Abstract
Although migrations are essential for soil microorganisms to exploit scarce and heterogeneously distributed resources, bacterial mobility in soil remains poorly studied due to experimental limitations. In this study, time-lapse images collected using live microscopy techniques captured collective and coordinated groups of B. subtilis cells exhibiting “crowd movement”. Groups of B. subtilis cells moved through transparent soil (nafion polymer with particle size resembling sand) toward plant roots and re-arranged dynamically around root tips in the form of elongating and retracting “flocks” resembling collective behaviour usually associated with higher organisms (e.g., bird flocks or fish schools). Genetic analysis reveals B. subtilis flocks are likely driven by the diffusion of extracellular signalling molecules (e.g., chemotaxis, quorum sensing) and may be impacted by the physical obstacles and hydrodynamics encountered in the soil like environment. Our findings advance understanding of bacterial migration through soil matrices and expand known behaviours for coordinated bacterial movement.
Similar content being viewed by others
From a microbial perspective, soil is a nutritionally poor environment. Essential nutrients and energy sources exhibit heterogeneous spatial and temporal distributions within the complex soil matrix, leading to transient oases of microbial activity (“hot spots” and “hot moments”; [1]). The ability of microorganisms to move and exploit rare and distant sources of nutrients, such as plant root exudates, is therefore important for the maintenance of microbial diversity and soil ecosystem functioning. Rhizosphere communities, in turn, are key drivers of terrestrial nutrient cycling, aiding in plant nutrient uptake and contributing to the maintenance of general soil health. They also play an important role in plant health and defence against pathogens [2].
The mobility of bacteria in soil is considered dependent on several life strategies, including sensing specific compounds released by the root and migrating towards them by means of chemotaxis [3, 4]. Different forms of movement are possible, such as swimming in water films or twitching and swarming over solid obstacles [5]. Microorganisms likely need to coordinate and act as a group, using quorum sensing to navigate the complex soil matrix [6, 7] and attach to the root surface to form biofilm [8, 9]. Current methods are inherently limited in their ability to track bacterial populations in soil [10], partially due to their microscopic size but also due to the opacity of the soil environment. Even with recent advances in molecular and imaging techniques (many of them destructive), soil bacteria have proved particularly elusive organisms and our understanding of how they migrate towards nutrient sources in soil is near inexistent. Most work on bacterial motility, though extensive, is still performed predominantly in liquid cultures [3, 11], various hydroponic systems [12, 13] and semi-solid synthetic surfaces [14]. We are still unable to determine the distance microorganisms can migrate through soils and the mechanisms they employ to navigate this heterogenous environment, both passively and actively.
Here we have advanced live imaging capabilities by capturing and analysing the movement of fluorescently labelled Bacillus subtilis strains around roots growing in a transparent ionomer (i.e., synthetic polymer with ionic properties which has a refractive index close to water) soil matrix. As a well-studied model organism, B. subtilis is known for its plant-root interactions, with some members of the genus used as biocontrol agents [15, 16]. In our system, time-lapse images were collected using a custom-built light sheet microscope, allowing analyses of 3D live datasets of root-microbe interactions. The study revealed an unexpected new form of coordinated movement used by B. subtilis to navigate the soil matrix and interact with the surfaces of plant roots.
Results
Whole plant-environment microscopy reveals coordinated movement of B. subtilis in soil
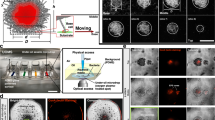
The custom-built experimental system simultaneously recorded the movement of B. subtilis cells, the topography of the soil-like matrix, and living plant roots at 30 min intervals over a 20 h period. The model system comprised four-day-old seedlings grown in transparent soil (Nafion), inoculated with a fluorescently labelled derivative of B. subtilis strain NCIB 3610, and images captured on a tailor-made light sheet microscope. To mimic the conditions for recruitment of bacteria from the surrounding soil environment, B. subtilis cells were inoculated locally using an inoculation patch infiltrated with approximately 2 million colony forming units (CFU), placed just beneath the soil surface level with the emerging root (Fig. 1A). This concentration of bacterial cells ensures an excess number of viable cells to colonise up to 0.2 g of root at an average concentration of 107 CFU g−1 likely encountered for B. subtilis [7] while at the lower end of bacterial concentrations found in agricultural soils [17]. Formation of filamentous flocks of bacteria occurred consistently within samples, converging systematically at the tip of the root (Fig. 1B). Filamentous flocks formed around the root tip and rearranged rapidly in the 20 h timeframe, elongating, retracting, and branching, suggesting a high level of coordination (Fig. 1C, Video S4). Because of the apparent sophistication of the coordination required to produce such flocking behaviour, akin to murmuration in birds or shoaling in fish, we have termed this form of bacterial mobility “crowd movement”.
A We have created an experimental system for studying bacterial movement towards plant roots, in soil. The experimental system consists of B. subtilis moving from an inoculation patch (yellow square) through the pore space between soil particles (grey) towards the plant root (brown). The system enabled us to track the movements of B. subtilis populations and revealed complex coordinated movement of bacteria through the pore structure. Note that diagram is not to scale. B B. subtilis (GFP labelled) was observed forming flocks using a form of coordinated mobility termed “crowd movement”. C Time lapse images (30 min intervals) show the distribution of bacteria (GFP labelled B. subtilis, green) in soil (Nafion, magenta) taking on the form of filamentous flocks that dynamically rearrange around the tomato root (red). Light scattering signal from air bubbles may be visible as seen to the left (middle and bottom) of the time lapse images (2 red spheres).
Crowd movement may be a mechanism to enhance colonisation of hosts
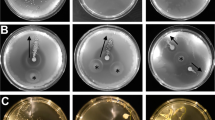
To study the functions of crowd movement, we investigated whether the phenomenon was plant-species specific. Similar patterns of B. subtilis movement around the roots of several plant hosts, including eudicotyledons such as tomato (Solanum lycopersicum var. Money maker) and lettuce (Lactuca sativa var. All Year Round), as well as monocotyledons such as winter wheat (Triticum aestivum var. filon) and Timothy grass (Phleum pratense) (Fig. 2A–D), were observed. Though many factors including bacterial preparation and several environmental factors are likely to affect the flocking behaviour, plant species and growth vigour has been observed as a deciding factor in flocking occurrence. All plant species tested showed flocking but varied in the likelihood of occurrence. Lettuce plants showed the most variable flocking response with about 25% of plants exhibiting flocks. Tomato plantlets showed a higher flocking success rate with about half of the plants, showing a degree of flock formation. The highest flock occurrence was observed with wheat plantlets, which show a near 100% occurrence of flocks within the time analysed. Only healthy plants were used for analysis, and natural variation in root elongation rate was statistically non-significant (Fig. S2).
Bacterial crowd movement has been captured around roots of several different plants, including (A) tomato (Solanum lycopersicum), (B) lettuce (Lactuca sativa), (C) timothy grass (Phleum pratense) and (D) wheat (Triticum aestivum). E Bacterial flocks form in the soil matrix and interact with the root tip (attaching/ detaching).
