Abstract
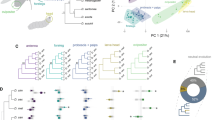
Despite numerous examples of chemoreceptor gene family expansions and contractions, how these relate to modifications in the sensory neuron populations in which they are expressed remains unclear. Drosophila melanogaster’s odorant receptor (Or) family is ideal for addressing this question because most Ors are expressed in distinct olfactory sensory neuron (OSN) types. Between-species changes in Or copy number may therefore indicate increases or reductions in the number of OSN populations. Here we investigated the Or67a subfamily, which exhibits copy number variation in D. melanogaster and its closest relatives: D. simulans, D. sechellia and D. mauritiana. These species’ common ancestor had three Or67a paralogues that had already diverged adaptively. Following speciation, two Or67a paralogues were lost independently in D. melanogaster and D. sechellia, with ongoing positive selection shaping the intact genes. Unexpectedly, the functionally diverged Or67a paralogues in D. simulans are co-expressed in a single neuron population, which projects to a glomerulus homologous to that innervated by Or67a neurons in D. melanogaster. Thus, while sensory pathway neuroanatomy is conserved, independent selection on co-expressed receptors has contributed to species-specific peripheral coding. This work reveals a type of adaptive change largely overlooked for olfactory evolution, raising the possibility that similar processes influence other cases of insect Or co-expression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All raw data generated for this study are available in the supplementary files or in the GitLab repository at https://gitlab.com/roman.arguello/or67a_dsim_trio.
Code availability
The code for this study is available in the GitLab repository at https://gitlab.com/roman.arguello/or67a_dsim_trio.
References
Gilad, Y., Wiebe, V., Przeworski, M., Lancet, D. & Pääbo, S. Loss of olfactory receptor genes coincides with the acquisition of full trichromatic vision in primates. PLoS Biol. 2, E5–E5 (2004).
Hughes, G. M. et al. The birth and death of olfactory receptor gene families in mammalian niche adaptation. Mol. Biol. Evol. 35, 1390–1406 (2018).
McBride, C. S. & Arguello, J. R. Five Drosophila genomes reveal nonneutral evolution and the signature of host specialization in the chemoreceptor superfamily. Genetics 177, 1395–1416 (2007).
Niimura, Y., Matsui, A. & Touhara, K. Extreme expansion of the olfactory receptor gene repertoire in African elephants and evolutionary dynamics of orthologous gene groups in 13 placental mammals. Genome Res. 24, 1485–1496 (2014).
Robertson, H. M. Molecular evolution of the major arthropod chemoreceptor gene families. Annu. Rev. Entomol. 64, 227–242 (2019).
Robertson, H. M. & Wanner, K. W. The chemoreceptor superfamily in the honey bee, Apis mellifera: expansion of the odorant, but not gustatory, receptor family. Genome Res. 16, 1395–1403 (2006).
McKenzie, S. K. et al. The genomic basis of army ant chemosensory adaptations. Mol. Ecol. https://doi.org/10.1111/mec.16198 (2021).
Zhao, H., Li, J. & Zhang, J. Molecular evidence for the loss of three basic tastes in penguins. Curr. Biol. 25, R141–R142 (2015).
Zhao, H., Yang, J. R., Xu, H. & Zhang, J. Pseudogenization of the umami taste receptor gene Tas1r1 in the giant panda coincided with its dietary switch to bamboo. Mol. Biol. Evol. 27, 2669–2673 (2010).
Guo, S. & Kim, J. Molecular evolution of Drosophila odorant receptor genes. Mol. Biol. Evol. 24, 1198–1207 (2007).
Nei, M. & Rooney, A. P. Concerted and birth-and-death evolution of multigene families. Annu. Rev. Genet. 39, 121–152 (2005).
Allen, A. M. et al. A single-cell transcriptomic atlas of the adult Drosophila ventral nerve cord. eLife https://doi.org/10.7554/eLife.54074 (2020).
Eschbach C. et al. Circuits for integrating learned and innate valences in the insect brain eLife 10:e62567 (2021).
