Abstract
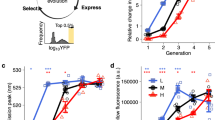
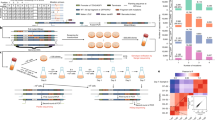
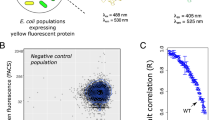
Protein abundance affects the evolution of protein genotypes, but we do not know how it affects the evolution of protein phenotypes. Here we investigate the role of protein abundance in the evolvability of green fluorescent protein (GFP) towards the novel phenotype of cyan fluorescence. We evolve GFP in E. coli through multiple cycles of mutation and selection and show that low GFP expression facilitates the evolution of cyan fluorescence. A computational model whose predictions we test experimentally helps explain why: lowly expressed proteins are under stronger selection for proper folding, which facilitates their evolvability on short evolutionary time scales. The reason is that high fluorescence can be achieved by either few proteins that fold well or by many proteins that fold less well. In other words, we observe a synergy between a protein’s scarcity and its stability. Because many proteins meet the essential requirements for this scarcity–stability synergy, it may be a widespread mechanism by which low expression helps proteins evolve new phenotypes and functions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data are available in the manuscript or the supplementary materials. SMRT sequencing data are available at the National Center for Biotechnology Information with a BioProject ID PRJNA833567 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA833567).
Code availability
Custom code used in this study is available in a public GitHub repository (https://github.com/dasmeh/Discrete_Time_Markov_Chain_Evolution).
References
Newman, J. R. et al. Single-cell proteomic analysis of S. cerevisiae reveals the architecture of biological noise. Nature 441, 840–846 (2006).
Silander, O. K. et al. A genome-wide analysis of promoter-mediated phenotypic noise in Escherichia coli. PLoS Genet. 8, e1002443 (2012).
Lehner, B. Selection to minimise noise in living systems and its implications for the evolution of gene expression. Mol. Syst. Biol. 4, 170 (2008).
Pál, C., Papp, B. & Lercher, M. J. An integrated view of protein evolution. Nat. Rev. Genet. 7, 337–348 (2006).
Bershtein, S., Goldin, K. & Tawfik, D. S. Intense neutral drifts yield robust and evolvable consensus proteins. J. Mol. Biol. 379, 1029–1044 (2008).
Socha, R. D., Chen, J. & Tokuriki, N. The molecular mechanisms underlying hidden phenotypic variation among metallo-β-lactamases. J. Mol. Biol. 431, 1172–1185 (2019).
Dasmeh, P. & Serohijos, A. W. Estimating the contribution of folding stability to nonspecific epistasis in protein evolution. Proteins 86, 1242–1250 (2018).
Bershtein, S. et al. Protein homeostasis imposes a barrier on functional integration of horizontally transferred genes in bacteria. PLoS Genet. 11, e1005612 (2015).
Laurent, J. M. et al. Protein abundances are more conserved than mRNA abundances across diverse taxa. Proteomics 10, 4209–4212 (2010).
Zhang, J. & Yang, J.-R. Determinants of the rate of protein sequence evolution. Nat. Rev. Genet. 16, 409–420 (2015).
Stefani, M. & Dobson, C. M. Protein aggregation and aggregate toxicity: new insights into protein folding, misfolding diseases and biological evolution. J. Mol. Med. 81, 678–699 (2003).
Yang, J. R., Zhuang, S. M. & Zhang, J. Impact of translational error‐induced and error‐free misfolding on the rate of protein evolution. Mol. Syst. Biol. 6, 421 (2010).
Moutinho, A. F., Trancoso, F. F. & Dutheil, J. Y. The impact of protein architecture on adaptive evolution. Mol. Biol. Evol. 36, 2013–2028 (2019).
Moutinho, A. F., Bataillon, T. & Dutheil, J. Y. Variation of the adaptive substitution rate between species and within genomes. Evol. Ecol. 34, 315–338 (2020).
Yip, S. H.-C. & Matsumura, I. Substrate ambiguous enzymes within the Escherichia coli proteome offer different evolutionary solutions to the same problem. Mol. Biol. Evol. 30, 2001–2012 (2013).
Larion, M., Moore, L. B., Thompson, S. M. & Miller, B. G. Divergent evolution of function in the ROK sugar kinase superfamily: role of enzyme loops in substrate specificity. Biochemistry 46, 13564–13572 (2007).