The interactions between B. subtilis flocks and the root tip were also visualised from time lapse image data under higher magnification. B. subtilis flocks interacted dynamically with the root surface preceding permanent biofilm formation on the root epidermis. At times, interactions led to temporary attachment followed by dispersion and formation of biofilm at other locations on the root surface (Fig. 2E). Both physiological states, bacteria forming biofilm, and dynamically re-arranging bacterial flocks could be present within a system at the same time. We hypothesise that B. subtilis cells use crowd movement to navigate the soil matrix, presumably in response to chemo-attractants which leads to subsequent dynamic interaction with the root tip (attachment and detachment) and may precede and/or facilitate root colonisation.
Morphodynamics of crowd movement
To understand when and how B. subtilis displays crowd movement, we studied how the shape of the flock dynamically forms and retracts (i.e., “morphodynamics”). We first defined a bacterial flock as a group of bacteria which appear to be connected and moving in a coordinated fashion. Groups of bacteria resulted in filamentous shapes which could be made up of a single branch or several branches (Fig. 3A). For quantitative analysis, single branches were traced on time-lapse image data and the morphodynamics characterised (Fig. 3B). We captured filamentous flocks appearing as early as 20 h post inoculation, reaching peak activity between 25 and 40 h, and then reducing substantially (Fig. 3C). The number of branches per flock varied over time and among samples, with an observed range between one and five (Fig. 3D).
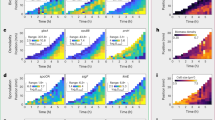
A A bacterial flock was defined as a group of bacteria which visually appear to be connected and moving dynamically in a coordinated fashion. Flocks of bacteria were filamentous and could occasionally define multiple branches. Several distinct, individual flocks could appear at the same time which do not appear to be connected to each other. The figure shows 3 individual flocks, represented by different colours. Two of the flocks have multiple branches (yellow and red trace) and one has only a single branch (blue trace). B Individual flock branches were traced to determine length, mean width and bacterial density over time with the first image in which flock was detected labelled as flock time zero (0 min). C Total combined length of individual flocks and (D) total number of flocks were measured around several tomato roots over time. E Length (left) and bacterial density (right) of individual flocks over time with different colours representing individual flocks. F Trajectories in the normalised length-density phase diagrams. Normalisations consisted of a centring step to place the centre of rotation of the trajectory, and a scaling step for magnitude of variations to be contained between −1 and 1 around the centre of rotation. To avoid overlapping of trajectories, individual curves were distributed along the x-axis. Each trajectory indicates the formation and disappearance of a single flock which could occasionally occur once (top panel) or multiple times (bottom panel). The graphs (C–F) show data collected from 1 to 3 flock branches traced from 8 individual and independent live systems.
We tracked various morphological characteristics of individual flock branches including length, elongation rate, width, and bacterial cell density. A single filamentous flock consisted of up to 107 cells and persisted for between 5 and 10 h, initially elongating, reaching a maximum length, then retracting and sometimes re-growing (Fig. 3E left). Similarly, the B. subtilis cell density of each filamentous flock increased initially, a peak density of approximately 109–1010 CFU per ml, and then decreased, sometimes repeating this process 2–3 times (Fig. 3E right).
The dynamics of filamentous flocks studied in the length-intensity phase space indicated the mechanism of formation likely results from an unstable feedback mechanism regulated by cell density (Fig. 3F). Initially, flock branches were seen elongating at low B. subtilis cell density. The increase in the length of the flock branch was then followed by an increase in density within the flock. As the B. subtilis cell density increased, the elongation rate increased, which in turn led to a decrease in B. subtilis cell density and subsequent retraction of the flock branch (Fig. 3F top). This cycle can be repeated multiple times for a single flock branch (Fig. 3F bottom).
Due to variation in the number of bacteria added to the system at the setup stage, as well as the population growth occurring within the system [18], we hypothesise that the number of cells within our system may vary considerably between samples at the time of flock occurrence. We suspect that cell density may influence the time of occurrence of the flocks but does not appear to impact the flock morphodynamics.
Crowd movement emerges from biophysical interactions in pore space
To better understand the mechanisms leading to the formation of bacterial flocks, we examined where the flocks formed in the pore space (Fig. 4A). A closer look at the soil matrix (Fig. 4B, top) revealed that B. subtilis flocks formed within the pore space (Fig. 4B, bottom) rather than on the surface of soil particles. This suggested a potential role for biophysical forces in the formation and regulation of these bacterial flocks.
A 3D image showing arrangement of bacteria in the soil matrix around the root. B Bacteria signal (green) and soil particle signal (white) from light sheet dataset showing the distribution of bacteria in the pore space rather than on particle surfaces. C Light sheet images showing highly branched flocks in high viscosity solutions (Percoll) and a single, more diffuse flock in low viscosity solution (water). D Viscosity of the solution used to saturate the soil matrix impacts the number of flock branches formed and the flock branch width but not the bacterial concentration (left to right). Barplots show the average values ± 2 SD for individual root samples (left) or for individual flocks (middle and right) for different diffusion coefficients. Each viscosity level data was collected from 5 separate live systems. Bars which share a letter are not significantly different from each other. E The phase diagram obtained in low viscosity soil solution showed a linear relationship between bacterial density characteristic of stable flock formation.
We investigated whether soil solution viscosity influenced the formation of bacterial flocks. Liquid density was adjusted by altering the composition of the colloidal suspension used as soil solution, which revealed a density-dependent influence on flock morphology (Fig. 4C). Increased viscosity (i.e., 78% Percoll, 6.6 mPa s−1, video S5) resulted in flocks that were significantly more branched (p = 0.027, Fig. 4D left) compared to those in water (1 mPa s−1, video S7). The flocks in the intermediate viscosity solution (39% Percoll, 2.2 mPa s−1, video S6) was not significantly different from those in either the high viscosity or water solution. Viscosity was indirectly related to flock branch width with decreasing viscosity resulting in increasing flock branch width from 0.33 ± 0.07 mm at high viscosity to 0.48 ± 0.15 mm at low viscosity (p = 0.031, Fig. 4D middle). Viscosity however, had no significant effect on the B. subtilis cell density of the flock (p = 0.251, Fig. 4D right). Additionally, in contrast to previous observations in high viscosity solution (Fig. 3F), filamentous flocks forming in low viscosity solution revealed linear trajectories in the length-intensity phase space (Fig. 4E). The elongation rate of the filamentous flock progressed steadily, slowing down gradually towards the final flock branch length.
We found that the theoretical doubling time calculated from flock cell density, though variable, could be as low as 50 min around the inflection point of bacterial flocks. To address whether flock development was a factor of bacterial proliferation, the growth rate / cellular doubling time of B. subtilis was estimated from in vitro growth at 20 °C in a defined medium (MSgg) as approximately 8 h. As the medium represents a more carbon-rich source than the microcosm, this implies that flock development occurred from accumulation of existing cells rather than generation of new cells.