Li, H. et al. Fly Cell Atlas: a single-nucleus transcriptomic atlas of the adult fruit fly. Science 375, eabk2432 (2022).
Zheng, Z. et al. A complete electron microscopy volume of the brain of adult Drosophila melanogaster. Cell 174, 730–743.e722 (2018).
Auer, T. O. et al. Olfactory receptor and circuit evolution promote host specialization. Nature 579, 402–408 (2020).
Ding, Y. et al. Neural evolution of context-dependent fly song. Curr. Biol. 29, 1089–1099.e1087 (2019).
Prieto-Godino, L. L. et al. Evolution of acid-sensing olfactory circuits in drosophilids. Neuron 93, 661–676.e666 (2017).
Seeholzer, L. F., Seppo, M., Stern, D. L. & Ruta, V. Evolution of a central neural circuit underlies Drosophila mate preferences. Nature https://doi.org/10.1038/s41586-018-0322-9 (2018).
Stern, D. L. et al. Genetic and transgenic reagents for Drosophila simulans, D. mauritiana, D. yakuba, D. santomea, and D. virilis. G3 (Bethesda) 7, 1339–1347 (2017).
Larsson, M. C. et al. Or83b encodes a broadly expressed odorant receptor essential for Drosophila olfaction. Neuron 43, 703–714 (2004).
Lewcock, J. W. & Reed, R. R. A feedback mechanism regulates monoallelic odorant receptor expression. Proc. Natl Acad. Sci. USA 101, 1069–1074 (2004).
Serizawa, S. et al. Negative feedback regulation ensures the one receptor–one olfactory neuron rule in mouse. Science 302, 2088–2094 (2003).
Couto, A., Alenius, M. & Dickson, B. J. Molecular, anatomical, and functional organization of the Drosophila olfactory system. Curr. Biol. 15, 1535–1547 (2005).
Gardiner, A., Barker, D., Butlin, R. K., Jordan, W. C. & Ritchie, M. G. Drosophila chemoreceptor gene evolution: selection, specialization and genome size. Mol. Ecol. 17, 1648–1657 (2008).
Obbard, D. J. et al. Estimating divergence dates and substitution rates in the Drosophila phylogeny. Mol. Biol. Evol. 29, 3459–3473 (2012).
Ometto, L. et al. Linking genomics and ecology to investigate the complex evolution of an invasive Drosophila pest. Genome Biol. Evol. 5, 745–757 (2013).
Ramasamy, S. et al. The evolution of olfactory gene families in Drosophila and the genomic basis of chemical–ecological adaptation in Drosophila suzukii. Genome Biol. Evol. 8, 2297–2311 (2016).
Dweck, H. K. M. et al. The olfactory logic behind fruit odor preferences in larval and adult Drosophila. Cell Rep. 23, 2524–2531 (2018).
Garrigan, D. et al. Genome sequencing reveals complex speciation in the Drosophila simulans clade. Genome Res. 22, 1499–1511 (2012).
Schrider, D. R., Ayroles, J., Matute, D. R. & Kern, A. D. Supervised machine learning reveals introgressed loci in the genomes of Drosophila simulans and D. sechellia. PLoS Genet. 14, e1007341 (2018).
Butterwick, J. A. et al. Cryo-EM structure of the insect olfactory receptor Orco. Nature 560, 447–452 (2018).
Del Mármol, J., Yedlin, M. A. & Ruta, V. The structural basis of odorant recognition in insect olfactory receptors. Nature https://doi.org/10.1038/s41586-021-03794-8 (2021).
McDonald, J. H. & Kreitman, M. Adaptive protein evolution at the adh locus in Drosophila. Nature 351, 652–654 (1991).
Smith, N. G. C. & Eyre-Walker, A. Adaptive protein evolution in Drosophila. Nature 415, 1022–1024 (2002).
Arguello, J. R. et al. Extensive local adaptation within the chemosensory system following Drosophila melanogaster’s global expansion. Nat. Commun. https://doi.org/10.1038/ncomms11855 (2016).
