Abstract
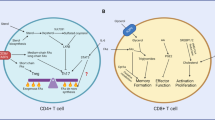
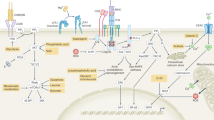
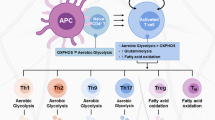
T cells orchestrate adaptive immunity against pathogens and other immune challenges, but their dysfunction can also mediate the pathogenesis of cancer and autoimmunity. Metabolic adaptation in response to immunological and microenvironmental signals contributes to T cell function and fate decision. Lipid metabolism has emerged as a key regulator of T cell responses, with selective lipid metabolites serving as metabolic rheostats to integrate environmental cues and interplay with intracellular signaling processes. Here, we discuss how extracellular, de novo synthesized and membrane lipids orchestrate T cell biology. We also describe the roles of lipids as regulators of intracellular signaling at the levels of transcriptional, epigenetic and post-translational regulation in T cells. Finally, we summarize therapeutic targeting of lipid metabolism and signaling, and conclude with a discussion of important future directions. Understanding the molecular and functional interplay between lipid metabolism and T cell biology will ultimately inform therapeutic intervention for human disease.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Chapman, N. M. & Chi, H. Metabolic adaptation of lymphocytes in immunity and disease. Immunity 55, 14–30 (2022).
Maseda, D., Ricciotti, E. & Crofford, L. J. Prostaglandin regulation of T cell biology. Pharmacol. Res. 149, 104456 (2019).
Trompette, A. et al. Dietary fiber confers protection against flu by shaping Ly6c− patrolling monocyte hematopoiesis and CD8+ T cell metabolism. Immunity 48, 992–1005 e1008 (2018).
Bachem, A. et al. Microbiota-derived short-chain fatty acids promote the memory potential of antigen-activated CD8+ T cells. Immunity 51, 285–297.e285 (2019).
Dudek, M. et al. Auto-aggressive CXCR6+ CD8 T cells cause liver immune pathology in NASH. Nature 592, 444–449 (2021). This paper shows a pathogenic role for the SCFA acetate in CD8+ T cell-mediated autoinflammation and tissue damage in the liver, which is in contrast to the protective roles of SCFAs in tissue homeostasis or anti-pathogen immunity.
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013).
Smith, P. M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
Park, J. et al. Short-chain fatty acids induce both effector and regulatory T cells by suppression of histone deacetylases and regulation of the mTOR-S6K pathway. Mucosal Immunol. 8, 80–93 (2015).
Haghikia, A. et al. Dietary fatty acids directly impact central nervous system autoimmunity via the small intestine. Immunity 43, 817–829 (2015).
Wang, H. et al. CD36-mediated metabolic adaptation supports regulatory T cell survival and function in tumors. Nat. Immunol. 21, 298–308 (2020). This study establishes a role for CD36-dependent lipid uptake in supporting Treg cell function in the TME.
Xu, S. et al. Uptake of oxidized lipids by the scavenger receptor CD36 promotes lipid peroxidation and dysfunction in CD8+ T cells in tumors. Immunity 54, 1561–1577 e1567 (2021). This study, along with ref. 13, shows that CD36-mediated lipid uptake triggers CD8+ T cell dysfunction in the TME.
Ma, X. et al. CD36-mediated ferroptosis dampens intratumoral CD8+ T cell effector function and impairs their antitumor ability. Cell Metab. 33, 1001–1012 e1005 (2021). This study, along with ref. 12, reveals that CD36-mediated lipid uptake triggers CD8+ T cell dysfunction in the TME.
Manzo, T. et al. Accumulation of long-chain fatty acids in the tumor microenvironment drives dysfunction in intrapancreatic CD8+ T cells. J. Exp. Med. 217, e20191920 (2020).
Wu, L. et al. Niche-selective inhibition of pathogenic Th17 cells by targeting metabolic redundancy. Cell 182, 641–654.e620 (2020).
Proto, J. D. et al. Hypercholesterolemia induces T cell expansion in humanized immune mice. J. Clin. Invest. 128, 2370–2375 (2018).
Ma, X. et al. Cholesterol induces CD8+ T cell exhaustion in the tumor microenvironment. Cell Metab. 30, 143–156.e145 (2019).