Mutations in genes controlling quorum sensing and mobility affect the morphology of flocks
We investigated the genetic mechanisms that may be required for flocks to form. Experiments were performed with fluorescently labelled strains of B. subtilis NCIB 3610 containing deletions of genes that encode the surface factors involved in movement and biofilm formation: flagella (hag) and the biofilm matrix exopolysaccharide (epsA-O) respectively. Additionally, the impact of removing the secreted biosurfactant surfactin (srfAA), as well as the DNA-binding response regulator ComA (comA), which forms part of the ComQXP quorum sensing system, were evaluated with live plant roots growing in transparent soil. For reference purposes, the colony biofilm architecture is shown for all the strains used in the analysis (Fig. S3).
The results obtained by imaging the movement of wild type and mutant strains revealed that crowd movement is a complex multicellular behaviour involving more than one mechanism. Observations derived from at least 8 replicates for each strain were divided into one of 3 categories: clearly defined, branching, filamentous flocks (Fig. 5A), a form of diffuse group movement resembling bacterial clouds or fronts with a distinct absence of branching (Fig. 5B) and “no crowd movement observed”. Our results show that there is no significant difference in flocking behaviour between either “wild-type” strain used to generate the mutants (compare WT1 (NRS 1473) with WT2 (NRS 5634), p = 0.897, video S8). Additionally, there is no statistically significant role for surfactin in flock formation (p = 0.057, video S9). However, when epsA-O (video S10) or comA (video S11) were deleted, a significant loss of co-ordination in flocks was occurred (p < 0.05). Moreover, in the absence of the flagellar filament (hag deletion strain, video S12), a complete absence of any form of crowd movement was noted (p < 0.05). Even in a dual inoculation experiment, the GFP labelled hag mutant (green) did not form flocks while the mKate2 labelled WT (magenta) did, suggesting conditions for crowd movement were met in the system and confirming active flagellar motion as a requirement (video S13) with no passive motility gained by non-motile cells within motile communities. Any biofilms on the growing root would likely be due to bacteria arriving at the root surface through passive dispersion and other non-flagellar driven bacterial movement.
Flock formation was evaluated in surfactin (srfA) mutant (NRS 6963), exopolysaccharide (EPS) mutant (NRS 3798), quorum sensing (comA) mutant (NRS 7387), and flagellar (hag) mutant (NRS 6959) around wheat roots. The flock formation was evaluated for WT2 which carries a comI mutation strain (NRS 5634) as the parental strain of the quorum sensing (comA) mutant. Results were divided into 2 morphological categories; (A) clearly defined, branched filamentous flocks which interact with the root or (B) diffuse fronts or clouds of bacteria with no detectable structure which move through the soil but may not necessarily interact with the root. C The percent occurrence of each of the categories were recorded for several plant-soil-strain systems (minimum of n = 8 for each strain). Selection of category was performed by way of a blind test by 3 individuals. Stars indicate mutant strains which show significantly different flocking behaviour from its corresponding parental strain at p < 0.05.
Mathematical models indicate crowd movement may be mediated by diffusion of extracellular signals
We designed a mathematical model to study whether the changes in flocking patterns observed in response to the soil solution are due to changes in the diffusivity of both bacteria and quorum sensing signal because the viscosity of a liquid is inversely proportional to diffusivity in a medium (Stokes-Einstein equation, [19]). The model predicted the near linear decrease in number of flock branches observed with the increase of the diffusion coefficients due to the change of soil solution (Fig. 6A). In contrast, the model did not completely predict the response observed in the width of flocks. Although the model correctly predicted the increase in flock branch width with the increase in diffusion coefficients, it could not predict the strong non-linearity of the response observed experimentally. This result suggests the existence of dependencies of the quorum sensing mechanism to environmental factors, for example mechanosensing of the physical properties of the liquid solution or complex dependency to cell density.
A Model predictions, showing the correlation between diffusion and the number of flock branches (blue, left y-axis) as well as the correlation between diffusion and the width of flock branch (red, right y-axis). Parameters of the model include the interaction radius of the bacteria (R = 0.5 mm), the diffusion coefficient (D = 0.005 mm2 s−1) and time (T = 10 s). Image inserts show from left to right the visual representation of the impact of increasing diffusion on flock branches as shown in the graph. B Image series depict the impact of the soil porosity on the formation of filamentous flocks in solutions. Soil porosity is controlled with an increasing number of regularly distributed obstacles with identical shape and size (from left to right), which increase the number of flock branches. C Image series showing the effect of interaction strength between bacteria (quorum sensing) on flock formation from high to low (left to right, K = 0.1, 0.7, and 1.0 respectively). Colour scale of flocks represents bacterial cell density, ranging from 20 (blue) to 100 (yellow) cells per mm2. Bacteria below the threshold are not plotted (white) and any values above 100 remain yellow.
We performed an additional simulation to assess factors that contribute to the morpho-dynamics of bacterial flocks. Soil heterogeneity was found to be important. The model predicted that physical obstacles affect the morphology of the bacterial flocks, with shorter distances between obstacles leading to more branched and more defined filamentous flocks, with more variation in cell density (Fig. 6B). The strength of the bacterial response to the quorum sensing signal (parameter K in Eq. (3)) controls the continuity of the bacterial flock, and the ability of the flock to maintain contact with the plant root. High values of the interaction strength parameter K were needed to generate long and continuous filamentous flocks. However, excessive values of interaction strength led to disconnected and eventually immobile islands of cells (Fig. 6C).
Discussion
We report on the collective and seemingly co-ordinated movement of bacteria in a soil-like matrix during the colonisation of living plant roots. This coordinated group response, referred to as crowd movement, resembles behaviour normally associated with higher organisms such as migrating herds [20, 21], flocking birds [22, 23], schools of fish [24] and swarms of insects [25, 26]. Many models have been created to explain the dynamics of this group behaviour using theories of attraction, short range repulsion, and alignment of velocities [22]. However, often the sets of rules used in these models are based on assumptions due to lack of experimental data [27]. With the advent of GPS tracing, advances have been made in similar research on higher organisms [28], but due to the size of microorganisms and the complex nature of the soil matrix, obtaining experimental data is difficult.
Evidence of the co-ordinated activity of microorganisms has been observed during the formation of biofilm [29] and fruiting bodies [30, 31], where unicellular organisms form multi-cellular structures and resources are produced for the common good of the community (public goods co-operation, [32]. One of the most frequently studied forms of collective behaviour in microorganisms is observed in myxobacteria, which not only form intricate multicellular fruiting bodies complete with division of labour, but also exhibit highly coordinated collective movement to enhance mobility and attack prey. Collective social movement in myxobacteria is driven predominantly by cell-to-cell contact brought about by high cell density and exopolysaccharide secretion [33]. Similarly, a form of social motility has been described for Flavobacterium johnsoniae in which EPS-dependant assembly into microcolonies results in a collective gliding motility [34].
