Abstract
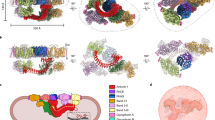
The cytoskeleton of a red blood cell (RBC) is anchored to the cell membrane by the ankyrin complex. This complex is assembled during RBC genesis and comprises primarily band 3, protein 4.2 and ankyrin, whose mutations contribute to numerous human inherited diseases. High-resolution structures of the ankyrin complex have been long sought-after to understand its assembly and disease-causing mutations. Here, we analyzed native complexes on the human RBC membrane by stepwise fractionation. Cryo-electron microscopy structures of nine band-3-associated complexes reveal that protein 4.2 stabilizes the cytoplasmic domain of band 3 dimer. In turn, the superhelix-shaped ankyrin binds to this protein 4.2 via ankyrin repeats (ARs) 6–13 and to another band 3 dimer via ARs 17–20, bridging two band 3 dimers in the ankyrin complex. Integration of these structures with both prior data and our biochemical data supports a model of ankyrin complex assembly during erythropoiesis and identifies interactions essential for the mechanical stability of RBC.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Cryo-EM density maps have been deposited in the Electron Microscopy Data Bank under accession numbers EMD-26148 (band 3 dimer), EMD-26145 (B2P1loose), EMD-26146 (B2P1vertical), EMD-26147 (B2P2vertical), EMD-26142 (B2P1diagonal), EMD-26143 (membrane part of B2P1diagonal), EMD-26144 (cytoplasmic part of B2P1diagonal), EMD-26149 (B2P1A1), EMD-26150 (cytoplasmic part of B2P1A1), EMD-26151 (B2P1A2), EMD-26152 (focused refinement of B2P1A2), EMD-26153 (B4P1A1) and EMD-26154 ((B2P1A1)2). Model coordinates have been deposited in the Protein Data Bank under accession numbers 7TW2 (band 3 dimer), 7TW0 (B2P1vertical), 7TW1 (B2P2vertical), 7TVZ (B2P1diagonal), 7TW3 (B2P1A1), 7TW5 (B2P1A2) and 7TW6 (B4P1A1). Other structures used in this study were retrieved from the PDB with accession codes 4YZF for the crystal structure of band 3 membrane domain, 1HYN for the crystal structure of band 3 cytoplasmic domain, 1N11 for ARs 13–24 of ankyrinR, 4RLV for ARs 1–24 of ankyrinB, 1L9N for transglutaminase 3 and 6KI1 for BicA. All other data needed to evaluate the conclusions in the paper are present in the paper and/or the supplementary materials. Source data are provided with this paper.
References
Shi, J. et al. Engineered red blood cells as carriers for systemic delivery of a wide array of functional probes. Proc. Natl Acad. Sci. USA 111, 10131–10136 (2014).
Bennett, V. & Baines, A. J. Spectrin and ankyrin-based pathways: metazoan inventions for integrating cells into tissues. Physiol. Rev. 81, 1353–1392 (2001).
Bennett, V. & Stenbuck, P. J. The membrane attachment protein for spectrin is associated with band 3 in human erythrocyte membranes. Nature 280, 468–473 (1979).
Mankelow, T. J., Satchwell, T. J. & Burton, N. M. Refined views of multi-protein complexes in the erythrocyte membrane. Blood Cells Mol. Dis. 49, 1–10 (2012).
Narla, J. & Mohandas, N. Red cell membrane disorders. Int J. Lab Hematol. 39(Suppl 1), 47–52 (2017).
Risinger, M. & Kalfa, T. A. Red cell membrane disorders: structure meets function. Blood 136, 1250–1261 (2020).
Korsgren, C. & Cohen, C. M. Associations of human erythrocyte band 4.2. Binding to ankyrin and to the cytoplasmic domain of band 3. J. Biol. Chem. 263, 10212–10218 (1988).
Kumpornsin, K., Jiemsup, S., Yongkiettrakul, S. & Chookajorn, T. Characterization of band 3–ankyrin–protein 4.2 complex by biochemical and mass spectrometry approaches. Biochem. Biophys. Res. Commun. 406, 332–335 (2011).
