Abstract
Background
CYP2C9 is a member of the cytochrome P450 (CYP) superfamily responsible for the metabolism of 16% of drugs that undergo oxidative metabolism. The activity of CYP2C9 exhibits marked inter-individual variability, which translates into prominent differences in the pharmacokinetics of CYP2C9 substrates, some of which are characterized by a narrow therapeutic window. Genetic polymorphisms in the gene encoding for CYP2C9 account for a fraction of the variability in CYP2C9 activity. The phenytoin metabolic ratio (PMR) is a marker of CYP2C9 activity in vivo, which correlates with CYP2C9 genetic polymorphisms.
Objective
The purpose of the current study was to evaluate the ability of the PMR to predict the oral clearance of (S)-warfarin (SWOCL) and its formation clearance towards its CYP2C9-mediated metabolites (SWCLf) [i.e., 6- and 7-hydroxy-(S)-warfarin].
Methods
The study was conducted in 150 healthy non-smoker subjects (segment 1) and 60 patients treated with warfarin (segment 2). In the first segment, the participants received on two separate occasions a single 300-mg dose of phenytoin and at least 7 days later a single dose of warfarin (5 or 10 mg). The same PMR procedure was performed in the second segment, except that it was performed either before warfarin initiation or after the patients had reached stable anticoagulation. The PMR was derived from the ratio of 5-(4-hydroxyphenyl)-5-phenyl-hydantoin content in a 24-hour urine collection to plasma phenytoin concentration 12- (PMR24/12) or 24- (PMR24/24) post-dosing. In segment 1, SWOCL was calculated from the ratio of (S)-warfarin dose to the warfarin area under the plasma concentration–time curve extrapolated to infinity and the SWCLf from the ratio of urine content of 6- and 7-hydroxy-(S)-warfarin to (S)-warfarin area under the (S)-warfarin plasma concentration–time curve until the last measured timepoint. In segment 2, estimated SWOCL was derived from the ratio of (S)-warfarin dose to the mid-interval plasma concentration of (S)-warfarin.
Results
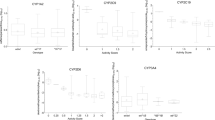
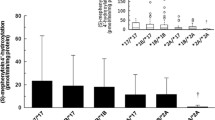
The PMR, SWOCL, and SWCLf varied significantly between carriers of different CYP2C9 genotypes in both healthy subjects (p < 0.001) and patients (p < 0.005). However, PMR and SWOCL values exhibited substantial intra-genotypic variability. PMR24/12 and PMR24/24 were significantly correlated with SWOCL both in healthy subjects (r = 0.62 and r = 0.67, respectively, p < 0.001) and in patients (r = 0.57 and r = 0.61, respectively, p < 0.001). In a multiple regression model that included all variables that correlated with SWOCL, PMR was the strongest predictor, explaining 44% and 38% of the variability in SWOCL among healthy subjects and patients, respectively, and accounting for 95.7% (44%/46%) and 90.5% (38%/42%) of the total explained variability in SWOCL among healthy subjects and patients, respectively.
Conclusions
The PMR is the strongest predictor of SWOCL, and as such, it exhibits a significant advantage over the CYP2C9 genotype. The inclusion of PMR in future dosing algorithms of CYP2C9 substrates characterized by a narrow therapeutic window should be encouraged and further investigated.







Similar content being viewed by others
References
Daly AK, Rettie AE, Fowler DM, Miners JO. Pharmacogenomics of CYP2C9: functional and clinical considerations. J Pers Med. 2017;8:1.
Zanger UM, Schwab M. Cytochrome P450 enzymes in drug metabolism: regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacol Ther. 2013;138:103–41.
Miners JO, Birkett DJ. Cytochrome P4502C9: an enzyme of major importance in human drug metabolism. Br J Clin Pharmacol. 1998;45:525–38.
Rettie AE, Jones JP. Clinical and toxicological relevance of CYP2C9: drug-drug interactions and pharmacogenetics. Ann Rev Pharmacol Toxicol. 2005;45:477–94.
Scordo MG, Pengo V, Spina E, Dahl ML, Gusella M, Padrini R. Influence of CYP2C9 and CYP2C19 genetic polymorphisms on warfarin maintenance dose and metabolic clearance. Clin Pharmacol Ther. 2002;72:702–10.
Veronese ME, Miners JO, Rees DL, Birkett DJ. Tolbutamide hydroxylation in humans: lack of bimodality in 106 healthy subjects. Pharmacogenetics. 1993;3:86–93.
PHARMGKB. Named alleles. https://www.pharmgkb.org/gene/PA126/haplotype. Accessed 22 May 2022.
