. R. et al. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 51, 584–591 (2019).
Clarke, S. L., Assimes, T. L. & Tcheandjieu, C. The propagation of racial disparities in cardiovascular genomics research. Circ. Genom. Precis. Med. 14, e003178 (2021).
Zdravkovic, S. et al. Heritability of death from coronary heart disease: a 36-year follow-up of 20 966 Swedish twins. J. Intern. Med. 252, 247–254 (2002).
Wienke, A., Holm, N. V., Skytthe, A. & Yashin, A. I. The heritability of mortality due to heart diseases: a correlated frailty model applied to Danish twins. Twin Res. 4, 266–274 (2001).
van der Harst, P. & Verweij, N. Identification of 64 novel genetic loci provides an expanded view on the genetic architecture of coronary artery disease. Circ. Res. 122, 433–443 (2018).
Koyama, S. et al. Population-specific and trans-ancestry genome-wide analyses identify distinct and shared genetic risk loci for coronary artery disease. Nat. Genet. 52, 1169–1177 (2020).
Webb, T. R. et al. Systematic evaluation of pleiotropy identifies 6 further loci associated with coronary artery disease. J. Am. Coll. Cardiol. 69, 823–836 (2017).
Coronary Artery Disease (C4D) Genetics Consortium. A genome-wide association study in Europeans and South Asians identifies five new loci for coronary artery disease. Nat. Genet. 43, 339–344 (2011).
Lu, X. et al. Genome-wide association study in Han Chinese identifies four new susceptibility loci for coronary artery disease. Nat. Genet. 44, 890–894 (2012).
Assimes, T. L. & Roberts, R. Genetics: implications for prevention and management of coronary artery disease. J. Am. Coll. Cardiol. 68, 2797–2818 (2016).
Buniello, A. et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47, D1005–D1012 (2019).
National Center for Health Statistics (US). Crude percentages of all types of heart disease for adults aged 18 and over, United States, 2015-2018. National Health Interview Survey. Generated interactively: Wed Jul 13 2022.
Institute for Health Metrics and Evaluation (IHME). GBD Compare Data Visualization (IHME, University of Washington, 2020). http://vizhub.healthdata.org/gbd-compare
Nikpay, M. et al. A comprehensive 1,000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat. Genet. 47, 1121–1130 (2015).
Ishigaki, K. et al. Large-scale genome-wide association study in a Japanese population identifies novel susceptibility loci across different diseases. Nat. Genet. 52, 669–679 (2020).
Barbalic, M. et al. Genome-wide association analysis of incident coronary heart disease (CHD) in African Americans: a short report. PLoS Genet. 7, e1002199 (2011).
Lettre, G. et al. Genome-wide association study of coronary heart disease and its risk factors in 8,090 African Americans: the NHLBI CARe Project. PLoS Genet. 7, e1001300 (2011).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224 (2018).
Inouye, M. et al. Genomic risk prediction of coronary artery disease in 480,000 adults: implications for primary prevention. J. Am. Coll. Cardiol. 72, 1883–1893 (2018).
Dikilitas, O. et al. Predictive utility of polygenic risk scores for coronary heart disease in three major racial and ethnic groups. Am. J. Hum. Genet. 106, 707–716 (2020).
Speliotes, E. K. et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 42, 937–948 (2010).
Vujkovic, M. et al. Discovery of 318 new risk loci for type 2 diabetes and related vascular outcomes among 1.4 million participants in a multi-ancestry meta-analysis. Nat. Genet. 52, 680–691 (2020).
Liu, D. J. et al. Exome-wide association study of plasma lipids in >300,000 individuals. Nat. Genet. 49, 1758–1766 (2017).
Hartiala, J. A. et al. Genome-wide analysis identifies novel susceptibility loci for myocardial infarction. Eur. Heart J. 42, 919–933 (2021).
Wainschtein, P. et al. Assessing the contribution of rare variants to complex trait heritability from whole-genome sequence data. Nat. Genet. 54, 263–273 (2022).
McPherson, R. et al. A common allele on chromosome 9 associated with coronary heart disease. Science 316, 1488–1491 (2007).
CARDIoGRAMplusC4D Consortium et al. Large-scale association analysis identifies new risk loci for coronary artery disease. Nat. Genet. 45, 25–33 (2013).
Shungin, D. et al. New genetic loci link adipose and insulin biology to body fat distribution. Nature 518, 187–196 (2015).
Huang, Y. et al. Sexual differences in genetic predisposition of coronary artery disease. Circ. Genom. Precis. Med. 14, e003147 (2021).