Davis, L., Lux, S. E. & Bennett, V. Mapping the ankyrin-binding site of the human erythrocyte anion exchanger. J. Biol. Chem. 264, 9665–9672 (1989).
Davis, L. H. & Bennett, V. Mapping the binding-sites of human erythrocyte ankyrin for the anion-exchanger and spectrin. J. Biol. Chem. 265, 10589–10596 (1990).
Kim, S. et al. Determination of structural models of the complex between the cytoplasmic domain of erythrocyte band 3 and ankyrin-R repeats 13–24. J. Biol. Chem. 286, 20746–20757 (2011).
Risinger, M. A., Dotimas, E. M. & Cohen, C. M. Human erythrocyte protein 4.2, a high copy number membrane protein, is N-myristylated. J. Biol. Chem. 267, 5680–5685 (1992).
Toye, A. M. et al. Protein-4.2 association with band 3 (AE1, SLCA4) in Xenopus oocytes: effects of three natural protein-4.2 mutations associated with hemolytic anemia. Blood 105, 4088–4095 (2005).
Korsgren, C., Lawler, J., Lambert, S., Speicher, D. & Cohen, C. M. Complete amino acid sequence and homologies of human erythrocyte membrane protein band 4.2. Proc. Natl Acad. Sci. USA 87, 613–617 (1990).
Zhang, D., Kiyatkin, A., Bolin, J. T. & Low, P. S. Crystallographic structure and functional interpretation of the cytoplasmic domain of erythrocyte membrane band 3. Blood 96, 2925–2933 (2000).
Arakawa, T. et al. Crystal structure of the anion exchanger domain of human erythrocyte band 3. Science 350, 680–684 (2015).
Michaely, P., Tomchick, D. R., Machius, M. & Anderson, R. G. Crystal structure of a 12 ANK repeat stack from human ankyrinR. EMBO J. 21, 6387–6396 (2002).
Stark, H. GraFix: stabilization of fragile macromolecular complexes for single particle cryo-EM. Methods Enzymol. 481, 109–126 (2010).
De Vecchis, D., Reithmeier, R. A. F. & Kalli, A. C. Molecular simulations of intact anion exchanger 1 reveal specific domain and lipid interactions. Biophys. J. 117, 1364–1379 (2019).
Wang, D. N. Band 3 protein: structure, flexibility and function. FEBS Lett. 346, 26–31 (1994).
Jiang, J. et al. Single particle electron microscopy analysis of the bovine anion exchanger 1 reveals a flexible linker connecting the cytoplasmic and membrane domains. PLoS ONE 8, e55408 (2013).
Romero, M. F., Chen, A. P., Parker, M. D. & Boron, W. F. The SLC4 family of bicarbonate (HCO3–) transporters. Mol. Asp. Med 34, 159–182 (2013).
Wang, W. et al. Cryo-EM structure of the sodium-driven chloride/bicarbonate exchanger NDCBE. Nat. Commun. 12, 5690 (2021).
Yu, X. et al. Dimeric structure of the uracil:proton symporter UraA provides mechanistic insights into the SLC4/23/26 transporters. Cell Res 27, 1020–1033 (2017).
Wang, C. et al. Structural mechanism of the active bicarbonate transporter from cyanobacteria. Nat. Plants 5, 1184–1193 (2019).
Muller-Berger, S. et al. Roles of histidine 752 and glutamate 699 in the pH dependence of mouse band 3 protein-mediated anion transport. Biochemistry 34, 9325–9332 (1995).
Chernova, M. N. et al. Electrogenic sulfate/chloride exchange in Xenopus oocytes mediated by murine AE1 E699Q. J. Gen. Physiol. 109, 345–360 (1997).
Karbach, D., Staub, M., Wood, P. G. & Passow, H. Effect of site-directed mutagenesis of the arginine residues 509 and 748 on mouse band 3 protein-mediated anion transport. Biochim. Biophys. Acta 1371, 114–122 (1998).
Bork, P., Holm, L. & Sander, C. The immunoglobulin fold. Structural classification, sequence patterns and common core. J. Mol. Biol. 242, 309–320 (1994).