Ma, X. et al. Cholesterol negatively regulates IL-9-producing CD8+ T cell differentiation and antitumor activity. J. Exp. Med. 215, 1555–1569 (2018).
Li, J., Lu, E., Yi, T. & Cyster, J. G. EBI2 augments Tfh cell fate by promoting interaction with IL-2-quenching dendritic cells. Nature 533, 110–114 (2016).
Yuan, J. et al. Potentiating CD8(+) T cell antitumor activity by inhibiting PCSK9 to promote LDLR-mediated TCR recycling and signaling. Protein Cell 12, 240–260 (2021).
Hang, S. et al. Bile acid metabolites control TH17 and Treg cell differentiation. Nature 576, 143–148 (2019). This study establishes that selective bile acids balance intestinal TH17 and Treg cell differentiation, which contributes to tissue homeostasis.
Song, X. et al. Microbial bile acid metabolites modulate gut RORγ+ regulatory T cell homeostasis. Nature 577, 410–415 (2020).
Li, W. et al. A bacterial bile acid metabolite modulates Treg activity through the nuclear hormone receptor NR4A1. Cell Host Microbe 29, 1366–1377 e1369 (2021).
Campbell, C. et al. Bacterial metabolism of bile acids promotes generation of peripheral regulatory T cells. Nature 581, 475–479 (2020).
Chen, M. L. et al. CAR directs T cell adaptation to bile acids in the small intestine. Nature 593, 147–151 (2021).
Kidani, Y. et al. Sterol regulatory element-binding proteins are essential for the metabolic programming of effector T cells and adaptive immunity. Nat. Immunol. 14, 489–499 (2013).
Su, W. et al. Protein prenylation drives discrete signaling programs for the differentiation and maintenance of effector Treg cells. Cell Metab. 32, 996–1011.e1017 (2020). This paper demonstrates that mevalonate metabolism-dependent post-translational modifications are essential for Treg cell activation and establishment of immunological tolerance.
Klein Geltink, R. I. et al. Mitochondrial priming by CD28. Cell 171, 385–397 e311 (2017).
Chapman, N. M., Boothby, M. R. & Chi, H. Metabolic coordination of T cell quiescence and activation. Nat. Rev. Immunol. 20, 55–70 (2020).
Yang, K. et al. T cell exit from quiescence and differentiation into Th2 cells depend on Raptor-mTORC1-mediated metabolic reprogramming. Immunity 39, 1043–1056 (2013).
Berod, L. et al. De novo fatty acid synthesis controls the fate between regulatory T and T helper 17 cells. Nat. Med. 20, 1327–1333 (2014).
Endo, Y. et al. Obesity drives Th17 cell differentiation by inducing the lipid metabolic kinase, ACC1. Cell Rep. 12, 1042–1055 (2015).
Endo, Y. et al. ACC1 determines memory potential of individual CD4+ T cells by regulating de novo fatty acid biosynthesis. Nat. Metab. 1, 261–275 (2019).
Lee, J. et al. Regulator of fatty acid metabolism, acetyl coenzyme a carboxylase 1, controls T cell immunity. J. Immunol. 192, 3190–3199 (2014).
Lim, S. A. et al. Lipid signalling enforces functional specialization of Treg cells in tumours. Nature 591, 306–311 (2021). This study establishes a relationship between lipid synthesis and PD-1 immune checkpoint signaling in Treg cell-mediated suppression of antitumor immunity.
Tian, M. et al. ACLY ubiquitination by CUL3-KLHL25 induces the reprogramming of fatty acid metabolism to facilitate iTreg differentiation. eLife 10, e62394 (2021).
Sofi, M. H. et al. Ceramide synthesis regulates T cell activity and GVHD development. JCI Insight 2, e91701 (2017).
Vaena, S. et al. Aging-dependent mitochondrial dysfunction mediated by ceramide signaling inhibits antitumor T cell response. Cell Rep. 35, 109076 (2021).
Li, G. et al. Nanoliposome C6-ceramide increases the anti-tumor immune response and slows growth of liver tumors in mice. Gastroenterology 154, 1024–1036 e1029 (2018).
Lacher, S. M. et al. HMG-CoA reductase promotes protein prenylation and therefore is indispensible for T-cell survival. Cell Death Dis. 8, e2824 (2017).
Timilshina, M. et al. Activation of mevalonate pathway via LKB1 is essential for stability of Treg cells. Cell Rep. 27, 2948–2961.e2947 (2019).
Yang, K. et al. Homeostatic control of metabolic and functional fitness of Treg cells by LKB1 signalling. Nature 548, 602–606 (2017).
Zeng, H. et al. mTORC1 couples immune signals and metabolic programming to establish Treg-cell function. Nature 499, 485–490 (2013).
Karmaus, P. W. F. et al. Metabolic heterogeneity underlies reciprocal fates of TH17 cell stemness and plasticity. Nature 565, 101–105 (2019).
Pearce, E. L. et al. Enhancing CD8 T-cell memory by modulating fatty acid metabolism. Nature 460, 103–107 (2009).
Borges da Silva, H. et al. The purinergic receptor P2RX7 directs metabolic fitness of long-lived memory CD8+ T cells. Nature 559, 264–268 (2018).
O’Sullivan, D. et al. Memory CD8+ T cells use cell-intrinsic lipolysis to support the metabolic programming necessary for development. Immunity 41, 75–88 (2014).
van der Windt, G. J. et al. Mitochondrial respiratory capacity is a critical regulator of CD8+ T cell memory development. Immunity 36, 68–78 (2012).
Cui, G. et al. IL-7-induced glycerol transport and TAG synthesis promotes memory CD8+ T cell longevity. Cell 161, 750–761 (2015).
Raud, B. et al. Etomoxir actions on regulatory and memory T cells are independent of cpt1a-mediated fatty acid oxidation. Cell Metab. 28, 504–515 e507 (2018).
Zhang, C. et al. STAT3 activation-induced fatty acid oxidation in CD8+ T effector cells is critical for obesity-promoted breast tumor growth. Cell Metab. 31, 148–161.e145 (2020).
Ringel, A. E. et al. Obesity shapes metabolism in the tumor microenvironment to suppress anti-tumor immunity. Cell 183, 1848–1866 e1826 (2020).
Pan, Y. et al. Survival of tissue-resident memory T cells requires exogenous lipid uptake and metabolism. Nature 543, 252–256 (2017). This paper establishes that fatty acid binding proteins are essential for regulating TRM cell accumulation in the skin, pointing to a role for extracellular fatty acid uptake and oxidization in orchestrating TRM cell persistence.
Frizzell, H. Organ-specific isoform selection of fatty acid-binding proteins in tissue-resident lymphocytes. Sci. Immunol. 5, eaay9283 (2020).
Lin, R. et al. Fatty acid oxidation controls CD8+ tissue-resident memory T-cell survival in gastric adenocarcinoma. Cancer Immunol. Res. 8, 479–492 (2020).
Field, C. S. et al. Mitochondrial integrity regulated by lipid metabolism is a cell-intrinsic checkpoint for Treg suppressive function. Cell Metab. 31, 422–437 e425 (2020).
Ecker, C. et al. Differential reliance on lipid metabolism as a salvage pathway underlies functional differences of T cell subsets in poor nutrient environments. Cell Rep. 23, 741–755 (2018).
Howie, D. et al. A novel role for triglyceride metabolism in Foxp3 expression. Front. Immunol. 10, 1860 (2019).
Liu, X. et al. Reprogramming lipid metabolism prevents effector T cell senescence and enhances tumor immunotherapy. Sci. Transl. Med. 13, eaaz6314 (2021).
Shin, J., O’Brien, T. F., Grayson, J. M. & Zhong, X. P. Differential regulation of primary and memory CD8 T cell immune responses by diacylglycerol kinases. J. Immunol. 188, 2111–2117 (2012).
Yang, W. et al. Potentiating the antitumour response of CD8+ T cells by modulating cholesterol metabolism. Nature 531, 651–655 (2016).
Wang, F., Beck-Garcia, K., Zorzin, C., Schamel, W. W. & Davis, M. M. Inhibition of T cell receptor signaling by cholesterol sulfate, a naturally occurring derivative of membrane cholesterol. Nat. Immunol. 17, 844–850 (2016).