Zore, T., Palafox, M. & Reue, K. Sex differences in obesity, lipid metabolism, and inflammation: a role for the sex chromosomes? Mol. Metab. 15, 35–44 (2018).
Salfati, E. et al. Susceptibility loci for clinical coronary artery disease and subclinical coronary atherosclerosis throughout the life-course. Circ. Cardiovasc. Genet. 8, 803–811 (2015).
Speliotes, E. K. et al. Genome-wide association analysis identifies variants associated with nonalcoholic fatty liver disease that have distinct effects on metabolic traits. PLoS Genet. 7, e1001324 (2011).
Natarajan, P. et al. Chromosome Xq23 is associated with lower atherogenic lipid concentrations and favorable cardiometabolic indices. Nat. Commun. 12, 2182 (2021).
Fletcher, R. et al. The role of the Niemann-Pick disease, type C1 protein in adipocyte insulin action. PLoS One 9, e95598 (2014).
Ghuran, A., van Der Wieken, L. R. & Nolan, J. Cardiovascular complications of recreational drugs. BMJ 323, 464–466 (2001).
Wirka, R. C. et al. Atheroprotective roles of smooth muscle cell phenotypic modulation and the TCF21 disease gene as revealed by single-cell analysis. Nat. Med. 25, 1280–1289 (2019).
Nelson, C. P. et al. Genetically determined height and coronary artery disease. N. Engl. J. Med. 372, 1608–1618 (2015).
Ong, J. S. et al. Height and overall cancer risk and mortality: evidence from a Mendelian randomisation study on 310,000 UK Biobank participants. Br. J. Cancer 118, 1262–1267 (2018).
Clarke, S. L. et al. Broad clinical manifestations of polygenic risk for coronary artery disease in the Women’s Health Initiative. Preprint at https://doi.org/10.1101/2021.06.15.21258993 (2022).
Xiao, B. et al. Inference of causal relationships based on the genetics of cardiometabolic traits and conditions unique to females in >50,000 participants. Preprint at https://doi.org/10.1101/2022.02.02.22269844 (2022).
Singh, K. K. et al. BRCA1 is a novel target to improve endothelial dysfunction and retard atherosclerosis. J. Thorac. Cardiovasc. Surg. 146, 949–960 (2013).
Wu, H. T. et al. Oncogenic functions of the EMT-related transcription factor ZEB1 in breast cancer. J. Transl. Med. 18, 51 (2020).
Ibrahim, N. et al. BRCA1-associated epigenetic regulation of p73 mediates an effector pathway for chemosensitivity in ovarian carcinoma. Cancer Res. 70, 7155–7165 (2010).
Bai, F. et al. BRCA1 suppresses epithelial-to-mesenchymal transition and stem cell dedifferentiation during mammary and tumor development. Cancer Res. 74, 6161–6172 (2014).
Fardi, M., Alivand, M., Baradaran, B., Farshdousti Hagh, M. & Solali, S. The crucial role of ZEB2: from development to epithelial-to-mesenchymal transition and cancer complexity. J. Cell. Physiol. https://doi.org/10.1002/jcp.28277 (2019).
Soini, Y. et al. Transcription factors zeb1, twist and snai1 in breast carcinoma. BMC Cancer 11, 73 (2011).
Cheng, P. et al. ZEB2 shapes the epigenetic landscape of atherosclerosis. Circulation 145, 469–485 (2022).
Tabas, I., Garcia-Cardena, G. & Owens, G. K. Recent insights into the cellular biology of atherosclerosis. J. Cell Biol. 209, 13–22 (2015).
Nagao, M. et al. Coronary disease-associated gene TCF21 inhibits smooth muscle cell differentiation by blocking the myocardin-serum response factor pathway. Circ. Res. 126, 517–529 (2020).
Lowrie Jr, D. J. Histology: An Essential Textbook (Thieme Publishers, 2020).
Ko, C. W., Qu, J., Black, D. D. & Tso, P. Regulation of intestinal lipid metabolism: current concepts and relevance to disease. Nat. Rev. Gastroenterol. Hepatol. 17, 169–183 (2020).
Fahed, A. C. et al. Transethnic transferability of a genome-wide polygenic score for coronary artery disease. Circ. Genom. Precis. Med. 14, e003092 (2021).
Gaziano, J. M. et al. Million Veteran Program: a mega-biobank to study genetic influences on health and disease. J. Clin. Epidemiol. 70, 214–223 (2016).