Satchwell, T. J., Shoemark, D. K., Sessions, R. B. & Toye, A. M. Protein 4.2: a complex linker. Blood Cells Mol. Dis. 42, 201–210 (2009).
Ahvazi, B., Kim, H. C., Kee, S. H., Nemes, Z. & Steinert, P. M. Three-dimensional structure of the human transglutaminase 3 enzyme: binding of calcium ions changes structure for activation. EMBO J. 21, 2055–2067 (2002).
Yee, V. C. et al. Three-dimensional structure of a transglutaminase: human blood coagulation factor XIII. Proc. Natl Acad. Sci. USA 91, 7296–7300 (1994).
Azim, A. C. et al. Human erythrocyte dematin and protein 4.2 (pallidin) are ATP binding proteins. Biochemistry 35, 3001–3006 (1996).
Risinger, M. A., Dotimas, E. M. & Cohen, C. M. Human erythrocyte protein 4.2, a high copy number membrane-protein, Is N-myristylated. J. Biol. Chem. 267, 5680–5685 (1992).
Malik, S., Sami, M. & Watts, A. A role for band 4.2 in human erythrocyte band 3 mediated anion transport. Biochemistry 32, 10078–10084 (1993).
Steck, T. L. & Yu, J. Selective solubilization of proteins from red blood cell membranes by protein perturbants. J. Supramol. Struct. 1, 220–232 (1973).
Korsgren, C. & Cohen, C. M. Purification and properties of human erythrocyte band 4.2. Association with the cytoplasmic domain of band 3. J. Biol. Chem. 261, 5536–5543 (1986).
Rybicki, A. C. et al. Human erythrocyte protein 4.2 deficiency associated with hemolytic anemia and a homozygous 40glutamic acid → lysine substitution in the cytoplasmic domain of band 3 (band 3Montefiore). Blood 81, 2155–2165 (1993).
Inoue, T. et al. Homozygous missense mutation (band 3 Fukuoka: G130R): a mild form of hereditary spherocytosis with near-normal band 3 content and minimal changes of membrane ultrastructure despite moderate protein 4.2 deficiency. Br. J. Haematol. 102, 932–939 (1998).
Wang, C. et al. Structural basis of diverse membrane target recognitions by ankyrins. eLife 3, e04353 (2014).
Bhattacharyya, R. et al. Mapping of a palmitoylatable band 3-binding domain of human erythrocyte membrane protein 4.2. Biochem. J. 340, 505–512 (1999).
Grey, J. L., Kodippili, G. C., Simon, K. & Low, P. S. Identification of contact sites between ankyrin and band 3 in the human erythrocyte membrane. Biochemistry 51, 6838–6846 (2012).
Rybicki, A. C., Schwartz, R. S., Hustedt, E. J. & Cobb, C. E. Increased rotational mobility and extractability of band 3 from protein 4.2-deficient erythrocyte membranes: evidence of a role for protein 4.2 in strengthening the band 3-cytoskeleton linkage. Blood 88, 2745–2753 (1996).
van den Akker, E. et al. Investigating the key membrane protein changes during in vitro erythropoiesis of protein 4.2 (-) cells (mutations Chartres 1 and 2). Haematologica 95, 1278–1286 (2010).
Satchwell, T. J. et al. Severe Ankyrin-R deficiency results in impaired surface retention and lysosomal degradation of RhAG in human erythroblasts. Haematologica 101, 1018–1027 (2016).
Satchwell, T. J. et al. Critical band 3 multiprotein complex interactions establish early during human erythropoiesis. Blood 118, 182–191 (2011).
Burton, N. M. & Bruce, L. J. Modelling the structure of the red cell membrane. Biochem. Cell Biol. 89, 200–215 (2011).
Low, P. S. Structure and function of the cytoplasmic domain of band 3: center of erythrocyte membrane-peripheral protein interactions. Biochim. Biophys. Acta 864, 145–167 (1986).
Van Dort, H. M., Moriyama, R. & Low, P. S. Effect of band 3 subunit equilibrium on the kinetics and affinity of ankyrin binding to erythrocyte membrane vesicles. J. Biol. Chem. 273, 14819–14826 (1998).