Mansourian, S. et al. Wild African Drosophila melanogaster are seasonal specialists on marula fruit. Curr. Biol. https://doi.org/10.1016/j.cub.2018.10.033 (2018).
Albalat, R. & Cañestro, C. Evolution by gene loss. Nat. Rev. Genet. 17, 379–391 (2016).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Hohenlohe, P. A., Phillips, P. C. & Cresko, W. A. Using population genomics to detect selection in natural populations: key concepts and methodological considerations. Int. J. Plant Sci. 171, 1059–1071 (2010).
Dobritsa, A. A., van der Goes van Naters, W., Warr, C. G., Steinbrecht, R. A. & Carlson, J. R. Integrating the molecular and cellular basis of odor coding in the Drosophila antenna. Neuron 37, 827–841 (2003).
Galizia, C. G., Münch, D., Strauch, M., Nissler, A. & Ma, S. Integrating heterogeneous odor response data into a common response model: a DoOR to the complete olfactome. Chem. Senses 35, 551–563 (2010).
Hallem, E. A. & Carlson, J. R. Coding of odors by a receptor repertoire. Cell 125, 143–160 (2006).
Hallem, E. A., Ho, M. G. & Carlson, J. R. The molecular basis of odor coding in the Drosophila antenna. Cell 117, 965–979 (2004).
Burchett, W. W., Ellis, A. R., Harrar, S. W. & Bathke, A. C. Nonparametric inference for multivariate data: the R package npmv. J. Stat. Softw. https://doi.org/10.18637/jss.v076.i04 (2017).
Khallaf, M. A. et al. Mate discrimination among subspecies through a conserved olfactory pathway. Sci. Adv. https://doi.org/10.1126/sciadv.aba5279 (2020).
Khallaf, M. A. et al. Large-scale characterization of sex pheromone communication systems in Drosophila. Nat. Commun. 12, 4165 (2021).
Stensmyr, M. C., Dekker, T. & Hansson, B. S. Evolution of the olfactory code in the Drosophila melanogaster subgroup. Proc. Biol. Sci. 270, 2333–2340 (2003).
Stensmyr, M. C. et al. A conserved dedicated olfactory circuit for detecting harmful microbes in Drosophila. Cell 151, 1345–1357 (2012).
Auer, T. O., Shahandeh, M. P. & Benton, R. Drosophila sechellia: a genetic model for behavioral evolution and neuroecology. Annu. Rev. Genet. https://doi.org/10.1146/annurev-genet-071719-020719 (2021).
Lachaise, D. et al. in Evolutionary Biology Vol. 22 (eds Hecht, M. K. et al.) Springer US. 159-225 (1988).
David, J. R., McEvey, S. F., Solignac, M. & Tsacas, L. Drosophila communities on Mauritius and the ecological niche of D. mauritiana (Diptera, Drosophilidae). Rev. Zool. Afr. J. Afr. Zool. 103, 107–116 (1989).
Wu, S. T. et al. Valence opponency in peripheral olfactory processing. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2120134119 (2022).
Hickner, P. V. et al. The making of a pest: insights from the evolution of chemosensory receptor families in a pestiferous and invasive fly, Drosophila suzukii. BMC Genomics 17, 648 (2016).
Monahan, K. & Lomvardas, S. Monoallelic expression of olfactory receptors. Annu. Rev. Cell Dev. Biol. 31, 721–740 (2015).
Rengarajan, S. & Hallem, E. A. Olfactory circuits and behaviors of nematodes. Curr. Opin. Neurobiol. 41, 136–148 (2016).
Dahanukar, A., Lei, Y.-T., Kwon, J. Y. & Carlson, J. R. Two Gr genes underlie sugar reception in Drosophila. Neuron 56, 503–516 (2007).
Dweck, H. K. M. & Carlson, J. R. Molecular logic and evolution of bitter taste in Drosophila. Curr. Biol. 30, e13 (2020).
Fujii, S. et al. Drosophila sugar receptors in sweet taste perception, olfaction, and internal nutrient sensing. Curr. Biol. 25, 621–627 (2015).