Collective movement of B. subtilis has commonly been described in liquid cultures [11, 35,36,37,38,39,40] as well as on and between solid surfaces [14, 41,42,43]. The collective movement of bacteria over solid surfaces, referred to as swarming, is associated with hyper-flagellation and peritrichously flagellated bacteria such as B. subtilis [14]. Swarming is also linked to the production of surfactants, an example of public goods that are controlled by quorum sensing, which reduces the tension between the moving cells and the viscous substrate [14]. The importance of surfactants during swarming is tightly coupled to the humidity of the surfaces and has been shown to not be essential at higher saturation levels [44, 45]. Studies have also shown that bacterial swarms may be directed by light in species that exhibit phototaxis [46, 47], suggesting the presence of possible biological response to stimuli. The impact of chemotaxis may even be increased within dense bacterial groups such as in swarms [48]. Swarming has not been demonstrated in natural environments to date [49] and the exact ecological function of this form of collective movement is still unclear [14]. Furthermore, there are debates over whether swarming is a co-ordinated response or if it occurs purely in response to the physical constraints and orientation of cells from collisions [41]. Brownian motion [38] and fluid flow [11, 39] have also been highlighted as the cause of microbial group movement and pattern formation, especially in high bacterial concentration suspensions.
Here we observed movements and behaviours that cannot be attributed solely to physical interactions, but rather strongly suggest the influence of biological drivers such as chemotaxis, quorum sensing, and mechanosensing. Our results show that groups of B. subtilis cells targeted the root tip, suggesting the directional impact of chemotaxis towards living plant roots. Around the root tip, the B. subtilis groups dynamically rearrange, which resulted in filamentous flocks that elongated over time and eventually disappeared by either retracting or dispersing. Flock elongation and retraction is linked to the B. subtilis cell concentration supporting the hypothesis that quorum sensing is involved in maintaining flocks. This is supported by the loss of co-ordination withing the flocks when the ComX-mediated quorum sensing system is disabled by mutation of comA. The necessity for monitoring cell density is also further confirmed by a mathematical model that shows how the strength of cell-to-cell communication is instrumental to the continuity of bacterial flocks. Indeed, when response to the hypothetical quorum sensing is too strong, the population of bacteria is unable to maintain filamentous structure and may lose connection to the nutrient source.
The dynamic changes in bacterial flock cell density are likely due to local densification and sparsification rather than cell growth and death. Though doubling times of various strains of B. subtilis are reported within the range of 30–120 min in a variety of nutrient rich media incubated at 30–35 °C [50, 51], a doubling time of 50 min, as calculated from our analysis, is highly unlikely under the nutrient and temperature conditions within our system. Microbial growth rates in soil, though still poorly understood [52], routinely show very slow doubling times at 20–25 °C in the rhizosphere of different plants [53,54,55].
After interacting dynamically with the root tip, the B. subtilis groups often disperse again and may adhere to form biofilm on more mature root zones. It is possible, that this observation occurs due to preferential attachment to mature root zones, like the root hair zone and elongation zone and to the heterogenous exudation along the root [56, 57]. Production of EPS by B. subtilis, although essential for the formation and maintenance of a stable biofilm on the root [58], is not essential for crowd movement but does appear to increase the coherence between the individual cells as seen with the social movement of myxobacteria [33] and flavobacteria [34].
Our mathematical model indicates that the diffusion of extracellular signals alone cannot explain the morphodynamics of bacterial flocks. The diffusivity of the medium is predicted to steadily increase the zone of influence of the flocking process, but experimental observation indicated that the width of the flock differed only in water, when viscosity was at a minimum. Such an effect could be obtained if the sensitivity to the extra cellular signal was, for example, dependent on viscous forces from the medium. The link between swimming velocity and viscosity of the surrounding liquid has been made more than half a century ago. Studies consistently show that bacterial swimming velocity increases with increasing viscosity up to approximately 2 mPa s−1 before decreasing [59,60,61]. Both the mid viscosity (2.2 mPa s−1) and the high viscosity (6.6 mPa s−1) solution used in our experiment fall well within the range associated with decreasing speed. It is possible crowd movement may also offer protection against shear stress in higher viscosity solutions. Indeed, studies in relatively low-density bacterial suspensions have shown that in viscous solutions bacteria resort to large scale collective motion [62], which may reduce the viscosity of the solution even to negative values [63]. Based on our findings, we propose that crowd movement in soil, like the collective behaviour of complex higher organisms, may be a form of energy conservation (e.g., hydrodynamics), and/or optimized search for resources [43], as well as protection from predation (e.g., nematodes) or antibiotics [64].
The pore size distribution in our system is very variable and ranges from 0.0 to 0.8 mm [18] and though pore size is likely a factor impacting the flock branch formation due to the physical barrier they represent; the pore size distribution would have impacted the flock branches equally regardless of viscosity. The importance of soil texture in ecosystem functioning is becoming ever more evident as a leading driver of trophic interactions in the micro-food web [65]. Pore size is frequently highlighted as playing a vital role in interspecies or interkingdom interactions by varying the accessibility between food/ prey and predators [66], which in turn may impact soil aggregate formation and ecosystem processes. Evidence suggests that, at least physically, Nafion provides a soil-like environment with a particle distribution resembling sand (250–1600 µm) and a water retention and cation exchange capacity comparable to vermiculite, in which plant root architecture is highly similar to natural soils or sand [67]. Similarly, the presence of solid surfaces and development of nutrient gradients should provide a more natural environment for soil microorganisms [53, 68].
It is still unclear how exactly microbial dynamics in Nafion, or other synthetic soils such as cryolite [69,70,71], compares to natural soils. Most natural soils due to clay and silt content, have smaller pores than Nafion or other synthetic soils, which likely means communication between bacterial cells is shorter ranged (i.e., smaller groups of bacteria, faster changes). In addition to this, unlike in our saturated system, natural soils frequently undergo cycles of drying and re-wetting, impacting connectivity of soil pores [72] and thus suggesting crowd movement may take place in thin water films and in fluid with higher viscosity [73]. Finally, numerous other chemical factors may also affect the physiology of the bacteria in natural soils, particularly variations in pH and aeration status. The observations reported in this study indicate the existence of highly coordinated movements of bacteria to navigate physically complex soil-like matrices with heterogenous nutrient availability. The challenge now is to better understand the mechanisms of crowd movement in natural systems with the aim to improve the understanding of soil ecosystems dynamics.
Conclusion
Microcosms consisting of plants grown in transparent synthetic soil with fluorescently labelled B. subtilis strains, coupled with light sheet image collection techniques, uncovered a co-ordinated form of microbial movement. Filamentous flocks formed by B. subtilis cells are attracted to the root, as well as to each other, resulting in the formation of a structure that is likely an energy efficient and protective mechanism that enables efficient translocation through the pores of the soil matrix. This work shed light on highly complex microbial movement in soil-like matrices. Such findings will greatly advance our knowledge of biodiversity and dynamics of soil ecosystems.