Veenstra DL, Blough DK, Higashi MK, Farin FM, Srinouanprachan S, Rieder MJ, et al. CYP2C9 haplotype structure in European American warfarin patients and association with clinical outcomes. Clin Pharmacol Ther. 2005;77:353–64.
Wang D, Sun X, Gong Y, Gawronski BE, Langaee TY, Shahin MH, et al. CYP2C9 promoter variable number tandem repeat polymorphism regulates mRNA expression in human livers. Drug Metab Dispos. 2012;40:884–91.
Dayer P, Leemann T, Striberni R. Dextromethorphan O-demethylation in liver microsomes as a prototype reaction to monitor cytochrome P-450 db1 activity. Clin Pharmacol Ther. 1989;45:34–40.
Silvado CE, Terra VC, Twardowschy CA. CYP2C9 polymorphisms in epilepsy: influence on phenytoin treatment. Pharmacogenomics Pers Med. 2018;11:51–8.
US FDA. FY2015 regulatory science research report: narrow therapeutic index drugs. https://www.fda.gov/industry/generic-drug-user-fee-amendments/fy2015-regulatory-science-research-report-narrow-therapeutic-index-drugs. Accessed 22 May 2022
US FDA. Quality and bioequivalence standards for narrow therapeutic index drugs. https://www.fda.gov/files/drugs/published/Quality-and-Bioequivalence-Standards-for-Narrow-Therapeutic-Index-Drugs.pdf. Accessed 22 May 2022.
Habet S. Narrow therapeutic index drugs: clinical pharmacology perspective. J Pharm Pharmacol. 2021;73:1285–91.
Jayachandran P, Okochi H, Frassetto LA, Park W, Fang L, Zhao L, et al. Evaluating within-subject variability for narrow therapeutic index drugs. Clin Pharmacol Ther. 2019;105:411–6.
Tamargo J, Le Heuzey JY, Mabo P. Narrow therapeutic index drugs: a clinical pharmacological consideration to flecainide. Eur J Clin Pharmacol. 2015;71:549–67.
Yu LX, Jiang W, Zhang X, Lionberger R, Makhlouf F, Schuirmann DJ, et al. Novel bioequivalence approach for narrow therapeutic index drugs. Clin Pharmacol Ther. 2015;97:286–91.
Lee CR, Pieper JA, Frye RF, Hinderliter AL, Blaisdell JA, Goldstein JA. Tolbutamide, flurbiprofen, and losartan as probes of CYP2C9 activity in humans. J Clin Pharmacol. 2003;43:84–91.
Yasar U, Eliasson E, Forslund-Bergengren C, Tybring G, Gadd M, Sjoqvist F, Dahl ML. The role of CYP2C9 genotype in the metabolism of diclofenac in vivo and in vitro. Eur J Clin Pharmacol. 2001;57:729–35.
Zgheib NK, Frye RF, Tracy TS, Romkes M, Branch RA. Evaluation of flurbiprofen urinary ratios as in vivo indices for CYP2C9 activity. Br J Clin Pharmacol. 2007;63:477–87.
Yasar U, Forslund-Bergengren C, Tybring G, Dorado P, Llerena A, Sjoqvist F, et al. Pharmacokinetics of losartan and its metabolite E-3174 in relation to the CYP2C9 genotype. Clin Pharmacol Ther. 2002;71:89–98.
Caraco Y, Muszkat M, Wood AJ. Phenytoin metabolic ratio: a putative marker of CYP2C9 activity in vivo. Pharmacogenetics. 2001;11:587–96.
Caraco Y. Abstracts of the 2002 annual meeting of the American Society for Clinical Pharmacology and Therapeutics. Atlanta, Georgia, USA. March 24-27, 2002. Clin Pharmacol Ther. 2002;71:P58.
Shaul C, Blotnick S, Muszkat M, Caraco Y. Abstracts of the 2002 annual meeting of the American Society for Clinical Pharmacology and Therapeutics. Atlanta, Georgia, USA. March 24–27, 2002. Clin Pharmacol Ther. 2002;71:P57.
Abu Ghosh Z, Alamia S, Shaul C, Caraco Y. Comparison of CYP2C9 activity in Ethiopian and non-Ethiopian jews using phenytoin as a probe. Front Pharmacol. 2020;11: 566842.
Caraco Y, Blotnick S, Muszkat M. CYP2C9 genotype-guided warfarin prescribing enhances the efficacy and safety of anticoagulation: a prospective randomized controlled study. Clin Pharmacol Ther. 2008;83:460–70.
Shaul C, Blotnick S, Muszkat M, Bialer M, Caraco Y. Quantitative assessment of CYP2C9 genetic polymorphisms effect on the oral clearance of S-warfarin in healthy subjects. Mol Diagn Ther. 2017;21:75–83.