Fu, G. et al. Metabolic control of TFH cells and humoral immunity by phosphatidylethanolamine. Nature 595, 724–729 (2021). This study establishes a context-specific role for de novo phosphatidylethanolamine synthesis in driving TFH cell generation and humoral immunity.
Corrado, M. et al. Dynamic cardiolipin synthesis is required for CD8+ T cell immunity. Cell Metab. 32, 981–995.e987 (2020).
Howie, D., Ten Bokum, A., Necula, A. S., Cobbold, S. P. & Waldmann, H. The role of lipid metabolism in T lymphocyte differentiation and survival. Front. Immunol. 8, 1949 (2017).
Cipolletta, D. et al. PPAR-γ is a major driver of the accumulation and phenotype of adipose tissue Treg cells. Nature 486, 549–553 (2012).
Bensinger, S. J. et al. LXR signaling couples sterol metabolism to proliferation in the acquired immune response. Cell 134, 97–111 (2008).
Xu, J., Wagoner, G., Douglas, J. C. & Drew, P. D. Liver X receptor agonist regulation of Th17 lymphocyte function in autoimmunity. J. Leukoc. Biol. 86, 401–409 (2009).
Vigne, S. et al. IL-27-induced Type 1 regulatory T-cells produce oxysterols that constrain IL-10 production. Front. Immunol. 8, 1184 (2017).
Qiu, J. et al. Acetate promotes T cell effector function during glucose restriction. Cell Rep. 27, 2063–2074.e2065 (2019).
Balmer, M. L. et al. Memory CD8+ T cells require increased concentrations of acetate induced by stress for optimal function. Immunity 44, 1312–1324 (2016).
Luu, M. et al. Regulation of the effector function of CD8+ T cells by gut microbiota-derived metabolite butyrate. Sci. Rep. 8, 14430 (2018).
Bailis, W. et al. Distinct modes of mitochondrial metabolism uncouple T cell differentiation and function. Nature 571, 403–407 (2019).
Thurnher, M. & Gruenbacher, G. T lymphocyte regulation by mevalonate metabolism. Sci. Signal. 8, re4 (2015).
Morrison, E. et al. Dynamic palmitoylation events following T-cell receptor signaling. Commun. Biol. 3, 368 (2020).
Wen, Z. et al. N-myristoyltransferase deficiency impairs activation of kinase AMPK and promotes synovial tissue inflammation. Nat. Immunol. 20, 313–325 (2019).
Saravia, J. et al. Homeostasis and transitional activation of regulatory T cells require c-Myc. Sci. Adv. 6, eaaw6443 (2020).
Bu, D. X. et al. Statin-induced Kruppel-like factor 2 expression in human and mouse T cells reduces inflammatory and pathogenic responses. J. Clin. Invest. 120, 1961–1970 (2010).
Ibitokou, S. A. et al. Early inhibition of fatty acid synthesis reduces generation of memory precursor effector T cells in chronic infection. J. Immunol. 200, 643–656 (2018).
Voss, K., Luthers, C. R., Pohida, K. & Snow, A. L. Fatty acid synthase contributes to restimulation-induced cell death of human CD4 T cells. Front. Mol. Biosci. 6, 106 (2019).
Wohlfert, E. A., Nichols, F. C., Nevius, E. & Clark, R. B. Peroxisome proliferator-activated receptor γ (PPARγ) and immunoregulation: enhancement of regulatory T cells through PPARγ-dependent and -independent mechanisms. J. Immunol. 178, 4129–4135 (2007).
Gocke, A. R. et al. Transcriptional modulation of the immune response by peroxisome proliferator-activated receptor- α agonists in autoimmune disease. J. Immunol. 182, 4479–4487 (2009).
Polak, P. E. et al. Protective effects of a peroxisome proliferator-activated receptor- β/δ agonist in experimental autoimmune encephalomyelitis. J. Neuroimmunol. 168, 65–75 (2005).
Marino, E. et al. Gut microbial metabolites limit the frequency of autoimmune T cells and protect against type 1 diabetes. Nat. Immunol. 18, 552–562 (2017).
Pompura, S. L. et al. Oleic acid restores suppressive defects in tissue-resident FOXP3 Tregs from patients with multiple sclerosis. J. Clin. Invest. 131, e138519 (2011).