Ganguly, A. et al. A molecular and cellular context-dependent role for Ir76b in detection of amino acid taste. Cell Rep. 18, 737–750 (2017).
Jiao, Y., Moon, S. J., Wang, X., Ren, Q. & Montell, C. Gr64f is required in combination with other gustatory receptors for sugar detection in Drosophila. Curr. Biol. 18, 1797–1801 (2008).
Kwon, J. Y., Dahanukar, A., Weiss, L. A. & Carlson, J. R. Molecular and cellular organization of the taste system in the Drosophila larva. J. Neurosci. 31, 15300–15309 (2011).
Scott, K. Gustatory processing in Drosophila melanogaster. Annu. Rev. Entomol. 63, 15–30 (2018).
Slone, J., Daniels, J. & Amrein, H. Sugar receptors in Drosophila. Curr. Biol. 17, 1809–1816 (2007).
Tauber, J. M. et al. A subset of sweet-sensing neurons identified by IR56d are necessary and sufficient for fatty acid taste. PLoS Genet. 13, 1–18 (2017).
Wilson, R. I. Early olfactory processing in Drosophila: mechanisms and principles. Annu. Rev. Neurosci. 36, 217–241 (2013).
Jafari, S., Henriksson, J., Yan, H. & Alenius, M. Stress and odorant receptor feedback during a critical period after hatching regulates olfactory sensory neuron differentiation in Drosophila. PLoS Biol. 19, e3001101 (2021).
Maguire, S. E., Afify, A., Goff, L. A. & Potter, C. J. Odorant-receptor-mediated regulation of chemosensory gene expression in the malaria mosquito Anopheles gambiae. Cell Rep. 38, 110494 (2022).
Mika, K. et al. Olfactory receptor-dependent receptor repression in Drosophila. Sci. Adv. https://doi.org/10.1126/sciadv.abe3745 (2021).
Goldman, A. L., der Goes van Naters, W., Lessing, D., Warr, C. G. & Carlson, J. R. Coexpression of two functional odor receptors in one neuron. Neuron 45, 661–666 (2005).
Ray, A., van Naters, W. V. D. G. & Carlson, J. R. A regulatory code for neuron-specific odor receptor expression. PLoS Biol. 6, e125 (2008).
Ray, A., van Naters, W. V. D. G., Shiraiwa, T. & Carlson, J. R. Mechanisms of odor receptor gene choice in Drosophila. Neuron 53, 353–369 (2007).
Vosshall, L. B. & Stocker, R. F. Molecular architecture of smell and taste in Drosophila. Annu. Rev. Neurosci. 30, 505–533 (2007).
Aguadé, M. Nucleotide and copy-number polymorphism at the odorant receptor genes Or22a and Or22b in Drosophila melanogaster. Mol. Biol. Evol. 26, 61–70 (2009).
Lebreton, S. et al. A Drosophila female pheromone elicits species-specific long-range attraction via an olfactory channel with dual specificity for sex and food. BMC Biol. 15, 88 (2017).
de Bruyne, M., Smart, R., Zammit, E. & Warr Coral, G. Functional and molecular evolution of olfactory neurons and receptors for aliphatic esters across the Drosophila genus. J. Comp. Physiol. A 196, 97–109 (2010).
Fishilevich, E. & Vosshall, L. B. Genetic and functional subdivision of the Drosophila antennal lobe. Curr. Biol. 15, 1548–1553 (2005).
McLaughlin, C. N. et al. Single-cell transcriptomes of developing and adult olfactory receptor neurons in Drosophila. eLife https://doi.org/10.7554/eLife.63856 (2021).
Mika, K. & Benton, R. Olfactory receptor gene regulation in insects: multiple mechanisms for singular expression. Front. Neurosci. 15, 738088 (2021).
Koutroumpa, F. A. et al. Shifts in sensory neuron identity parallel differences in pheromone preference in the European corn borer. Front. Ecol. Evol. https://doi.org/10.3389/fevo.2014.00065 (2014).