Hunter-Zinck, H. et al. Genotyping array design and data quality control in the Million Veteran Program. Am. J. Hum. Genet. 106, 535–548 (2020).
Genomes Project Consortiumet al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Loh, P. R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 48, 1443–1448 (2016).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Fang, H. et al. Harmonizing genetic ancestry and self-identified race/ethnicity in genome-wide association studies. Am. J. Hum. Genet. 105, 763–772 (2019).
Byrd, J. B. et al. Data quality of an electronic health record tool to support VA cardiac catheterization laboratory quality improvement: the VA Clinical Assessment, Reporting, and Tracking System for Cath Labs (CART) program. Am. Heart J. 165, 434–440 (2013).
Maddox, T. M. et al. A national clinical quality program for Veterans Affairs catheterization laboratories (from the Veterans Affairs clinical assessment, reporting, and tracking program). Am. J. Cardiol. 114, 1750–1757 (2014).
Maddox, T. M. et al. Nonobstructive coronary artery disease and risk of myocardial infarction. JAMA 312, 1754–1763 (2014).
Yang, J. et al. Genetic variance estimation with imputed variants finds negligible missing heritability for human height and body mass index. Nat. Genet. 47, 1114–1120 (2015).
Evans, L. M. et al. Comparison of methods that use whole genome data to estimate the heritability and genetic architecture of complex traits. Nat. Genet. 50, 737–745 (2018).
Lee, S. H., Wray, N. R., Goddard, M. E. & Visscher, P. M. Estimating missing heritability for disease from genome-wide association studies. Am. J. Hum. Genet. 88, 294–305 (2011).
Lee, S. H. et al. Estimating the proportion of variation in susceptibility to schizophrenia captured by common SNPs. Nat. Genet. 44, 247–250 (2012).
Visscher, P. M. et al. Statistical power to detect genetic (co)variance of complex traits using SNP data in unrelated samples. PLoS Genet. 10, e1004269 (2014).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Magi, R. et al. Trans-ethnic meta-regression of genome-wide association studies accounting for ancestry increases power for discovery and improves fine-mapping resolution. Hum. Mol. Genet. 26, 3639–3650 (2017).
Loley, C. et al. No association of coronary artery disease with X-chromosomal variants in comprehensive international meta-analysis. Sci. Rep. 6, 35278 (2016).
Graham, S. E. et al. The power of genetic diversity in genome-wide association studies of lipids. Nature 600, 675–679 (2021).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Coram, M. A. et al. Leveraging multi-ethnic evidence for mapping complex traits in minority populations: an empirical Bayes approach. Am. J. Hum. Genet. 96, 740–752 (2015).
Choi, S. W. & O’Reilly, P. F. PRSice-2: Polygenic Risk Score software for biobank-scale data. Gigascience 8, giz082 (2019).
Denny, J. C. et al. PheWAS: demonstrating the feasibility of a phenome-wide scan to discover gene–disease associations. Bioinformatics 26, 1205–1210 (2010).
Wu, P. et al. Mapping ICD-10 and ICD-10-CM codes to phecodes: workflow development and initial evaluation. JMIR Med. Inform. 7, e14325 (2019).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 10, e1004383 (2014).
Global Lipids Genetics Consortiumet al. Discovery and refinement of loci associated with lipid levels. Nat. Genet. 45, 1274–1283 (2013).
Evangelou, E. et al. Genetic analysis of over 1 million people identifies 535 new loci associated with blood pressure traits. Nat. Genet. 50, 1412–1425 (2018).
Erzurumluoglu, A. M. et al. Meta-analysis of up to 622,409 individuals identifies 40 novel smoking behaviour associated genetic loci. Mol. Psychiatry 25, 2392–2409 (2020).
Yengo, L. et al. Meta-analysis of genome-wide association studies for height and body mass index in approximately 700000 individuals of European ancestry. Hum. Mol. Genet. 27, 3641–3649 (2018).
Xue, A. et al. Genome-wide association analyses identify 143 risk variants and putative regulatory mechanisms for type 2 diabetes. Nat. Commun. 9, 2941 (2018).
Maples, B. K., Gravel, S., Kenny, E. E. & Bustamante, C. D. RFMix: a discriminative modeling approach for rapid and robust local-ancestry inference. Am. J. Hum. Genet. 93, 278–288 (2013).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
de Leeuw, C. A., Stringer, S., Dekkers, I. A., Heskes, T. & Posthuma, D. Conditional and interaction gene-set analysis reveals novel functional pathways for blood pressure. Nat. Commun. 9, 3768 (2018).
