Abstract
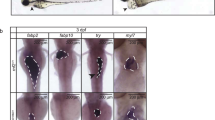
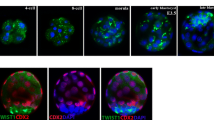
To create an organism, it is vital to assemble enough cells of the various differentiated types with the correct spatial arrangement within the embryo. Circadian clocks development is closely correlated with all cellular differentiation. However, the expression of its emergence during mammalian development are not fully understood. To determine whether embryonic development is influenced by circadian rhythm, it is necessary to observe the ontogeny of the circadian clock gene. We first measured the expression of key circadian genes in whole embryos and maternal major tissues of 25 female mice using RT-PCR and immunohistochemical analysis. Our results indicated that mouse embryos begin to express key circadian genes and have the capacity to express active circadian regulatory cycles during development. But circadian molecular rhythms can’t be built in embryo. At E15, the expression of Bmal1, Clock and Per1 mRNA in whole embryo were increased, especially Per1. In the meanwhile, immunohistochemical analysis shows a small number of PER1 positive cells were observed in the bottom of right atrium. From E16 to E17, CLOCK and PER1 positive cells were observed in the airway smooth muscle, the wall of left atrium and skeletal muscle of body wall. It is interesting that CLOCK and PER1 positive cells could not be detected in the liver. By using RT-PCR, we continue to observe the expression of myogenic regulatory factor in embryos and also analyse the relationship of embryo development and maternal rhythms. From E12, the expression of myogenin increased quickly. The expression of Tcap at E15 significantly increased. myogenin may play a direct role in contributing Tcap expression. The expression of MAZ is always the highest than myogenin and Tcap in embryos. MAZ may concern with the development of skeletal muscle. The clock gene is a positive regulator of myogenesis and the development of organ. In contrast to embryonic tissues, circadian variation was present for Bmal1, Clock and Per1 at maternal tissues. Our results indicate that circadian clock genes seem to function differently in different tissues of embryo and maternal mice. Synchrony does not occur during embryo development despite exposure to maternal rhythms. But development of embryo may be affected by maternal tissues of mice.





Similar content being viewed by others
References
Álvaro-Blanco J, Urso K, Chiodo Y, Martín-Cortázar C, Kourani O, Gómez-Del Arco P, Rodríguez-Martínez M, Calonge E, Alcamí J, Redondo JM, Iglesias T, Campanero MR (2017) MAZ induces MYB expression during the exit from quiescence via the E2F site in the MYB promoter. Nucleic Acids Res 45:9960–9975. https://doi.org/10.1093/nar/gkx641
Amano T, Matsushita A, Hatanaka Y, Watanabe T, Oishi K, Ishida N, Anzai M, Mitani T, Kato H, Kishigami S, Saeki K, Hosoi Y, Iritani A, Matsumoto K (2009) Expression and functional analyses of circadian genes in mouse oocytes and preimplantation embryos: Cry1 is involved in the meiotic process independently of circadian clock regulation. Biol Reprod 80:473–483. https://doi.org/10.1095/biolreprod.108.069542
Amaral IPG, Johnston IA (2012) Circadian expression of clock and putative clock-controlled genes in skeletal muscle of the zebrafish. Am J Physiol Regul Integr Comp Physiol 302:R193–R206. https://doi.org/10.1152/ajpregu.00367.2011
Andrews JL, Zhang X, Mccarthy JJ, McDearmon EL, Hornberger TA, Russell B, Campbell KS, Arbogast S, Reid MB, Walker JR, Hogenesch JB, Takahashi JS, Esser KA (2010) CLOCK and BMAL1 regulate MyoD and are necessary for maintenance of skeletal muscle phenotype and function. Proc Natl Acad Sci USA 107:19090–19095. https://doi.org/10.1073/pnas.1014523107
Aoyama S, Shibata S (2017) The role of circadian rhythms in muscular and osseous physiology and their regulation by nutrition and exercise. Front Neurosci 11:63. https://doi.org/10.3389/fnins.2017.00063
Bass J (2012) Circadian topology of metabolism. Nature 491:348–356. https://doi.org/10.1038/nature11704
Bos JM, Poley RN, Ny M, Tester DJ, Xu X, Vatta M, Towbin JA, Gersh BJ, Ommen SR, Ackerman MJ (2006) Genotype-phenotype relationships involving hypertrophic cardiomyopathy-associated mutations in titin, muscle LIM protein, and telethonin. Mol Genet Metab 88:78–85. https://doi.org/10.1016/j.ymgme.2005.10.008
Candasamy AJ, Haworth RS, Cuello F, Ibrahim M, Aravamudhan S, Krüger M, Holt MR, Terracciano CMN, Mayr M, Gautel M, Avkiran M (2014) Phosphoregulation of the titin-cap protein telethonin in cardiac myocytes. J Biol Chem 289:1282–1293. https://doi.org/10.1074/jbc.M113.479030
Conerly ML, Yao Z, Zhong JW, Groudine M, Tapscott SJ (2016) Distinct activities of Myf5 and MyoD indicate separate roles in skeletal muscle lineage specification and differentiation. Dev Cell 36:375–385. https://doi.org/10.1016/j.devcel.2016.01.021
Harfmann BD, Schroder EA, Esser KA (2015) Circadian rhythms, the molecular clock, and skeletal muscle. J Biol Rhythms 30:84–94. https://doi.org/10.1177/0748730414561638
Kondratov RV, Kondratova AA, Gorbacheva VY, Vykhovanets OV, Antoch MP (2006) Early aging and age-related pathologies in mice deficient in BMAL1, the core component of the circadian clock. Genes Dev 20:1868–1873. https://doi.org/10.1101/gad.1432206
Landgraf D, Achten C, Dallmann F, Oster H (2015) Embryonic development and maternal regulation of murine circadian clock function. Chronobiol Int 32:416–427. https://doi.org/10.3109/07420528.2014.986576
Mayeuf-Louchart A, Staels B, Duez H (2015) Skeletal muscle functions around the clock. Diabetes Obes Metab 17(Suppl 1):39–46. https://doi.org/10.1111/dom.12517
Podobed PS, Alibhai FJ, Chow CW, Martino TA (2014) Circadian regulation of myocardial sarcomeric Titin-cap (Tcap, telethonin):identification of cardiac clock-controlled genes using open access bioinformatics data. PLoS ONE 9:e104907. https://doi.org/10.1371/journal.pone.0104907
Qiao M, Huang J, Wu H, Wu J, Peng X, Mei S (2014) Molecular characterization, transcriptional regulation and association analysis with carcass traits of porcine TCAP gene. Gene 538:273–279. https://doi.org/10.1016/j.gene.2014.01.043
Robinson I, Reddy AB (2014) Molecular mechanisms of the circadian clockwork in mammals. FEBS 588:2477–2483. https://doi.org/10.1016/j.febslet.2014.06.005
Sato T, Rocancourt D, Marques L, Thorsteinsdóttir S, Buckingham M (2010) A Pax3/Dmrt2/Myf5 regulatory cascade functions at the onset of myogenesis. PLoS Genet 6:e1000897. https://doi.org/10.1371/journal.pgen.1000897
Song J, Murakami H, Tsutsui H, Ugai H, