Abstract
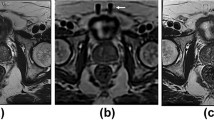
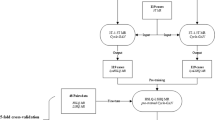
This study aimed to assess the clinical feasibility of employing synthetic diffusion-weighted (DW) images with different b values (50, 400, 800 s/mm2) for the prostate cancer patients with the help of three models, namely CycleGAN, Pix2PiX, and DC2Anet. DW images of 170 prostate cancer patients were used to train and test models. Here, 119 patients were assigned to the training set and 51 patients to the testing set according to a ratio of 7:3. To generate synthetic b value DW images based on CycleGAN, Pix2Pix, and DC2Anet networks, three experiments were performed as follows: generating synthetic DW images with b values of 400 and 800 s/mm2 from acquired DW images with b value of 50 s/mm2; generating synthetic DW images with b value of 800 8 s/mm2 from acquired DW images with b value of 400 s/mm2. Five metrics were used to compare real and synthetic b values. These metrics included Mean Absolute Error (MAE), Root Mean Squared Error (RMSE), Pearson’s Correlation Coefficient (PCC), Peak-Signal-to-Noise-Ratio (PSNR), and Structural Similarity Index Measure (SSIM). As well as, ADC values for different b values were computed using the mono-exponentially mode. The whole prostate volume was manually segmented by drawing regions of interest (ROIs) in each slice of the ADC maps. P values less than 0.05 were considered statistically significant. Based on the quantitative evaluation and for all metrics, especially for generating b values of 400 and 800 s/mm2 from a b value of 50 s/mm2, the DC2Anet model was found accurate and it outperformed CycleGAN and Pix2Pix models (P < 0.05). It is necessary to mention that the agreement between synthetic ADC (sADC) and real ADC (rADC) was satisfactory. No significant difference was observed in the one-way ANOVA between sADC and rADC in the whole prostate volume (P > 0.05). Our results showed the significant potential of the three used models for generating images with different b values in the case of prostate cancer. The results demonstrated that the used three models were accurate and robust for generating DW images and also, they outperformed other methods mentioned in the literature review.











Similar content being viewed by others
Availability of data and code
The dataset and code used and/or analyzed during the current study are available from the corresponding author on reasonable request only for personal uses, this dataset can be downloaded from a public repository: https://wiki.cancerimagingarchive.net/pages/viewpage.action?pageId=23691656
References
P. Rawla, World J. Oncol. 10, 63 (2019)
P.J.L. De Visschere, C. Standaert, J.J. Fütterer, G.M. Villeirs, V. Panebianco, J. Walz et al., Eur. Urol. Oncol. 2, 47–76 (2019)
S. Heydarheydari, V. Dehlaghi, A. Haghparast, Acta Med. Iran. 54, 343–4 (2016)
S.M. Rezaeijo, B. Hashemi, B. Mofid, M. Bakhshandeh, A. Mahdavi, M.S. Hashemi, Radiat. Oncol. 16, 1–16 (2021)
D.C. Johnson, R.E. Reiter, Transl. Androl. Urol. 6, 472 (2017)
G. Gaunay, V. Patel, P. Shah, D. Moreira, S.J. Hall, M.A. Vira et al., Asian J. Urol. 4, 68–74 (2017)
J.O. Barentsz, J. Richenberg, R. Clements, P. Choyke, S. Verma, G. Villeirs et al., Eur. Radiol. 22, 746–757 (2012)
M. Kasel-Seibert, T. Lehmann, R. Aschenbach, F.V. Guettler, M. Abubrig, M.-O. Grimm et al., Eur. J. Radiol. 85, 726–731 (2016)
T. Barrett, B. Turkbey, P.L. Choyke, Clin. Radiol. 70, 1165–1176 (2015)
C.V. Dinh, P. Steenbergen, G. Ghobadi, S.W. Heijmink, F.J. Pos, K. Haustermans et al., Phys. Med. 32, 446–451 (2016)
C. Debus, R. Floca, M. Ingrisch, I. Kompan, K. Maier-Hein, A. Abdollahi et al., BMC Bioinform. 20, 1–18 (2019)
R. Shimofusa, H. Fujimoto, H. Akamata, K. Motoori, S. Yamamoto, T. Ueda et al., J. Comput. Assist. Tomogr. 29, 149–153 (2005)
R. De Robertis, P.T. Martini, E. Demozzi, F. Dal Corso, C. Bassi, P. Pederzoli et al., World J. Radiol. 7, 319 (2015)
Y. Itou, K. Nakanishi, Y. Narumi, Y. Nishizawa, H. Tsukuma, J. Magn. Reson. Imaging 33, 167–172 (2011)
H.K. Agarwal, F.V. Mertan, S. Sankineni, M. Bernardo, J. Senegas, J. Keupp et al., J. Magn. Reson. Imaging 45, 125–131 (2017)
G. Manenti, M. Nezzo, F. Chegai, E. Vasili, E. Bonanno, G. Simonetti, Prostate Cancer (2014). https://doi.org/10.1155/2014/868269
A. Wetter, F. Nensa, C. Lipponer, N. Guberina, T. Olbricht, M. Schenck et al., Acta Radiol. 56, 1009–1015 (2015)
R. Bourne, E. Panagiotaki, Diagnostics 6, 21 (2016)
A. Kamil, T. Shaikh, Literature Review of Generative models for Image-to-Image translation problems. 2019 International Conference on Computational Intelligence and Knowledge Economy (ICCIKE) (IEEE, 2019), p. 340–5.