Serohijos, A. W., Rimas, Z. & Shakhnovich, E. I. Protein biophysics explains why highly abundant proteins evolve slowly. Cell Rep. 2, 249–256 (2012).
Leuenberger, P. et al. Cell-wide analysis of protein thermal unfolding reveals determinants of thermostability. Science 355, eaai7825 (2017).
Serohijos, A. W., Lee, S. R. & Shakhnovich, E. I. Highly abundant proteins favor more stable 3D structures in yeast. Biophys. J. 104, L1–L3 (2013).
Bloom, J. D., Labthavikul, S. T., Otey, C. R. & Arnold, F. H. Protein stability promotes evolvability. Proc. Natl. Acad. Sci. USA 103, 5869–5874 (2006).
Zheng, J., Guo, N. & Wagner, A. Selection enhances protein evolvability by increasing mutational robustness and foldability. Science 370, eabb5962 (2020).
Tokuriki, N., Stricher, F., Serrano, L. & Tawfik, D. S. How protein stability and new functions trade off. PLoS Comput. Biol. 4, e1000002 (2008).
Keseler, I. M. et al. The EcoCyc database: reflecting new knowledge about Escherichia coli K-12. Nucleic Acids Res. 45, D543–D550 (2017).
Zaslaver, A. et al. A comprehensive library of fluorescent transcriptional reporters for Escherichia coli. Nat. Methods 3, 623–628 (2006).
Wu, Z. et al. Expression level is a major modifier of the fitness landscape of a protein coding gene. Nat. Ecol. Evol. 6, 103–115 (2022).
Pakula, A. A. & Sauer, R. T. Genetic analysis of protein stability and function. Annu. Rev. Genet. 23, 289–310 (1989).
Sarkisyan, K. S. et al. Local fitness landscape of the green fluorescent protein. Nature 533, 397–401 (2016).
Mitchell, R. L. Permanence of the log-normal distribution. J. Opt. Soc. Am. 58, 1267–1272 (1968).
Tokuriki, N. & Tawfik, D. S. Stability effects of mutations and protein evolvability. Curr. Opin. Struct. Biol. 19, 596–604 (2009).
Zheng, J., Payne, J. L. & Wagner, A. Cryptic genetic variation accelerates evolution by opening access to diverse adaptive peaks. Science 365, 347–353 (2019).
Crameri, A., Whitehorn, E. A., Tate, E. & Stemmer, W. P. Improved green fluorescent protein by molecular evolution using DNA shuffling. Nat. Biotechnol. 14, 315–319 (1996).
Fukuda, H., Arai, M. & Kuwajima, K. Folding of green fluorescent protein and the cycle3 mutant. Biochemistry 39, 12025–12032 (2000).
Nam, S. H., Oh, K. H., Kim, G. J. & Kim, H. S. Functional tuning of a salvaged green fluorescent protein variant with a new sequence space by directed evolution. Protein Eng. 16, 1099–1105 (2003).
Heim, R. & Tsien, R. Y. Engineering green fluorescent protein for improved brightness, longer wavelengths and fluorescence resonance energy transfer. Curr. Biol. 6, 178–182 (1996).
Drummond, D. A., Raval, A. & Wilke, C. O. A single determinant dominates the rate of yeast protein evolution. Mol. Biol. Evol. 23, 327–337 (2006).
Plotkin, J. B. & Fraser, H. B. Assessing the determinants of evolutionary rates in the presence of noise. Mol. Biol. Evol. 24, 1113–1121 (2007).
Maddamsetti, R. Universal constraints on protein evolution in the long-term evolution experiment with Escherichia coli. Genome Biol. Evol. https://doi.org/10.1093/gbe/evab070 (2021).
LaBar, T. & Adami, C. Evolution of drift robustness in small populations. Nat. Commun. 8, 1012 (2017).
Raser, J. M. & O’shea, E. K. Noise in gene expression: origins, consequences, and control. Science 309, 2010–2013 (2005).
Hardin, J. & Wilson, J. A note on oligonucleotide expression values not being normally distributed. Biostatistics 10, 446–450 (2009).
Ham, L., Brackston, R. D. & Stumpf, M. P. Extrinsic noise and heavy-tailed laws in gene expression. Phys. Rev. Lett. 124, 108101 (2020).
Furusawa, C., Suzuki, T., Kashiwagi, A., Yomo, T. & Kaneko, K. Ubiquity of log-normal distributions in intra-cellular reaction dynamics. Biophysics 1, 25–31 (2005).