Karner, T., Kellner, I., Schultze, A., Breer, H. & Krieger, J. Co-expression of six tightly clustered odorant receptor genes in the antenna of the malaria mosquito Anopheles gambiae. Front. Ecol. Evol. https://doi.org/10.3389/fevo.2015.00026 (2015).
Younger, M. A. et al. Non-canonical odor coding in the mosquito. Preprint at bioRxiv https://doi.org/10.1101/2020.11.07.368720 (2022).
Task, D. et al. Chemoreceptor co-expression in Drosophila melanogaster olfactory neurons. eLife https://doi.org/10.7554/eLife.72599 (2022).
Dippel, S. et al. Morphological and transcriptomic analysis of a beetle chemosensory system reveals a gnathal olfactory center. BMC Biol. 14, 90 (2016).
Vosshall, L. B., Wong, A. M. & Axel, R. An olfactory sensory map in the fly brain. Cell 102, 147–159 (2000).
Port, F. & Bullock, S. L. Augmenting CRISPR applications in Drosophila with tRNA-flanked sgRNAs. Nat. Methods 13, 852–854 (2016).
Port, F., Chen, H.-M., Lee, T. & Bullock, S. L. Optimized CRISPR/Cas tools for efficient germline and somatic genome engineering in Drosophila. Proc. Natl Acad. Sci. USA 111, E2967–E2976 (2014).
Gratz, S. J. et al. Highly specific and efficient CRISPR/Cas9-catalyzed homology-directed repair in Drosophila. Genetics 196, 961–971 (2014).
Kondo, S. et al. Neurochemical organization of the Drosophila brain visualized by endogenously tagged neurotransmitter receptors. Cell Rep. 30, 284–297.e285 (2020).
Bischof, J., Maeda, R. K., Hediger, M., Karch, F. & Basler, K. An optimized transgenesis system for Drosophila using germ-line-specific phiC31 integrases. Proc. Natl Acad. Sci. USA 104, 3312–3317 (2007).
Han, C., Jan, L. Y. & Jan, Y.-N. Enhancer-driven membrane markers for analysis of nonautonomous mechanisms reveal neuron–glia interactions in Drosophila. Proc. Natl Acad. Sci. USA 108, 9673–9678 (2011).
Gohl, D. M. et al. A versatile in vivo system for directed dissection of gene expression patterns. Nat. Methods 8, 231–237 (2011).
Benton, R. & Dahanukar, A. Electrophysiological recording from Drosophila olfactory sensilla. Cold Spring Harb. Protoc. https://doi.org/10.1101/pdb.prot5630 (2011).
Ebrahim, S. A. M. et al. Drosophila avoids parasitoids by sensing their semiochemicals via a dedicated olfactory circuit. PLoS Biol. 13, 1–18 (2015).
Lin, C.-C. & Potter, C. J. Re-classification of Drosophila melanogaster trichoid and intermediate sensilla using fluorescence-guided single sensillum recording. PLoS ONE 10, e0139675 (2015).
R Core Team R: A Language and Environment for Statistical Computing v.4.1.0 https://www.R-project.org/ (R Foundation for Statistical Computing, 2021).
Wickham, H. ggplot2: Elegant graphics for data analysis. R package v.3.3.0 (2016).
Ligges, U. & Mächler, M. Scatterplot3d—an R package for visualizing multivariate data. J. Stat. Softw. 8, 1–20 (2003).
Stekhoven, D. J. & Bühlmann, P. MissForest—non-parametric missing value imputation for mixed-type data. Bioinformatics 28, 112–118 (2012).
Saina, M. & Benton, R. Visualizing olfactory receptor expression and localization in Drosophila. Methods Mol. Biol. 1003, 211–228 (2013).
Silbering, A. F. et al. Complementary function and integrated wiring of the evolutionarily distinct Drosophila olfactory subsystems. J. Neurosci. 31, 13357–13375 (2011).
Sánchez-Alcañiz, J. A., Zappia, G., Marion-Poll, F. & Benton, R. A mechanosensory receptor required for food texture detection in Drosophila. Nat. Commun. 8, 14192 (2017).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Xu, B. & Yang, Z. PAMLX: a graphical user interface for PAML. Mol. Biol. Evol. 30, 2723–2724 (2013).
