Abstract
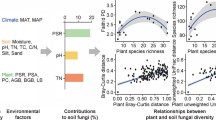
We collected solonchaks samples of 5 subtypes and 13 genera from Gansu Province in China and analyzed the diversity of fungal communities with the methods of ITS sequencing. We found that ACE, Chao 1, Simpson and Shannon indices of the fungal community varied significantly in soil samples and were generally inversely correlated with pH. The dominant fungal groups were Ascomycota and Basidiomycota among the different solonchaks. Redundancy analysis revealed that soil organic matters was the most important driving force for fungal composition (29.9%), and the second most influencing factor was sulfate ion (9.1%). Our results suggest that Ascomycota was the main indicator phylum reflecting changes of the microbial groups and soil organic matter as a key factor drove the composition of fungal community in 5 subtypes and 13 genera of solonchaks in Gansu Province.





Similar content being viewed by others
REFERENCES
S. D. Bao, Soil Agricultural Chemistry Analysis (China Agriculture Press, Beijing, 2000) [in Chinese].
R. D. Bardgett and W. H. van der Putten, “Belowground biodiversity and ecosystem functioning,” Nature 515, 505–511 (2014). https://doi.org/10.1038/nature13855
N. A. Bokulich, S. Subramanian, J. J. Faith, et al., “Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing,” Nat. Methods 10, 57–59 (2012). https://doi.org/10.1038/nmeth.2276
G. Bradfield, P. Somerfield, T. Meyn, M. Holby, D. Babcock, and B. I. H. Segel, “Regulation of sulfate transport in filamentous fungi,” Plant Physiol. 46, 720–727 (1970). https://doi.org/10.2307/4262253
M. Buée, M. Reich, C. Murat, et al., “454 Pyrosequencing analyses of forest soils reveal an unexpectedly high fungal diversity,” New Phytol. 184, 449–456 (2010). https://doi.org/10.1111/j.1469-8137.2009.03003.x
J. G. Caporaso, J. Kuczynski, J. Stombaugh, et al., “QIIME allows analysis of high-throughput community sequencing data,” Nat. Methods 7, 335–336 (2010).
Z. Cheng, Y. Chen, and F. Zhang, “Effect of reclamation of abandoned salinized farmland on soil bacterial communities in arid northwest China,” Sci. Total Environ. 630, 799–808 (2018).
E. Coller, A. Cestaro, R. Zanzotti, et al., “Microbiome of vineyard soils is shaped by geography and management,” Microbiome 7, 140 (2019). https://doi.org/10.1186/s40168-019-0758-7
A. M. Cupples, “Principles and applications of soil microbiology,” J. Environ. Qual. 34, 11–68 (2005). https://doi.org/10.2134/jeq2005.0731dup
B. W. De, L. B. Folman, R. C. Summerbell, and L. Boddy, “Living in a fungal world: impact of fungi on soil bacterial niche development,” FEMS Microbiol. Rev. 4, 795–811 (2005). https://doi.org/10.1016/j.femsre.2004.11.005
R. E. Drenovsky, D. Vo, K. J. Graham, and K. M. Scow, “Soil water content and organic carbon availability are major determinants of soil microbial community composition,” Microb. Ecol. 48, 424–430 (2004). https://doi.org/10.1007/s00248-003-1063-2
R. C. Edgar, “MUSCLE: multiple sequence alignment with high accuracy and high throughput,” Nucleic Acids Res. 32, 1792–1797 (2004). https://doi.org/10.2460/ajvr.69.1.82
R. C. Edgar, B. J. Haas, J. C. Clemente, C. Quince, and R. Knight, “UCHIME improves sensitivity and speed of chimera detection,” Bioinformatics 27 (16), 2194–2200 (2011). https://doi.org/10.1093/bioinformatics/btr381
R. C. Edgar, “UPARSE: highly accurate OTU sequences from microbial amplicon reads,” Nat. Methods 10, 996–998 (2013). https://doi.org/10.1038/NMETH.2604
E. Egidi, M. Delgado-Baquerizo, J. M. Plett, et al., “A few ascomycota taxa dominate soil fungal communities worldwide,” Nat. Commun. 10, 2369 (2019). https://doi.org/10.1038/s41467-019-10373-z
Soil Challenge Badge (UN Food and Agriculture Organization, Rome, 2014).