Materials and methods
Transparent soil and custom-made microcosms
Transparent soil was prepared from Nafion precursor beads (Ion Power Inc., USA) as previously described in Downie et al. [67] with minor modifications [18]. In short, Nafion transparent soil pellets were broken up to particle sizes of 250–1250 μm using in a cryogenic mill (SPEX SamplePrep 6770). Following this, the particles were converted to anionic form by soaking in 6 M KOH, 35% 5 M DMSO at 80 °C, followed by 3 M nitric acid at room temperature before titrating with Hoaglands basal medium (WVR ICNA092621822 at 1.6 g L−1). Sulphorhodamine B was added to the Nafion particles for the option of imaging the soil particles with a 561 nm laser. Transparent soil was sterilized by autoclaving for 20 min at 121 °C before use. The custom-made microcosms comprised of a 3-sided polydimethylsiloxane frame (PDMS, SYLGARDTM 184 Silicone Elastomer Base, Length 50 × outside-width 25 and inside-width 20 × Thickness 3 mm) between 2 standard microscopy glass slides (VWR, microscope slides at Length 76 × Width 26 × Thickness 1 mm) (see 65 for details on chamber construction).
Whole plant-environment light sheet microscopy
A non-commercial fluorescent light sheet microscope (FLSM) system with dual illumination was used to collect large field of view time-lapse data from live microcosms [18]. A laser source with up to 4 different wavelengths was used in conjunction with Powell and cylindrical lenses to generate uniform and overlapping light sheets which allow the illumination of only a thin slice of the sample (50 μm) at a time with a detection limit of approximately 105 CFU ml−1. Bacterial flocks with densities lower than the detection limit cannot be observed. An automated filter changer was built into the microscope to distinguish between the different scattering and fluorescent signals. This combined system allows a large variety of signals to be collected non-destructively on a single sample using a combination of light scattering and fluorescent based measurements. The objectives used (Mitutoyo Plan Apo Infinity Corrected 2X, 5X, 10X, Edmund Optics, UK) have a long working distance and a large field of view which allows the capture of images on large (several cm) samples in real time in situ. The system also included a computer-controlled growth light and a Peltier element-based temperature controlling system. Integrated software enabled volume data reconstruction and the resulting data consisted of several gigabytes. Frames were captured every 30 min and a large volume of the sample (6.1 × 22.9 × 8.0 mm) was scanned at a resolution of 12 μm.
Plants
Plants used include eudicots such as Tomato (Solanum lycopersicum var. Money maker) and Lettuce, (Lactuca sativa var. All Year Round) as well as monocots such as winter wheat (Triticum aestivum var. filum) and timothy grass (Phleum pratense). All seeds were surface sterilized by soaking in 10% bleach for 15 min before being thoroughly rinsed using sterile distilled H2O. Seeds were germinated on distilled water agar for 1–4 days, depending on species (until about 2 mm root is visible), before transferring a single seedling into a sterile microcosm filled with transparent soil, saturated with half strength Murashige and Skoog (MS) culture medium (without carbon source) later referred to as “MS medium” only.
Bacterial strains and growth media
The B. subtilis strains used in this study are listed in Table S1 and were grown for routine culturing purposes on lysogeny broth (LB: 1% (w/v) Bacto-peptone, 1% (w/v) NaCl, 0.5% (w/v) yeast extract and 1.5% (w/v) agar) plates or in liquid cultures at 37 °C. Antibiotic resistance cassette marked gene deletions or reporter fusions were moved between strains using either SSP1 mediated phage transduction [74] or genetic competence with plasmid DNA as indicated. To construct strains using genetic competency an altered version of 10 x Modified Competency (MC) media was used using the method previously described (10.7 g K2HPO4, 5.2 g KH2PO4, 20 g dextrose, 0.88 g sodium citrate dehydrate, 2.2 g L-glutamic acid monopotassium salt, and 1 g tryptone per 100 ml) [75]. For biofilm experiments, the strains for analysis were grown on MSgg agar plates (5 mM potassium phosphate (pH 7), 100 mM MOPS (pH 7), 2 mM MgCl2, 700 μM CaCl2, 50 μM MnCl2, 50 μM FeCl3, 1 μM ZnCl2, 2 μM thiamine, 0.5% (v/v) glycerol, 0.5% (w/v) glutamate, 1.5% (w/v) agar) [7]. Antibiotics were added as required at the following concentrations: tetracycline (12.5 μg/ml); spectinomycin (100 μg/ml); chloramphenicol (5 μg/ml); kanamycin (10 μg/ml). For antibiotic selection of pBL7 [76] in E. coli strain MC1061 ampicillin was used at a concentration of 100 µg/ml.
Colony biofilm assays
B. subtilis strains were streaked on LB agar plates which were incubated at 37 °C for ~16 h. A single colony was taken and grown in 3 ml of LB broth at 37 °C with agitation to an OD600 of ~1. At this point, 5 µl of each cell suspension was spotted onto MSgg media plates. The samples were incubated at 30 °C for 48 h before imaging. Colony biofilm imaging was performed using a Leica MZ16 FA stereoscope and LAS version 2.7.1. Scale bar in the upper images 10 mm and in the lower images 5 mm (Fig. S3).
Preparation of cells for microscopy analysis
All strains were grown in MSgg medium [31] for about 24 h at 18 °C, whilst shaking at 200 rpm (increase in OD600 of 1–1.5 orders of magnitude). After incubation, the MSgg solution was removed by centrifugation followed by reconstitution of the bacterial pellet in MS to remove any carbon contained in the bacterial solution. Approximately 2 million CFU were then inoculated onto a 2 mm by 2 mm sterile, fibreglass filter paper (inoculation patch) which was inserted into the microcosm, just beneath the soil surface, level with the newly emerging root (approximately 2 mm long). The insertion of the inoculation patch and the plant represents the biological time zero. The complete biological system was incubated at 21 °C for 20–24 h (light cycles of 16 h light at 60 µmol m−2 s−1 and 8 h dark) before replacing the MS medium in the system with Percoll (GE Healthcare) for index matching and live imaging in the self-developed FLSM system. The start of image collection (at 20–24 h post inoculation) represents the imaging time zero.
Bacterial growth curves
Fresh bacterial suspensions of B. subtilis NCIB 3610 were made in MSgg and diluted to a OD600 of 0.06. 20 replicates of 200 μl were added to 100-well microwell plates (Honeycomb; Thermo Fisher, USA). Growth rates were determined at 20 °C by measuring the OD600 every 15 min for 40 h with intermittent shaking using an automated plate reader (Bioscreen C plate reader; Oy Growth Curves Ab Ltd., Finland).
Viscous soil solutions
High viscosity soil samples were saturated with 78% Percoll solution (6.6 mPa s−1), mid viscosity samples with 34% Percoll (2.2 mPa s−1) and low viscosity samples with water (1 mPa s−1). Viscosity of all solutions was determined using a Cannon-Fenske viscometer (VFOC.931.v4, vidra FOC, Spain). Time lapse images were collected using the self-developed FLSM system described above.