Casellas, J. & Varona, L. Modeling skewness in human transcriptomes. PLoS ONE 7, e38919 (2012).
Bengtsson, M., Ståhlberg, A., Rorsman, P. & Kubista, M. Gene expression profiling in single cells from the pancreatic islets of Langerhans reveals lognormal distribution of mRNA levels. Genome Res. 15, 1388–1392 (2005).
Bódi, Z. et al. Phenotypic heterogeneity promotes adaptive evolution. PLoS Biol. 15, e2000644 (2017).
Zhang, Z., Qian, W. & Zhang, J. Positive selection for elevated gene expression noise in yeast. Mol. Syst. Biol. 5, 299 (2009).
Acar, M., Mettetal, J. T., & Van Oudenaarden, A. Stochastic switching as a survival strategy in fluctuating environments. Nat. Genet. 40, 471–475 (2008).
Kacser, H., Burns, J. A., Kacser, H. & Fell, D. The control of flux. Biochem. Soc. Trans. 23, 341–366 (1995).
Chen, P. & Shakhnovich, E. I. Lethal mutagenesis in viruses and bacteria. Genetics 183, 639–650 (2009).
Soskine, M. & Tawfik, D. S. Mutational effects and the evolution of new protein functions. Nat. Rev. Genet. 11, 572–582 (2010).
Hoekstra, H. E. et al. Strength and tempo of directional selection in the wild. Proc. Natl. Acad. Sci. USA 98, 9157–9160 (2001).
Oz, T. et al. Strength of selection pressure is an important parameter contributing to the complexity of antibiotic resistance evolution. Mol. Biol. Evol. 31, 2387–2401 (2014).
Jahn, L. J., Munck, C., Ellabaan, M. M. H. & Sommer, M. O. A. Adaptive laboratory evolution of antibiotic resistance using different selection regimes lead to similar phenotypes and genotypes. Front. Microbiol. 8, 816–816 (2017).
Zimmer, M. Green fluorescent protein (GFP): applications, structure, and related photophysical behavior. Chem. Rev. 102, 759–782 (2002).
Cormack, B. P., Valdivia, R. H. & Falkow, S. FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173, 33–38 (1996).
Warren, D. J. Preparation of highly efficient electrocompetent Escherichia coli using glycerol/mannitol density step centrifugation. Anal. Biochem. 413, 206–207 (2011).
Khersonsky, O. & Tawfik, D. S. Enzyme promiscuity: a mechanistic and evolutionary perspective. Annu. Rev. Biochem. 79, 471–505 (2010).
Aharoni, A. et al. The ‘evolvability’ of promiscuous protein functions. Nat. Genet. 37, 73–76 (2005).
Rhoads, A. & Au, K. F. PacBio sequencing and its applications. Genom. Proteom. Bioinform. 13, 278–289 (2015).
Chaisson, M. J. & Tesler, G. Mapping single molecule sequencing reads using basic local alignment with successive refinement (BLASR): application and theory. BMC Bioinform. 13, 238 (2012).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547 (2018).
Acknowledgements
This project has received funding from the European Research Council under grant agreement number 739874. We would also like to acknowledge support by Swiss National Science Foundation grant 31003A_172887, by the University Priority Research Program in Evolutionary Biology and by the flow cytometry facility and the functional genomics centre at the University of Zurich. S.K. thanks B.R. Iyengar and M. Olombrada for discussions and support.
Author information
Authors and Affiliations
Contributions
S.K., P.D. and A.W. were involved in the conceptualization of the study. S.K. and P.D. performed the experiments. P.D. performed computational modelling and formulated theoretical predictions. J.Z. provided the resources and training for working with fluorescent proteins. S.K., P.D., J.Z. and A.W. contributed to data analysis and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Ecology & Evolution thanks Jian-Rong (Philip) Yang, Rohan Maddamsetti and Claus Wilke for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–14, Tables 1–5 and information for the theoretical analysis and simulations of GFP evolution.
Rights and permissions
About this article
Cite this article
Karve, S., Dasmeh, P., Zheng, J. et al. Low protein expression enhances phenotypic evolvability by intensifying selection on folding stability. Nat Ecol Evol 6, 1155–1164 (2022). https://doi.org/10.1038/s41559-022-01797-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-022-01797-w
This article is cited by
-
Rapid evolutionary change in trait correlations of single proteins
Nature Communications (2024)