J. C. Frankland, N. Magan, and G. M. Gadd, Fungi and Environmental Change (Cambridge University Press, Cambridge, 1996). https://doi.org/10.1016/s0269-915x(98)80116-x
M. Frąc, S. E. Hannula, M. Bełka, and M. Jędryczka, “Fungal biodiversity and their role in soil health,” Front. Microbiol. 9, 707 (2018). https://doi.org/10.3389/fmicb.2018.00707
Gansu Province Soil Survey Office, Gansu Soil (China Agricultural Press, Beijing, 1993) [in Chinese].
R. Jeewon, Q. S. Yeung, D. N. Wannasinghe, et al., “Hidden mycota of pine needles: molecular signatures from PCR-DGGE and ribosomal DNA phylogenetic characterization of novel phylotypes,” Sci. Rep. 8, 18053 (2018). https://doi.org/10.1038/s41598-018-36573-z
A. Kumar and J. P. Verma, “Does plant-microbe interaction confer stress tolerance in plants: a review?” Microbiol. Res. 207, 41–52 (2018). https://doi.org/10.1016/j.micres.2017.11.004
U. Kõljalg, R. H. Nilsson, K. Abarenkov, et al., “Towards a unified paradigm for sequence-based identification of fungi,” Mol. Ecol. 22, 5271–5277 (2013). https://doi.org/10.1111/mec.12481
C. L. Lauber, M. S. Strickland, M. A. Bradford, and N. Fierer, “The influence of soil properties on the structure of bacterial and fungal communities across land-use types,” Soil Biol. Biochem. 40, 2407–2415 (2008). https://doi.org/10.1016/j.soilbio.2008.05.021
C. Loredana, L. P. Giuseppe, V. A. Livia, et al., “Spatial microbial community structure and biodiversity analysis in “extreme” hypersaline soils of a semiarid Mediterranean area,” Appl. Soil Ecol. 93 (10), 120–129 (2015). https://doi.org/10.1016/j.apsoil.2015.04.014
P. D. Li, R. Jeewon, B. Aruna, et al., “Metabarcoding reveals differences in fungal communities between unflooded versus tidal flat soil in coastal saline ecosystem,” Sci. Total Environ. 690 (1), 911–922 (2019). https://doi.org/10.1016/j.scitotenv.2019.06.473
W. Li, M. M. Wang, X. M. Bian, J. J. Guo, and L. Cai, “A high-level fungal diversity in the intertidal sediment of Chinese seas presents the spatial variation of community composition,” Front. Microbiol. 7, 2098 (2016). https://doi.org/10.3389/fmicb.2016.02098
C. Liu, J. Xu, N. Ding, et al., “The effect of long-term reclamation on enzyme activities and microbial community structure of saline soil at Shangyu, China,” Environ. Earth Sci. 69, 151–159 (2013). https://doi.org/10.1007/s12665-012-1943-1
F. T. Maestre, M. Delgado-Baquerizo, T. C. Jeffries, et al., “Increasing aridity reduces soil microbial diversity and abundance in global drylands,” Proc. Natl. Acad. Sci. U.S.A. 112, 15684–15689 (2015). https://doi.org/10.1073/pnas.1516684112
R. Munns and M. Tester, “Mechanisms of salinity tolerance,” Annu. Rev. Plant Biol. 59, 651–681 (2008). https://doi.org/10.1146/annurev.arplant.59.032607.092911
Z. Mustafa, M. A. Pervez, C. M. Ayyub, et al., “Morpho-physiological characterization of chilli genotypes under NaCl salinity,” Plant, Soil Environ. 33 (2), 133–141 (2014).