Image processing and data analysis
Composite time lapse volume data was created by deconvolution, stitching and fusing of individual light sheet microscope scans, using the Matlab software (MathWorks, USA). Algorithms are detailed by Liu et al. [18] Filamentous flocks were traced and measured, manually. Only flocks of bacteria that persisted more than 1 h and were longer than 900 µm were analysed. A segmented line was traced manually along the maximum of fluorescence intensity in the image. The start and end point of the segmented line was defined as either the point where fluorescence intensity in the image is similar to the image background, or when the root is reached. The starting point is chosen as the one closest to the root surface. The segments were then used to transform the image coordinates so that the flock branches become linear, before fitting a Gaussian ridge on the transformed image to extract diameter and intensity values of individual flock branches. Statistical analysis and plots were created using R 3.1.2 [77], image processing using Fiji ImageJ [78] and then analysed for length, width, and B. subtilis cell density using custom made python scripts (https://zenodo.org/record/4946262).
Bacterial flock cell density estimation - calibration of system
Estimates of local cell densities (CFU ml−1) were calculated using a correlation equation obtained from 2 step calibration experiments performed and published previously [18]. In short, we captured images from of a series of homogenous bacterial suspensions at known concentrations (OD600) and determined the correlation with signal intensity of the image captured. The second correlation was obtained between the OD600 absorbance value of a bacterial solution and CFU per ml, by plating dilutions and counting the colonies formed after overnight incubation. The estimate of total number of bacterial cells within a flock was determined using the average local density (CFU ml−1) and the area of the flock.
Mathematical model
The ability of flocks to disappear and regroup elsewhere indicates the process is largely due to local densification and dispersion of the population rather than growth itself. The observed flocks orientate towards the root, converging at the tip. Therefore, kin attraction must also exist, and this occurs in directions not linked to the root chemotactic gradient. We hypothesize that the existence of a trade-off between the cost of motility and the cost of carbon deprivation.
To understand the paths taken by bacteria, and how these lead to the formation of flocking patterns in soil, we propose to model the trade-off between kin attraction and root attraction using the following equations:
The change of bacterial density ρ with time (left hand side) is linked to a directed transport term (first term on the right-hand side) and a dispersive term (second term on the right-hand side) with bacterial diffusion coefficient D. The directed transport of the bacteria is then decomposed into two terms which determine the direction of the movement. v models the chemotactic response towards the root while I models the attraction of bacteria towards each other in a density dependent manner:
where φ is the concentration of nutrients released by the root, C the chemotactic sensitivity parameter, and \(\rho \ast \eta\) is the interaction potential created by the density of surrounding bacteria, with η a kernel function encoding the interaction with neighbours through a specified radius. Since the kernel function encode the effect of a diffusible extracellular signal, it is expressed as a function of the diffusion coefficient D, i.e., \(\eta = f\left( {x,\sqrt D } \right)\). The strength of the interaction K therefore defines the morphological properties of the flocks (from diffuse to filamentous flocks and to aggregated morphologies). Therefore, in this chemotactic model, the direction of bacterial velocity is dependent on the gradient of the bacterial density.
The model was implemented with a finite volume scheme for advection and diffusion on a 2D cartesian grid. The scheme was modified to include a nonlocal estimation of the concentration distribution. The concentration of nutrients φ is determined by solving an eikonal equation with the root as a target (https://zenodo.org/record/4946262).
Data availability
The link https://zenodo.org/record/4946262 provides the codes for numerical simulations of models of bacterial crowd movements. Processed image data and code for analysis of bacterial flocks is available at: https://github.com/LionelDupuy/CrowdMovement.
References
Kuzyakov Y, Razavi BS. Rhizosphere size and shape: Temporal dynamics and spatial stationarity. Soil Biol Biochem. 2019;135:343–60.
Teixeira PJ, Colaianni NR, Fitzpatrick CR, Dangl JL. Beyond pathogens: Microbiota interactions with the plant immune system. Curr Opin Microbiol. 2019;49:7–17.
Alirezaeizanjani Z, Großmann R, Pfeifer V, Hintsche M, Beta C. Chemotaxis strategies of bacteria with multiple run modes. Sci Adv. 2020;6:eaaz6153.
Gao S, Wu H, Yu X, Qian L, Gao X. Swarming motility plays the major role in migration during tomato root colonization by Bacillus subtilis SWR01. Biol Control. 2016;98:11–17.
Mitchell JG, Kogure K. Bacterial Motility: Links to the environment and a driving force for microbial physics. FEMS Microbiol Ecol. 2006;55:3–16.
Kalamara M, Spacapan M, Mandic-Mulec I, Stanley-Wall NR. Social behaviours by Bacillus subtilis: Quorum sensing, kin discrimination and beyond. Mol Microbiol. 2018;110:863–78.
Posada LF, Álvarez JC, Romero-Tabarez M, de-Bashan L, Villegas-Escobar V. Enhanced molecular visualization of root colonization and growth promotion by Bacillus subtilis EA-CB0575 in different growth systems. Microbiol Res. 2018;217:69–80.
Beauregard PB, Yunrong C, Vlamakis H, Losick R, Kolter R. Bacillus subtilis Biofilm induction by plant polysaccharides. Proc Natl Acad Sci USA. 2013;110:1621–30.
Allard-Massicotte R, Tessier L, Lécuyer F, Lakshmanan V, Lucier J. Bacillus subtilis early colonization of Arabidopsis thaliana roots involves multiple chemotaxis receptors. mBio 2016;7:1–10.
Massalha H, Korenblum E, Malitsky S, Shapiro OH, Aharoni A. Live imaging of root-bacteria interactions in a microfluidics setup. Proc Natl Acad Sci USA. 2017;114:4549–54.
Koch DL, Subramanian G. Collective hydrodynamics of swimming microorganisms: Living fluids. Annu Rev Fluid Mech. 2011;43:637–59.
Wioland H, Lushi E, Goldstein RE. Directed collective motion of bacteria under channel confinement. New J Phys. 2016;18:eaaz6153.
Petroff A, Libchaber A. Erratum: Hydrodynamics and collective behavior of the tethered bacterium Thiovulum majus. Proc Natl Acad Sci USA. 2016;111:5. E537-E545
Kearns DB. A field guide to bacterial swarming motility. Nat Rev Microbiol. 2010;8:634–44.
Bais HP, Fall R, Vivanco JM. Biocontrol of Bacillus subtilis against infection of arabidopsis roots by Pseudomonas syringae is facilitated by biofilm formation and surfactin production. Plant Physiol. 2004;134:307–19.
De Souza R, Ambrosini A, Passaglia LMP. Plant growth-promoting bacteria as inoculants in agricultural soils. Genet Mol Biol. 2015;38:401–19.