Mathewson, N. D. et al. Gut microbiome-derived metabolites modulate intestinal epithelial cell damage and mitigate graft-versus-host disease. Nat. Immunol. 17, 505–513 (2016).
Gopalakrishnan, V. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359, 97–103 (2018).
Matson, V. et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108 (2018).
Spencer, C. N. et al. Dietary fiber and probiotics influence the gut microbiome and melanoma immunotherapy response. Science 374, 1632–1640 (2021). This study suggests that high dietary fiber intake improves responsiveness to immunotherapy via interplay with microbiota-derived production of the SCFA propionate.
Longo, J., van Leeuwen, J. E., Elbaz, M., Branchard, E. & Penn, L. Z. Statins as anticancer agents in the era of precision medicine. Clin. Cancer Res. 26, 5791–5800 (2020).
Calle, R. A. et al. ACC inhibitor alone or co-administered with a DGAT2 inhibitor in patients with non-alcoholic fatty liver disease: two parallel, placebo-controlled, randomized phase 2a trials. Nat. Med. 27, 1836–1848 (2021).
Pecin, I. & Reiner, Z. Novel experimental agents for the treatment of hypercholesterolemia. J. Exp. Pharm. 13, 91–100 (2021).
Payandeh, J. & Volgraf, M. Ligand binding at the protein–lipid interface: strategic considerations for drug design. Nat. Rev. Drug Discov. 20, 710–722 (2021).
Lopes, N. et al. Distinct metabolic programs established in the thymus control effector functions of γδ T cell subsets in tumor microenvironments. Nat. Immunol. 22, 179–192 (2021).
Kobayashi, T. et al. Increased lipid metabolism impairs NK cell function and mediates adaptation to the lymphoma environment. Blood 136, 3004–3017 (2020).
Michelet, X. et al. Metabolic reprogramming of natural killer cells in obesity limits antitumor responses. Nat. Immunol. 19, 1330–1340 (2018).
Fu, S. et al. Impaired lipid biosynthesis hinders anti-tumor efficacy of intratumoral iNKT cells. Nat. Commun. 11, 438 (2020).
Ko, J. S. et al. Palmitate inhibits arthritis by inducing t-bet and gata-3 mRNA degradation in iNKT cells via IRE1α-dependent decay. Sci. Rep. 7, 14940 (2017).
Muri, J., Thut, H., Bornkamm, G. W. & Kopf, M. B1 and marginal zone B cells but not follicular B2 cells require Gpx4 to prevent lipid peroxidation and ferroptosis. Cell Rep. 29, 2731–2744.e2734 (2019).
Bibby, J. A. et al. Cholesterol metabolism drives regulatory B cell IL-10 through provision of geranylgeranyl pyrophosphate. Nat. Commun. 11, 3412 (2020).
Acknowledgements
This work was supported by US National Institutes of Health grants AI105887, AI131703, AI140761, AI150241, AI150514 and CA253188, Alliance for Lupus Research grant, and ALSAC (to H.C.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
S.A.L. and W.S. conceived, designed and wrote the manuscript. N.M.C. co-wrote the manuscript. H.C. co-wrote the manuscript and provided overall direction.
Corresponding author
Ethics declarations
Competing interests
H.C. is a consultant for Kumquat Biosciences.
Peer review
Peer review information
Nature Chemical Biology thanks Ping-Chih Ho and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lim, S.A., Su, W., Chapman, N.M. et al. Lipid metabolism in T cell signaling and function. Nat Chem Biol 18, 470–481 (2022). https://doi.org/10.1038/s41589-022-01017-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-022-01017-3
This article is cited by
-
Cancer immunometabolism: advent, challenges, and perspective
Molecular Cancer (2024)
-
Nanoparticles in tumor microenvironment remodeling and cancer immunotherapy
Journal of Hematology & Oncology (2024)
-
Landscape of unconventional γδ T cell subsets in cancer
Molecular Biology Reports (2024)
-
Lipid metabolism and its implications in tumor cell plasticity and drug resistance: what we learned thus far?
Cancer and Metastasis Reviews (2024)
-
Discovery of a novel lipid metabolism-related gene signature to predict outcomes and the tumor immune microenvironment in gastric cancer by integrated analysis of single-cell and bulk RNA sequencing
Lipids in Health and Disease (2023)