Yi, S. J. et al. Red cell membranes of ankyrin-deficient nb/nb mice lack band 3 tetramers but contain normal membrane skeletons. Biochemistry 36, 9596–9604 (1997).
Ipsaro, J. J. & Mondragon, A. Structural basis for spectrin recognition by ankyrin. Blood 115, 4093–4101 (2010).
Bruce, L. J. et al. A band 3-based macrocomplex of integral and peripheral proteins in the RBC membrane. Blood 101, 4180–4188 (2003).
Ho, C. M. et al. Malaria parasite translocon structure and mechanism of effector export. Nature 561, 70–75 (2018).
Liu, S. et al. Structure of the yeast spliceosomal postcatalytic P complex. Science 358, 1278–1283 (2017).
Yan, C., Wan, R., Bai, R., Huang, G. & Shi, Y. Structure of a yeast step II catalytically activated spliceosome. Science 355, 149–155 (2017).
Fitzpatrick, A. W. P. et al. Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 547, 185–190 (2017).
Bennett, V. Isolation of an ankyrin-band 3 oligomer from human erythrocyte membranes. Biochim. Biophys. Acta 689, 475–484 (1982).
Liu, X., Li, M., Xia, X., Li, X. & Chen, Z. Mechanism of chromatin remodelling revealed by the Snf2-nucleosome structure. Nature 544, 440–445 (2017).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. Processing of structurally heterogeneous cryo-EM data in RELION. Methods Enzymol. 579, 125–157 (2016).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Wang, N. et al. Structural basis of human monocarboxylate transporter 1 inhibition by anti-cancer drug candidates. Cell 184, 370–383 e13 (2021).
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr D. Biol. Crystallogr 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D. Biol. Crystallogr 66, 213–221 (2010).
Madeira, F. et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 47, W636–W641 (2019).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324 (2014).
Acknowledgements
We thank T. Nguyen, J. Zhen and A. Stevens for editorial assistance. This project is supported by grants from the US NIH (R01GM071940 to Z.H.Z.). We acknowledge the use of resources at the Electron Imaging Center for Nanomachines supported by UCLA and grants from the NIH (1S10OD018111 and 1U24GM116792) and the National Science Foundation (DBI-1338135 and DMR-1548924).
Author information
Authors and Affiliations
Contributions
Z.H.Z. conceived the project. X.X. and S.L. prepared samples and acquired and analyzed cryo-EM data. X.X. engineered and isolated the recombinant proteins and performed biochemistry analyses. S.L. and X.X. built the models. X.X., S.L. and Z.H.Z. interpreted the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural and Molecular Biology thanks Ash Toye, Yifan Cheng, and Werner KÃhlbrandt for their contribution to the peer review of this work. Primary Handling editor: Florian Ullrich, in collaboration with the Nature Structural & Molecular Biology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Purification and cryo-EM reconstruction of the erythrocyte membrane proteins.
(a) Workflow of the stepwise fractionation of erythrocyte membrane proteins. (b) The second gel-filtration chromatography profile of the low-salt fraction. The result from SDS-PAGE analysis of the peak fractions is inserted in the upper left corner. The peak fractions were applied to SDS-PAGE and visualized by Coomassie blue staining. Dashed blue box on the gel and blue bar on the chromatogram indicate the fractions collected for cryo-EM. (c) The first gel-filtration chromatography profile of the high-salt fraction. Green and magenta boxes on the gel and green and magenta bars on the chromatogram indicate the fractions collected for the protein 4.2 complex and ankyrin complex, respectively. (d) The second gel-filtration chromatography profile and corresponding gel of the protein 4.2 complex. Dashed green box on the gel and green bar on the chromatogram indicate the fractions collected for cryo-EM. (e) The gel-filtration chromatography profile of the ankyrin complex after Grafix purification. Magenta bar on the chromatogram indicates the fractions collected for cryo-EM.
Extended Data Fig. 2 Cryo-EM analysis of the low-salt fraction (band 3).