Roy K, Ghosh D, DeBruyn JM, Dasgupta T, Wommack KE, Liang X, et al. Temporal dynamics of soil virus and bacterial populations in agricultural and early plant successional soils. Front Microbiol. 2020;11:1–13.
Liu Y, Patko D, Engelhardt IC, George TS, Stanley-Wall NP, Ladmiral V. et al. Whole plant-environment microscopy reveals how Bacillus subtilis utilises the soil pore space to colonise plant roots. Proc Natl Acad Sci USA. 2021;118:e2109176118.
Einstein A. On the motion of small particles suspended in liquids at rest required by the molecular-kinetic theory of heat. Ann Phys. 1905;17:549–60.
Shellard A, Mayor R. Rules of Collective Migration: From the wildebeest to the neural crest: Rules of neural crest migration. Philos Trans R Soc B Biol Sci. 2020;375:1–9.
Torney CJ, Lamont M, Debell L, Angohiatok RJ, Leclerc LM, Berdahl AM. Inferring the rules of social interaction in migrating caribou. Philos Trans R Soc B Biol Sci. 2018;373:20170385.
Ballerini MN, Cabibbo R, Candelier A, Cavagna E, Cisbani I, Giardina V, et al. Interaction ruling animal collective behavior depends on topological rather than metric distance: Evidence from a field study. Proc Natl Acad Sci USA. 2008;105:1232–37.
Cavagna A, Cimarelli A, Giardina I, Parisi G, Santagati R, Stefanini F, et al. Scale-free correlations in starling flocks. Proc Natl Acad Sci USA. 2010;107:11865–70.
Katz Y, Tunstrøm C, Ioannou CC, Huepe C, Couzin ID. Inferring the structure and dynamics of interactions in schooling fish. Proc Natl Acad Sci USA. 2011;108:18720–25.
Buhl JD, Sumpter JT, Couzin ID, Hale JJ, Despland E, Miller ER, et al. From disorder to order in marching locusts. Science 2006;312:1402–6.
Seeley TD, Visscher PK. Quorum Sensing during nest-site selection by honeybee swarms. Behav Ecol Sociobiol. 2004;56:594–601.
Zhang HP, Be’er A, Florin EL, Swinney HL. Collective motion and density fluctuations in bacterial colonies. Proc Natl Acad Sci USA. 2010;107:13626–30.
Hughey LF, Hein AM, Strandburg-Peshkin A, Jensen FH. Challenges and solutions for studying collective animal behaviour in the wild. Philos Trans R Soc B Biol Sci. 2018;373:1–13.
Nadell CD, Xavier JB, Foster KR. The sociobiology of biofilms. FEMS Microbiol Rev. 2009;33:206–24.
Velicer GJ, Vos M. Sociobiology of the myxobacteria. Ann Rev Microbiol. 2009;63:599–623.
Branda SS, González-Pastor JE, Ben-Yehuda S, Losick R, Kolter R. Fruiting body formation by Bacillus subtilis. Proc Natl Acad Sci USA. 2001;98:11621–26.
Cordero OX, Wildschutte H, Kirkup B, Proehl S, Ngo L, Hussain F, et al. Antibiotic production and resistance. Sci Rep. 2012;337:1228–31.
Muñoz-Dorado J, Marcos-Torres FJ, García-Bravo E, Moraleda-Muñoz A, Pérez J. Myxobacteria: Moving, killing, feeding, and surviving together. Front Microbiol. 2016;7:1–18.
Li C, Hurley A, Hu W, Warrick JW, Lozano GL, Ayuso JM, et al. Social motility of biofilm-like microcolonies in a gliding bacterium. Nat Commun. 2021;12:1–12.
Sokolov A, Aranson IS, Kessler JO, Goldstein RE. Concentration dependence of the collective dynamics of swimming bacteria. Phys Rev Lett. 2007;98:158102.
Cisneros LH, Cortez R, Dombrowski C, Goldstein RE, Kessler JO. Fluid dynamics of self-propelled microorganisms, from individuals to concentrated populations. Exp Fluids. 2007;43:737–53.
Tuval I, Cisneros L, Dombrowski C, Wolgemuth CW, Kessler JO, Goldstein RE. Bacterial swimming and oxygen transport near contact lines. Proc Natl Acad Sci USA. 2005;102:2277–82.
Li G, Tam L, Tang JX. Amplified effect of brownian motion in bacterial near-surface swimming. Proc Natl Acad Sci USA. 2008;105:18355–59.
Lushi E, Wioland H, Goldstein RE. Fluid flows created by swimming bacteria drive self-organization in confined suspensions. Proc Natl Acad Sci USA. 2014;111:9733–38.
Ryan SD, Sokolov A, Berlyand L, Aranson IS. Correlation properties of collective motion in bacterial suspensions. New J Phys. 2013;15:105021.
Damton NC, Turner L, Rojevsky S, Berg HC. Dynamics of bacterial swarming. Biophys J. 2010;98:2082–90.
Ingham CJ, Jacob EB. Swarming and complex pattern formation in Paenibacillus vortex studied by imaging and tracking cells. BMC Microbiol. 2008;8:1–16.
Ariel G, Rabani A, Benisty S, Partridge JD, Harshey RM, Be’Er A. Swarming bacteria migrate by lévy walk. Nat Commun. 2015;6:8396.
Hamze K, Autret S, Hinc K, Laalami S, Julkowska D, Briandet R, et al. Single-cell analysis in situ in a Bacillus subtilis swarming community identifies distinct spatially separated subpopulations differentially expressing Hag (Flagellin), including specialized swarmers. Microbiol. 2011;157:2456–69.
Ghelardi E, Salvetti S, Ceragioli M, Gueye SA, Celandroni F, Senesi S. Contribution of surfactin and swrA to flagellin expression, swimming, and surface motility in Bacillus subtilis. Appl Environ Microbiol. 2012;78:6540–44.
Wilde A, Mullineaux CW. Light-controlled motility in prokaryotes and the problem of directional light perception. FEMS Microbiol Rev. 2017;41:900–22.
Zhang J, Luo Y, Poh CL. Blue light-directed cell migration, aggregation, and patterning. J Mol Biol. 2020;432:3137–48.
Tian T, Sun B, Shi H, Gao T, He Y, Li Y, et al. Sucrose triggers a novel signalling cascade promoting Bacillus subtilis rhizosphere colonization. ISME J 2021;15:2723–37.
Harshey RM, Partridge JD. Shelter in a swarm. J Mol Biol. 2015;427:3683–94.
Burdett IDJ, Kirkwood TBL, Whalley JB. Growth kinetics of individual Bacillus subtilis cells and correlation with nucleoid extension. J Bacteriol. 1986;167:219–30.
Sharpe ME, Hauser PM, Sharpe RG, Errington J. Bacillus subtilis cell cycle as studied by fluorescence microscopy: Constancy of cell length at initiation of DNA replication and evidence for active nucleoid partitioning. J Bacteriol. 1998;180:547–55.