Z. Shen, S. K. Zhou, Y. Chen, B. Georgescu, X. Liu, T. Huang, One-to-one Mapping for Unpaired Image-to-image Translation. Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision (2020), p. 1170–9.
H. Bickel, S.H. Polanec, G. Wengert, K. Pinker, W. Bogner, T.H. Helbich et al., J. Magn. Reson. Imaging 50, 1754–1761 (2019)
B.H. Choi, H.J. Baek, J.Y. Ha, K.H. Ryu, Moon J. Il, S.E. Park et al., Korean J. Radiol. 21, 1036 (2020)
C.-B. Jin, H. Kim, M. Liu, I.H. Han, Lee J. Il, J.H. Lee et al., Appl. Sci. 9, 2521 (2019)
K. Clark, B. Vendt, K. Smith, J. Freymann, J. Kirby, P. Koppel et al., J. Digit. Imaging 26, 1045–1057 (2013)
G. Litjens, O. Debats, J. Barentsz, N. Karssemeijer, H. Huisman, IEEE Trans. Med. Imaging 33, 1083–1092 (2014)
G. Litjens, O. Debats, J. Barentsz, N. Karssemeijer, H. Huisman. Cancer imaging archive wiki (2017). https://doi.org/10.7937/K9TCIA
Q. Yang, X. Li, BMC Bioinform. 22, 1–17 (2021)
A. Creswell, T. White, V. Dumoulin, K. Arulkumaran, B. Sengupta, A.A. Bharath, IEEE Signal Process. Mag. 35, 53–65 (2018)
P. Isola, J.-Y. Zhu, T. Zhou, A. A. Efros. Image-to-image translation with conditional adversarial networks. Proceedings of the IEEE conference on computer vision and pattern recognition (2017), p. 1125–1134.
J.-Y. Zhu, T. Park, P. Isola, A. A. Efros. Unpaired image-to-image translation using cycle-consistent adversarial networks. Proceedings of the IEEE international conference on computer vision (2017), p. 2223–2232.
C. Cavaro-Ménard, L. Zhang, P. Le Callet. Diagnostic quality assessment of medical images: Challenges and trends. 2010 2nd European Workshop on Visual Information Processing (EUVIP) (IEEE, 2010), p. 277–284.
L. Hu, D. Zhou, Y. Zha, L. Li, H. He, W. Xu et al., Radiology 3, e200237 (2021)
P. Sahoo, R.C. Rockne, A. Jung, P.K. Gupta, R.K.S. Rathore, R.K. Gupta, Prostate Cancer 2020, 5091218 (2020)
M.C. Maas, J.J. Fütterer, T.W.J. Scheenen, Invest. Radiol. 48, 779–786 (2013)
Acknowledgements
Not applicable.
Funding
This work was supported by the Ahvaz Jundishapur University of Medical Sciences under Grant [CRC-0024].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rezaeijo, S.M., Entezari Zarch, H., Mojtahedi, H. et al. Feasibility Study of Synthetic DW-MR Images with Different b Values Compared with Real DW-MR Images: Quantitative Assessment of Three Models Based-Deep Learning Including CycleGAN, Pix2PiX, and DC2Anet. Appl Magn Reson 53, 1407–1429 (2022). https://doi.org/10.1007/s00723-022-01482-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00723-022-01482-y