(a) Representative cryo-EM image of the low-salt fraction from 9455 images collected. (b) Selected 2D class averages of the cryo-EM particle images. (c) Flow chart of cryo-EM data processing. (d) Gold-standard Fourier shell correlation (FSC) curve for 3D reconstruction. (e) Angular distribution of cryo-EM reconstructions used for final refinement. (f) Density of the membrane domain. (g) Atomic model of band 3 cytoplasmic domain fitted into the cryo-EM density. A lower map threshold is used in (g) compared to that of (f) to better present the cytoplasmic domain.
Extended Data Fig. 3 Image processing for the cryo-EM data of the high-salt fraction 1 (band 3-protein 4.2 complex).
(a) Representative cryo-EM image of the high-salt fraction 1 from 20842 images collected. (b) Selected 2D class averages of the cryo-EM particle images. (c) Flow chart of cryo-EM data processing. (d), Gold-standard Fourier shell correlation (FSC) curves for 3D reconstructions. (e) Local resolution of the overall map of B2P1diagonal complex. (f) Angular distribution of cryo-EM reconstruction of B2P1diagonal complex used for final refinement. (g-h) Local resolutions of the focused refinement maps of the cytoplasmic part and membrane part of B2P1diagonal complex. (i) Representative cryo-EM density maps of the B2P1diagonal complex.
Extended Data Fig. 4 Cryo-EM analysis of the high-salt fraction 2 (ankyrin complex).
(a) Representative cryo-EM image of the high-salt fraction 2 from 21187 images collected. (b) Selected 2D class averages of cryo-EM particle images. (c) Flow chart of cryo-EM data processing. (d) Gold-standard Fourier shell correlation (FSC) curves for 3D reconstructions. (e) Local resolution of the overall map of B2P1A1 complex. (f) Angular distribution of cryo-EM reconstruction of B2P1A1 complex. (g) Representative cryo-EM density maps of the B2P1A1 complex showing the fragments of ankyrin and protein 4.2 at their binding interface. (h) Representative cryo-EM density maps of the B2P1A2 complex showing the fragments of ankyrin and band 3 at their binding interface.
Extended Data Fig. 5 Structural analysis of the band 3 membrane domain.
(a) Superposition of the band 3 membrane domain in B2P1diagonal complex and reported crystal structure (PDB: 4YZF)16. (b) Topology of the transmembrane helices of band 3. (c) Density of the DDM molecule at the interface of the core and gate domain. (d) Enlarged view of the substrate binding site in B2P1diagonal complex. Four water molecules were tentatively modelled into the cryo-EM density of band 3 near the substrate binding site. (e) Comparison of the substrate binding site in band 3 with that in bicarbonate transporter BicA (PDB: 6KI1)25.
Extended Data Fig. 6 Sequence alignment of band 3 from different species.
Sequences of human band 3 (P02730), mouse band 3 (P04919), rabbit band 3 (G1SLY0), bovine band 3 (Q9XSW5), horse band 3 (Q2Z1P9), chicken band 3 (P15575), frog band 3 (F6XSL8) and fish band 3 (Q7ZZJ7). The sequence alignment is done using the Clustal Omega server72; the figure is generated by ESPript 373. Cyan triangles represent the band 3 residues interacting with protein 4.2; magenta bars indicate the regions of band 3 interacting with ankyrin; reported disease mutations on human band 3 are labeled as red circles.
Extended Data Fig. 7 Structure of protein 4.2.
(a) Structure of the core domain shown in ribbon. Residue numbers of its N and C terminus are labeled. (b) Superposition of protein 4.2 with transglutaminase (gray, PDB: 1L9N)31, showing the missing catalytic triad in protein 4.2. (c-e) Structures of the three Ig-like domains and illustrations of their secondary structure. (f) The electrostatic surface of protein 4.2, showing its membrane binding site (blue dashed circle) and band 3 binding interface (orange dashed circle). (g-h) Sequence conservation of protein 4.2 among mammals mapped to the structure. Orientation in (g) is the same as that in (f). Orange dashed circle shows the band 3 binding interface; magenta dashed box shows the ankyrin binding interface.
Extended Data Fig. 8 Sequence alignment of protein 4.2 from different species.