Rousk J, Bååth E. Growth of saprotrophic fungi and bacteria in soil. FEMS Microbiol Ecol. 2011;78:17–30.
Bennett RA, Lynch JM. Bacterial growth and development in the rhizosphere of gnotobiotic cereal plants. Microbiol. 1981;125:95–102.
Felici C, Vettori L, Giraldi E, Forino LMC, Toffanin A, Tagliasacchi AM, et al. Single and co-inoculation of Bacillus subtilis and Azospirillum brasilense on Lycopersicon Esculentum: Effects on plant growth and rhizosphere microbial community. Appl Soil Ecol. 2008;40:260–70.
Arkhipova TN, Galimsyanova NF, Kuzmina LY, Vysotskaya LB, Sidorova LV, Gabbasova IM, et al. Effect of seed bacterization with plant growth-promoting bacteria on wheat productivity and phosphorus mobility in the rhizosphere. Plant Soil Environ. 2019;65:313–19.
Marschner P, Crowley D, Rengel Z. Rhizosphere interactions between microorganisms and plants govern iron and phosphorus acquisition along the root axis - model and research methods. Soil Biol Biochem. 2011;43:883–94.
Lagos ML, Maruyama F, Nannipieri P, Mora ML, Jorquera MA. Current Overview on the study of bacteria in the rhizosphere by modern molecular techniques: A Mini-Review. J Soil Sci Plant Nutr. 2015;15:504–23.
Gerwig J, Kiley TB, Gunka K, Stanley-Wall N, Stülke J. The protein tyrosine kinases epsB and ptkA differentially affect biofilm formation in Bacillus Subtilis. Microbiol. 2014;160:682–91.
Shoesmith JG. The measurement of bacterial motility. J Gen Microbiol. 1960;22:528–35.
Schneider WR, Doetsch RN. Effect of viscosity on bacterial motility. J Bacteriol. 1974;117:696–701.
Kaiser GE, Doetsch RN. Enhanced translational motion of Leptospira in viscous environments. Nature 1975;255:656–57.
Ryan SD, Haines BM, Berlyand L, Ziebert F, Aranson IS. Viscosity of bacterial suspensions: Hydrodynamic interactions and self-induced noise. Phys Rev E Stat Nonlin Soft Matter Phys. 2011;E83:050904.
López HM, Gachelin J, Douarche C, Auradou H, Clément E. Turning bacteria suspensions into superfluids. Phys Rev Lett. 2015;115:028301.
Butler MT, Wang Q, Harshey RM. Cell density and mobility protect swarming bacteria against antibiotics. Proc Natl Acad Sci USA. 2010;107:3776–81.
Erktan A, Or D, Scheu S. The physical structure of soil: Determinant and consequence of trophic interactions. Soil Biol Biochem. 2020;148:107876.
Rønn R, Thomsen IK, Jensen B. Naked amoebae, flagellates and nematodes in soil of different texture. Eur J Soil Biol. 1995;31:135–41.
Downie H, Holden N, Otten W, Spiers AJ, Valentine TA, Dupuy LX. Transparent soil for imaging the rhizosphere. PLoS ONE. 2012;7:1–6.
Mills AL. Keeping in Touch: Microbial life on soil particle surfaces. Adv Agron. 2003;78:1–43.
Downie HF, Valentine TA, Otten W, Spiers AJ, Dupuy LX. Transparent soil microcosms allow 3D spatial quantification of soil microbiological processes in vivo. Plant Signal Behav. 2014;9:e970421.
O’Callaghan FE, Braga RA, Neilson R, MacFarlane SA, Dupuy LX. New live screening of plant-nematode interactions in the rhizosphere. Sci Rep. 2018;8:1–17.
Sharma K, Palatinszky M, Nikolov G, Berry D, Shank EA. Transparent soil microcosms for live-cell imaging and non-destructive stable isotope probing of soil microorganisms. ELife 2020;9:1–28.
Bickel S, Or D. Soil bacterial diversity mediated by microscale aqueous-phase processes across biomes. Nat Commun. 2020;11:1–9.
Farré M, Sanchís J, Barceló D. Analysis and assessment of the occurrence, the fate and the behavior of nanomaterials in the environment. Trends Anal Chem. 2011;30:517–27.
Verhamme DT, Kiley TB, Stanley-Wall NR. DegU co-ordinates multicellular behaviour exhibited by Bacillus subtilis. Mol Microbiol. 2007;65:554–68.
Konkol MA, Blair KM, Kearns DB. Plasmid-encoded comi inhibits competence in the ancestral 3610 strain of Bacillus subtilis. J Bacteriol. 2013;195:4085–93.
Stanley NR, Lazazzera BA. Defining the genetic differences between wild and domestic strains of Bacillus subtilis that affect poly-γ-DL-glutamic acid production and biofilm formation. Mol Microbiol. 2005;57:1143–58.
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. 2020. URL https://www.R-project.org/.
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, et al. (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods. 2012;9:676–82.
Acknowledgements
The James Hutton Institute & Scotland’s Rural College were supported by funds from the Rural and Environment Science and Analytical Services Division of the Scottish Government. We thank Margarita Kalamara, Adam Ostrowski and the NSW laboratory for sharing their strains with us, Elliot Erskine, Jacqueline Marshall from JHI James Hutton Institute for advice on B. subtilis culture and other laboratory requirements, and Kathryn Wright from the James Hutton Institute for help with confocal microscopy. This work was funded by the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (Grant agreement No. 647857-SENSOILS). We also acknowledge the funding from the Spanish Ministry of Science and Innovation (MICINN) under de project MICROCROWD (PID2020-112950RR-I00). Work in the NSW laboratory is funded by the Biotechnology and Biological Science Research Council (BBSRC) [BB/P001335/1, BB/R012415/1].
Author information
Authors and Affiliations
Contributions
Funding Acquisition: LD. Conceptualisation: LXD, TJD, NH, IE. Experimental Work: IE, DP, YL, GHM, TS. Data Analysis: IE, LD, MM. Mathematical Modelling: MM, MP, LXD. Supervision: TSG, MM, MP, NRS-W, NH, TJD, LXD. Validation: LE, MK, NSW, FAD. Visualisation: LE, MK. Writing – original draft: IE, LXD. Writing – review & editing: MM, TSG, MM, MP, NRS-W, NH, TJD, LXD.
Corresponding author
Ethics declarations
Competing interests
The material is original research, has not been previously published and has not been submitted for publication elsewhere while under consideration. The authors declare no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Engelhardt, I.C., Patko, D., Liu, Y. et al. Novel form of collective movement by soil bacteria. ISME J 16, 2337–2347 (2022). https://doi.org/10.1038/s41396-022-01277-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-022-01277-w
This article is cited by
-
Microbes in porous environments: from active interactions to emergent feedback
Biophysical Reviews (2024)
-
Characterization and environmental applications of soil biofilms: a review
Environmental Chemistry Letters (2024)
-
Smart soils track the formation of pH gradients across the rhizosphere
Plant and Soil (2023)