Zar JH. Biostatistical analysis. 5th ed. Essex: Pearson; 2014.
Vogl S, Lutz RW, Schonfelder G, Lutz WK. CYP2C9 genotype vs. metabolic phenotype for individual drug dosing: a correlation analysis using flurbiprofen as probe drug. PLoS ONE. 2015;10:e0120403.
Fekete F, Mangó K, Déri M, Incze E, Minus A, Monostory K. Impact of genetic and non-genetic factors on hepatic CYP2C9 expression and activity in Hungarian subjects. Sci Rep. 2021;11:17081.
Mostafa S, Kirkpatrick CMJ, Byron K, Sheffield L. An analysis of allele, genotype and phenotype frequencies, actionable pharmacogenomic (PGx) variants and phenoconversion in 5408 Australian patients genotyped for CYP2D6, CYP2C19, CYP2C9 and VKORC1 genes. J Neural Transm (Vienna). 2019;126:5–18.
Temesvari M, Kobori L, Paulik J, Sarvary E, Belic A, Monostory K. Estimation of drug-metabolizing capacity by cytochrome P450 genotyping and expression. J Pharmacol Exp Ther. 2012;341:294–305.
Yamazaki H, Inoue K, Shimada T. Roles of two allelic variants (Arg144Cys and Ile359Leu) of cytochrome P4502C9 in the oxidation of tolbutamide and warfarin by human liver microsomes. Xenobiotica. 1998;28:103–15.
Scripture CD, Pieper JA. Clinical pharmacokinetics of fluvastatin. Clin Pharmacokinet. 2001;40:263–81.
Dai DP, Wang SH, Geng PW, Hu GX, Cai JP. In vitro assessment of 36 CYP2C9 allelic isoforms found in the Chinese population on the metabolism of glimepiride. Basic Clin Pharmacol Toxicol. 2014;114:305–10.
Maekawa K, Harakawa N, Sugiyama E, Tohkin M, Kim SR, Kaniwa N, et al. Substrate-dependent functional alterations of seven CYP2C9 variants found in Japanese subjects. Drug Metab Dispos. 2009;37:1895–903.
Niinuma Y, Saito T, Takahashi M, Tsukada C, Ito M, Hirasawa N, et al. Functional characterization of 32 CYP2C9 allelic variants. Pharmacogenomics J. 2014;14:107–14.
Fuhr U, Jetter A, Kirchheiner J. Appropriate phenotyping procedures for drug metabolizing enzymes and transporters in humans and their simultaneous use in the “cocktail” approach. Clin Pharmacol Ther. 2007;81:270–83.
King BP, Khan TI, Aithal GP, Kamali F, Daly AK. Upstream and coding region CYP2C9 polymorphisms: correlation with warfarin dose and metabolism. Pharmacogenetics. 2004;14:813–22.
Mosher CM, Tai G, Rettie AE. CYP2C9 amino acid residues influencing phenytoin turnover and metabolite regio- and stereochemistry. J Pharmacol Exp Ther. 2009;329:938–44.
Marcath LA, Pasternak AL, Hertz DL. Challenges to assess substrate-dependent allelic effects in CYP450 enzymes and the potential clinical implications. Pharmacogenomics J. 2019;19:501–15.
Acknowledgements
Dr. Chanan Shaul is a post-doctoral fellow at the Hebrew University of Jerusalem under the supervision of Prof. Yoseph Caraco and Prof. Meir Bialer. The technical support of Mrs. Ester Zadka and Mrs. Ilanit Linzer is greatly appreciated.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was not supported by any funding source.
Conflicts of interest/competing interests
HS, SB, LA, MM, and YC do not have any conflicts of interest to disclose. MB received consultancy fees from Boehringer Ingelheim, Medison, Pharma 2B, Rekah-Vitamned, and Xenon Pharmaceuticals.
Ethics approval
The study protocols were approved by the Hadassah Institutional Review Board.
Consent to participate
Following a detailed explanation, all participants gave their written consent to participate in the study.
Consent for publication
Not applicable.
Availability of data and material
Not applicable.
Code availability
Not applicable.
Author contributions
CS, YC, and MB wrote the manuscript, LA, MB, and YC designed the research, CS and LA performed the research, and CS, SB, and LA analyzed the data.
Rights and permissions
About this article
Cite this article
Shaul, C., Blotnick, S., Adar, L. et al. Phenytoin Metabolic Ratio, a Marker of CYP2C9 Activity, is Superior to the CYP2C9 Genotype as a Predictor of (S)-Warfarin Clearance. Clin Pharmacokinet 61, 1187–1198 (2022). https://doi.org/10.1007/s40262-022-01141-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40262-022-01141-2