Zhu, X. & Stephens, M. Large-scale genome-wide enrichment analyses identify new trait-associated genes and pathways across 31 human phenotypes. Nat. Commun. 9, 4361 (2018).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Barbeira, A. N. et al. Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics. Nat. Commun. 9, 1825 (2018).
Acknowledgements
This research is based on data from the Million Veteran Program, Office of Research and Development, Veterans Health Administration, and was supported by Veterans Administration awards I01-01BX003362, I01-BX004821 (K.-M.C., P.S.T.), I01-BX003340 (K.Cho, P.W.F.W.) and VA HSR RES 13-457 (VA Informatics and Computing Infrastructure). The content of this manuscript does not represent the views of the Department of Veterans Affairs or the United States Government. The eMERGE Network was initiated and funded by the National Human Genome Research Institute (NHGRI) through the following grants: Phase III: U01HG8657 (Kaiser Permanente Washington/University of Washington); U01HG8685 (Brigham and Women’s Hospital); U01HG8672 (Vanderbilt University Medical Center); U01HG8666 (Cincinnati Children’s Hospital Medical Center); U01HG6379 (Mayo Clinic); U01HG8679 (Geisinger Clinic); U01HG8680 (Columbia University Health Sciences); U01HG8684 (Children’s Hospital of Philadelphia); U01HG8673 (Northwestern University); U01HG8701 (Vanderbilt University Medical Center serving as the Coordinating Center); U01HG8676 (Partners Healthcare/Broad Institute); and U01HG8664 (Baylor College of Medicine); Phase II: U01HG006828 (Cincinnati Children’s Hospital Medical Center/Boston Children’s Hospital); U01HG006830 (Children’s Hospital of Philadelphia); U01HG006389 (Essentia Institute of Rural Health, Marshfield Clinic Research Foundation and Pennsylvania State University); U01HG006382 (Geisinger Clinic); U01HG006375 (Group Health Cooperative/University of Washington); U01HG006379 (Mayo Clinic); U01HG006380 (Icahn School of Medicine at Mount Sinai); U01HG006388 (Northwestern University); U01HG006378 (Vanderbilt University Medical Center); and U01HG006385 (Vanderbilt University Medical Center serving as the Coordinating Center). Phase II: U01HG004438 (CIDR) and U01HG004424 (the Broad Institute) serving as Genotyping Centers. Phase I: U01-HG-004610 (Group Health Cooperative/University of Washington); U01-HG-004608 (Marshfield Clinic Research Foundation and Vanderbilt University Medical Center); U01-HG-04599 (Mayo Clinic); U01HG004609 (Northwestern University); U01-HG-04603 (Vanderbilt University Medical Center, also serving as the Administrative Coordinating Center); U01HG004438 (CIDR) and U01HG004424 (the Broad Institute) serving as Genotyping Centers. The Population Architecture Using Genomics and Epidemiology (PAGE) program is funded by the NHGRI with co-funding from the National Institute on Minority Health and Health Disparities (NIMHD), supported by U01HG007416 (CALiCo), U01HG007417 (ISMMS), U01HG007397 (MEC), U01HG007376 (WHI), and U01HG007419 (Coordinating Center). The MultiEthnic Study (MEC) was supported by U01 CA164973. The Women’s Health Initiative (WHI) program is funded by the National Heart, Lung, and Blood Institute, National Institutes of Health, US Department of Health and Human Services through contracts HHSN268201100046C, HHSN268201100001C, HHSN268201100002C, HHSN268201100003C, HHSN268201100004C and HHSN271201100004C. Scientific Computing Infrastructure at Fred Hutch is funded by ORIP grant S10OD028685. Funding support for the ‘Exonic variants and their relation to complex traits in minorities of the WHI study is provided through the NHGRI PAGE program (U01HG004790). The Atherosclerosis Risk in Communities (ARIC) Study is carried out as a collaborative study supported by National Heart, Lung, and Blood Institute contracts (HHSN268201100005C, HHSN268201100006C, HHSN268201100007C, HHSN268201100008C, HHSN268201100009C, HHSN268201100010C, HHSN268201100011C and HHSN268201100012C). The Cardiovascular Health Study (CHS) was supported by NHLBI contracts HHSN268201200036C, HHSN268200800007C, HHSN268201800001C, N01HC55222, N01HC85079, N01HC85080, N01HC85081, N01HC85082, N01HC85083, N01HC85086, 75N92021D00006; and NHLBI grants U01HL080295, R01HL085251, R01HL087652, R01HL105756, R01HL103612, R01HL120393 and U01HL130114 with additional contribution from the National Institute of Neurological Disorders and Stroke (NINDS). Additional support was provided through R01AG023629 from the National Institute on Aging (NIA). A full list of principal CHS investigators and institutions can be found at CHS-NHLBI.org. The provision of genotyping data was supported in part by the National Center for Advancing Translational Sciences, CTSI grant UL1TR001881, and the National Institute of Diabetes and Digestive and Kidney Disease Diabetes Research Center (DRC) grant DK063491 to the Southern California Diabetes Endocrinology Research Center. BioBank Japan (BBJ) was supported by the Tailor-Made Medical Treatment Program of the Ministry of Education, Culture, Sports, Science, and Technology and Japan Agency for Medical Research (AMED) under grant numbers JP17km0305002 and JP17km0305001. Healthy Aging in Neighborhoods of Diversity across the Life Span (HANDLS) was funded by the Interlaboratory Proposal Funding of the Intramural Research Program of the National Institute on Aging (NIA), the National Institutes of Health (NIH), Baltimore, Maryland. Funding number: [AG000989]. X.Z. was supported by the Stein Fellowship from Stanford University and Institute for Computational and Data Sciences Seed Grant from The Pennsylvania State University. S.M.D., J.A.L. and K.M.L. were supported by the US Department of Veterans Affairs (IK2-CX001780). Y.L. is supported by NIH R56HL150186. S.Ko. and K.It. were supported by AMED under Grant Numbers JP20km0405209 and JP20ek0109487. K.E.N. is supported by NIH R01HL142302. R.D. is supported by NIH R35GM124836 and R01HL139865. F.C. is supported by NCI T32CA229110. B.F.V. was supported by the NIH R01DK101478 and a Linda Pechenik Montague Investigator Award. P.N. is supported by grants from the NIH/NHLBI (R01HL142711, R01HL148050, R01HL127564, R01HL151152), NIH/NHGRI (U01HG011719), Fondation Leducq (TNE-18CVD04) and Massachusetts General Hospital (Fireman Chair). Support for title page creation and format was provided by AuthorArranger, a tool developed at the National Cancer Institute. The authors thank C. D. Bustamante for his review and feedback of specific cross-population analyses involving the 9p21 region. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Consortia
Contributions
Concept and design: C.T., X.Z., A.T.H., S.L.C., V.N., D.J.R., K.-M.C., J.A.L., S.M.D., P.W.F.W., H.T., Y.V.S., P.S.T., C.J.O., T.L.A. Acquisition, analysis or interpretation of data: C.T., X.Z., A.T.H., S.L.C., V.N., S.M., B.R.G., K.M.L., R.S., K.M.K., H.F., F.C., Y.L., S.Ko., N.L.T., M.Vu., S.R., M.E.P., T.M.M., S.W.W., A.G.B., M.G.L., S.P., J.Hu., N.S.-A., Y.-L.H., G.L.W., S.B., C.K., J.Ha., R.J.F.L., R.D., M.Ve., K.Cha., K.E.N., C.L.A., M.G., C.A.H., L.L.M., L.R.W., J.C.B., H.L., B.S., L.A.La., A.G., O.D., I.J.K., I.B.S., G.P.J., A.S.G., S.H., B.N., K.It., K.Is., Y.K., S.S.V., M.D.R., R.L.K., A.B., L.A.Lo., S.Ka., E.R.H., D.R.M., J.S.L., D.S., P.D.R., K.Cho, J.M.G., J.E.H., B.F.V., D.J.R., K.-M.C., J.A.L., S.M.D., P.W.F.W, H.T., Y.V.S., P.S.T., C.J.O., T.L.A. Drafting of the manuscript: C.T., T.L.A. Critical revision of the manuscript for important intellectual content: X.Z., A.T.H., S.L.C., V.N., M.Vu., D.K., S.R., M.G.L., R.D., K.E.N., C.K., J.C.B., I.J.K., M.D.R., P.N., B.F.V., J.A.L., S.M.D., P.W.F.W, H.T., Y.V.S., P.S.T, C.J.O.