Signor, S. A., New, F. N. & Nuzhdin, S. A large panel of Drosophila simulans reveals an abundance of common variants. Genome Biol. Evol. 10, 189–206 (2018).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Grenier, J. K. et al. Global diversity lines—a five-continent reference panel of sequenced Drosophila melanogaster strains. G3 (Bethesda) https://doi.org/10.1534/g3.114.015883 (2015).
Pupko, T., Pe’er, I., Shamir, R. & Graur, D. A fast algorithm for joint reconstruction of ancestral amino acid sequences. Mol. Biol. Evol. 17, 890–896 (2000).
Chakraborty, M. et al. Evolution of genome structure in the Drosophila simulans species complex. Genome Res. 31, 380–396 (2021).
Miller, D. E., Staber, C., Zeitlinger, J. & Hawley, R. S. Highly contiguous genome assemblies of 15 Drosophila species generated using nanopore sequencing. G3 (Bethesda) 8, 3131–3141 (2018).
Smit, AFA, Hubley, R & Green, P. RepeatMasker Open-4.0.2013-2015. http://www.repeatmasker.org
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Bailey, T. L., Williams, N., Misleh, C. & Li, W. W. MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 34, W369–W373 (2006).
Acknowledgements
We thank M. Cardoso-Moreira, L.L. Prieto-Godino, J. A. Sánchez-Alcañiz, L. Keller, M. Long and members of the Arguello lab for comments on earlier versions of the manuscript; M. Khallaf, M. Knaden, A. Svatos and J. Weissflog at the Max Planck Institute for Chemical Ecology for the synthesized (R)-actinidine; and J. Carlson and D. Stern for sharing transgenic fly lines. T.O.A. was supported by a Human Frontier Science Program Long-Term Fellowship (no. LT000461/2015-L) and a Swiss National Science Foundation Ambizione Grant (no. PZ00P3 185743). Research in R.B.’s laboratory was supported by ERC Consolidator and Advanced Grants (nos 615094 and 833548, respectively) and the Swiss National Science Foundation. Research in J.R.A.’s lab is supported by the Swiss National Science Foundation grants no. PP00P3_176956 and no. 310030_201188.
Author information
Authors and Affiliations
Contributions
J.R.A. conceived the study. J.R.A., T.O.A. and R.A.-O. designed the experiments with input from R.B. J.R.A., T.O.A., R.A.-O. and S.C. carried out the experiments. J.R.A., T.O.A. and R.A.-O. analysed the data. J.R.A and T.O.A. prepared the original draft of the manuscript with revisions from R.A.-O. and R.B. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Jean-Christophe Billeter, Carolyn McBride and Coral Warr for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Microsynteny for chromosomal regions containing Or67a.D and Or67a.P.
Alignment for the chromosome 3L interval containing Or67a.D and Or67a.P (and flanking genes, light blue) for six species. High sequence identity is indicated with black alignment blocks with low sequence identity indicated with light grey alignment blocks. Thin horizontal lines are alignment gaps. Red annotations indicate locations of transposable elements. Chromosome position on the horizontal axis are relative to the extracted interval. See Supplementary Files 1 for the alignment in a flat file.
Extended Data Fig. 2 Microsynteny for chromosomal region containing Or67a.3R.
Alignment for the chromosome 3R interval containing Or67a.3R (and flanking genes, light blue) for six species. High sequence identity is indicated with black alignment blocks with low sequence identity indicated with light grey alignment blocks. Thin horizontal lines are alignment gaps. Red annotations indicate locations of transposable elements. Chromosome position on the horizontal axis are relative to the extracted interval. See Supplementary Files 2 for the alignment in a flat file.
Extended Data Fig. 3 Genetic diversity in regions containing D. melanogaster’s Or67a.D and Or67a.3R deletions.