L. L. Nan and Q. E. Guo, “Soil properties under major halophytic vegetation communities in arid regions,” Wuhan Univ. J. Nat. Sci. 23 (5), 376–386 (2018). https://doi.org/10.1007/s11859-018-1337-7
L. L. Nan and Q. E. Guo, “Soil properties of Alhagi sparsifolia community in saline-sodic badlands in west China,” Acta Ecol. Sin. 38 (5), 339–344 (2018). https://doi.org/10.1016/j.chnaes.2018.01.010
L. L. Nan, Q. E. Guo, and S. Y. Cao, “Archaeal community diversity in different types of saline-alkali soil in arid regions of Northwest China,” J. Biosci. Bioeng. 130 (4), 382–389 (2020). https://doi.org/10.1016/j.jbiosc.2020.06.001
T. Nemse, B. Colomb, P. Hinsinger, and C. A. Watson, “Soil phosphorus management in organic cropping systems: From current practices to avenues for a more efficient use of p resources”, in Organic Farming, Prototype for Sustainable Agricultures (Springer-Verlag, New York, 2014), pp. 22–25. https://doi.org/10.1007/978-94-007-7927-3_2
R. H. Nilsson, L. Tedersoo, M. Ryberg, et al., “A comprehensive, automatically updated fungal ITS sequence dataset for reference-based chimera control in environmental sequencing efforts,” Microb. Environ. 30 (2), 145–150 (2015). https://doi.org/10.1264/jsme2.ME14121
I. H. Segel and M. J. Johnson, “Accumulation of intracellular inorganic sulfate by Penicillium chrysogenum,” J. Bacteriol. 81, 91–106 (1961). https://doi.org/10.1002/path.1700810111
L. Tedersoo, M. Bahram, S. Põlme, et al., “Global diversity and geography of soil fungi,” Science 346, 1256688 (2014). https://doi.org/10.1126/science.1256688
M. G. A. van der Heijden, R. D. Bardgett, and N. M. van Straalen, “The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems,” Ecol. Lett. 11, 296–310 (2008). https://doi.org/10.1111/j.1461-0248.2007.01139.x
D. Vu, M. Groenewald, M. de Vries, et al., “Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation,” Stud. Mycol. 92, 135–154 (2019). https://doi.org/10.1016/j.simyco.2018.05.001
S. Wang, L. Sun, N. Ling, et al., “Exploring soil factors determining composition and structure of the bacterial communities in saline-alkali soils of Songnen Plain,” Front. Microbiol. 10, 2902 (2020). https://doi.org/10.3389/fmicb.2019.02902
T. J. White, T. Bruns, S. Lee, et al., “Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics,” in PCR Protocols: A Guide to Methods and Applications (Academic, San Diego, 1990), pp. 315–322.
J. J. Worrall, S. E. Anagnost, and R. D. Zabel, “Comparison of wood decay among diverse lignicolous fungi,” Mycologia 89 (2), 199–219 (1997).
F. G. Zhang, Y. Q. Huo, X. X. Xu, et al., “Trichoderma improves the growth of Leymus chinensis,” Biol. Fertil. Soils 54, 685–696 (2018). https://doi.org/10.1007/s00374-018-1292-7
M.-M. Zhang, N. Wang, Y.-B. Hu, and G.-Y. Sun, “Changes in soil physicochemical properties and soil bacterial community in mulberry (Morus alba L.)/alfalfa (Medicago sativa L.) intercropping system,” MicrobiologyOpen 7 (2), e00555 (2018). https://doi.org/10.1002/mbo3.555
Funding
The study was supported by Project of Department of Science and Technology of Gansu Province (Nos. 20YF3FA011), National Natural Science Foundation of China (nos. 41363004 and 31460630), and the Key Research and Development Program of Gansu Academy of Agricultural Sciences (no. 2019GAAS24).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
All the authors certify that all co-authors have read the manuscript and agreed with its submission.
Rights and permissions
About this article
Cite this article
Quanen Guo, Nan, L., Cao, S. et al. Fungal Community Diversity in Solonchaks of Gansu Province in China. Eurasian Soil Sc. 55, 511–519 (2022). https://doi.org/10.1134/S106422932204010X
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S106422932204010X