Sequences of human protein 4.2 (P16452), mouse protein 4.2 (P49222), rabbit protein (G1TDR3), bovine protein 4.2 (O46510), horse protein 4.2 (F6ZDW1), chicken protein 4.2 (E1BQZ4), frog protein 4.2 (XP_018090678.1) and human transglutaminase 3 (Q08188). The sequence alignment is done using the Clustal Omega server72; the figure is generated in ESPript 373. Salmon triangles represent protein 4.2 residues interacting with band 3; magenta bars indicate the regions of protein 4.2 interacting with ankyrin; reported disease mutations on human protein 4.2 are labeled as red circles; black stars indicate the catalytic residues of human transglutaminase 3.
Extended Data Fig. 9 Conformational changes of band 3 and protein 4.2 during the assembly process.
(a, b) Density map of the B2P1vertical complex and B2P1diagonal complex. Red box indicates the anchorage site of protein 4.2 N-termini to the membrane. (c) Rotation of protein 4.2 (red arrow) from vertical (transparent grey surface) to diagonal conformation (green surface). The two complexes are superposed according to the membrane domains of band 3. The cytoplasmic domains of band 3 are omitted for clarity. Angles between the membrane (grey bar) and protein 4.2 are labeled. (d-e) Rotation of the cytoplasmic domain of band 3 (red arrow) from B2P1vertical to B2P1diagonal complex viewed from the cytoplasmic side. The membrane domain of band 3 is shown as transparent surface and cytoplasmic domain as ribbon. Protein 4.2 is indicated as a green oval for clarity. (f) Density map of the B2P1diagonal complex sharpened with B-factor of −50 Å2 showing the interaction of the CM-linker with protein 4.2. (g-h) Ribbon representation of the CM-linker region. Residues of protein 4.2 interacting with the CM-linker are labeled.
Extended Data Fig. 10 Analysis of the ankyrin-containing complexes.
(a) Atomic model of B4P1A1 complex in ribbon superposed with the density map at a low threshold. Red arrow indicates the density of the membrane domain of the second band 3 dimer. (b) Analytical gel filtration assay showing the assembly of B4P1A1 complex in vitro. Dashed boxes show the position of B4P1A1 complex. Experiments were repeated for two times with similar results. (c) Atomic model of B2P1A2 complex in ribbon superposed with the density map at a low threshold. The density for the second band 3 dimer is indicated by arrow. (d) Analytical gel filtration assay showing the assembly of B2P1A2 complex in vitro. Dashed boxes show the position of B2P1A2 complex. The gel-filtration and SDS-PAGE results of protein 4.2 complex, band 3 cytoplasmic domain and ARs 1–24 are the same as that in (b). Second band 3 dimer may be incorporated into this complex, resulting in the B4P1A2. Experiments were repeated for two times with similar results. (e) Atomic model of (B2P1A1)2 complex in ribbon superposed with the density map. Boxes show the position of individual B2P1A1 complexes. (f) Reported disease mutations mapped on the structure of B4P1A1 complex.
Supplementary information
Source data
Source Data Fig. 4
Uncropped gels for Fig. 4c
Source Data Extended Data Fig. 10
Uncropped gels for Extended Data Fig. 10
Rights and permissions
About this article
Cite this article
Xia, X., Liu, S. & Zhou, Z.H. Structure, dynamics and assembly of the ankyrin complex on human red blood cell membrane. Nat Struct Mol Biol 29, 698–705 (2022). https://doi.org/10.1038/s41594-022-00779-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00779-7
This article is cited by
-
Structural and functional insights into the lipid regulation of human anion exchanger 2
Nature Communications (2024)
-
Substrate binding and inhibition of the anion exchanger 1 transporter
Nature Structural & Molecular Biology (2023)
-
Structural insights into the conformational changes of BTR1/SLC4A11 in complex with PIP2
Nature Communications (2023)
-
The structural basis of the pH-homeostasis mediated by the Cl−/HCO3− exchanger, AE2
Nature Communications (2023)
-
The band 3–ankyrin multiprotein complex comes in from the cold
Nature Structural & Molecular Biology (2022)