Corresponding authors
Ethics declarations
Competing interests
A.B. and L.A.Lo. are employees of Regeneron Pharmaceuticals. R.D. has received grants from AstraZeneca, grants and non-financial support from Goldfinch Bio, is a scientific co-founder, consultant and equity holder for Pensieve Health and a consultant for Variant Bio. T.M.M. is an employee of the Healthcare Innovation Lab at BJC HealthCare/Washington University School of Medicine, an advisor of Myia Labs, and a compensated director of the JF Maddox Foundation in New Mexico. S.Ka. is an employee of Verve Therapeutics, holds equity in Verve Therapeutics and Maze Therapeutics, and has served as a consultant for Acceleron, Eli Lilly, Novartis, Merck, Novo Nordisk, Novo Ventures, Ionis, Alnylam, Aegerion, Haug Partners, Noble Insights, Leerink Partners, Bayer Healthcare, Illumina, Color Genomics, MedGenome, Quest and Medscape. D.J.R. is on the Scientific Advisory Board of Alnylam, Novartis and Verve Therapeutics. M.D.R. is on the scientific advisory board for Goldfinch Bio and Cipherome. C.J.O. became an employee of Novartis after the initial submission of the manuscript. P.N. reports investigator-initiated grants from Amgen, Apple, AstraZeneca, Boston Scientific and Novartis, personal fees from Apple, AstraZeneca, Blackstone Life Sciences, Invitae, Foresite Labs, Novartis and Roche/Genentech, and is a co-founder of TenSixteen Bio, a shareholder of geneXwell, TenSixteen Bio and Vertex, a scientific advisory board member of geneXwell and TenSixteen Bio, and reports spousal employment at Vertex, all unrelated to the present work. S.M.D. receives research support from RenalytixAI to his institution and consulting fees from Calico Labs. A.G.B. is a scientific co-founder and equity holder in TenSixteen Bio. All other authors have no competing interests.
Peer review
Peer review information
Nature Medicine thanks Riyaz Patel and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Michael Basson, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 LocusZoom plots of loci reaching genome-wide significance in Black participants and Hispanic participants.
Sets of LocusZoom plots for five loci in Black participants and 3 loci in Hispanic participants reaching genome-wide significance after two-stage meta-analysis with external cohorts. Each set of plots show the association results for a locus for all three populations using the same chromosome location scale (x-axis) but not the same p-value scale (y-axis). P values are derived from inverse variance-weighted meta-analysis using METAL and are two-sided.
Extended Data Fig. 2 Allele frequencies and association results at the 9p21 locus among Black in the Million Veteran Program stratified by local ancestry status.
Top panels show plots of corresponding allelic frequencies at the 9p21 susceptibility locus observed in MVP white participants vs. subgroups of MVP Black participants with a. two African chromosomes (chr), b. one African chr, and c. no African chr at the locus. Corresponding LocusZoom plots for each group are in the panels immediately below. Association testing was performed using logistic regression with adjustment on sex and principal component as implemented in PLINK. P values were derived from a Wald test and are two-sided.
Extended Data Fig. 3 LocusZoom plots of SNP association at the 9p21 susceptibility locus for CAD.
Top panel plots the results for MVP GWAS of all Hispanic participants + Stage 2 cohort meta-analysis. P values are derived from inverse variance weighted meta-analysis using METAL and are two-sided. Bottom panel plots the subset of MVP Hispanic participants with no African derived chromosomes at 9p21 based on local ancestry assessment using RFMix (5,298 cases/20,556 controls). Association testing was performed using logistic regression with adjustment on sex and principal component as implemented in PLINK. P values were derived from a Wald test and are two-sided.
Supplementary information
Supplementary Information
Supplementary Methods and list of Consortia authors
Supplementary Table 1
Supplementary Tables 1–37 and list of abbreviations used in the Tables
Rights and permissions
About this article
Cite this article
Tcheandjieu, C., Zhu, X., Hilliard, A.T. et al. Large-scale genome-wide association study of coronary artery disease in genetically diverse populations. Nat Med 28, 1679–1692 (2022). https://doi.org/10.1038/s41591-022-01891-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-022-01891-3
This article is cited by
-
Principles and methods for transferring polygenic risk scores across global populations
Nature Reviews Genetics (2024)
-
Genetic ancestry and diagnostic yield of exome sequencing in a diverse population
npj Genomic Medicine (2024)
-
To advance science we need to address ‘otherness’
Nature Human Behaviour (2024)
-
EGR1 transcriptionally regulates SVEP1 to promote proliferation and migration in human coronary artery smooth muscle cells
Molecular Biology Reports (2024)
-
Convergence of coronary artery disease genes onto endothelial cell programs
Nature (2024)