Nucleotide diversity (Π) and Tajima’s D over D. melanogaster’s chromosomal regions containing the intact Or67a.P gene and the deleted Or67a.D and Or67a.3R. The regions containing the deleted Or67a paralogs do not show differences in genetic diversity in comparison to the surrounding regions, as would be expected if the deletions were adaptive and swept in the population. The black line in each panel is the smoothed curve fit with LOESS, with the grey ribbon around it displaying the standard error. The sample size (number of genomes) = 84.
Extended Data Fig. 4 Evolution of odour sensitivity across Or67a orthologs.
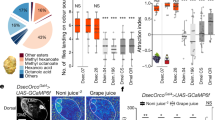
The full set of dose-response experiments for the subset of odours that evoked high or intermediate responses in our initial screen of nine odours (Fig. 2b). For simplicity, the level of significance indicated above each concentration’s comparison is only for the single comparison with the largest difference (see Supplementary Table 4 for the full set of tests; *p < 0.05, **p < 0.01, ***p < 0.001). Colours correspond to those in Fig. 2d. p-values were calculated with a two-sided Dunn test; correction for multiple comparisons was done using the Holm method. For sample sizes and all test results see Supplementary Table 4.
Extended Data Fig. 5 Transgenic tools for Or67a expression analyses in D. simulans and D. melanogaster.
a, Schematics of the wild-type and knock-in Gal4 transcriptional reporter alleles at the DsimOr67a.3R (top) and DsimOr67a.D/P (bottom) loci. The first was created via a two-step process (CRISPR/Cas9 engineering + PhiC31 mediated integration) while the latter was resulting from a direct CRISPR/Cas9 mediated insertion. The red lines indicate sgRNA cutting sites; three and two sgRNAs were used to target the Or67a.3R and Or67a.P locus, respectively. Note that the intercalated sequence was removed upon donor vector integration. b, Antennal co-expression of the melOr67a.P-GFP transcriptional reporter and Or67a.P RNA in D. melanogaster. Scale bar = 25 μm. c, Antennal co-expression of the simOr67a.P-GFP and simOr67a.D-RFP transcriptional reporters (top) and the simOr67a.3R-GFP and simOr67a.D-RFP transcriptional reporters (bottom) in D. melanogaster. Scale bar = 25 μm. d, Pairing of the simOr67a.P-GFP promoter transcriptional reporter and melOr85f-Gal4, UAS-RFP expression in neighboring neurons in the antenna of D. melanogaster. Scale bar = 25 μm. Inset scale bar = 5 μm. (b-d) Experiments were repeated at least three times for each staining on independent days and pictures show representative examples for each condition.
Supplementary information
Supplementary Tables
Supplementary Table 1. Codeml table of models tested and likelihoods. Supplementary Table 2. Codeml table of likelihood ratio tests. Supplementary Table 3. Relative effects of odours on receptor responses based on the npmv analysis. Supplementary Table 4. Dose–response test results and sample sizes. Supplementary Table 5. Pairwise identity between Or67a gene sequences. Supplementary Table 6. Pairwise identity between Or67a promoter sequences. Supplementary Table 7. Table with MEME results. Supplementary Table 8. Oligonucleotides used in this study.
Supplementary Data 1
Alignment for the 3L region containing Or67a.D and Or67a.P.
Supplementary Data 2
Alignment for the 3R region containing Or67a.3R.
Supplementary Data 3
D. simulans and D. melanogaster polymorphism summaries.
Supplementary Data 4
Odour response data from electrophysiology experiments: 10−2.
Supplementary Data 5
Odour response data from electrophysiology experiments: dose–responses.
Supplementary Data 6
Odour response data from electrophysiology experiments: fluorescence-guided single-sensillum recordings.
Supplementary Data 7
D. simulans Sanger sequences generated in this study.
Supplementary Data 8
D. simulans Sanger sequences generated in this study.
Supplementary Data 9
D. simulans Sanger sequences generated in this study.
Rights and permissions
About this article
Cite this article
Auer, T.O., Álvarez-Ocaña, R., Cruchet, S. et al. Copy number changes in co-expressed odorant receptor genes enable selection for sensory differences in drosophilid species. Nat Ecol Evol 6, 1343–1353 (2022). https://doi.org/10.1038/s41559-022-01830-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-022-01830-y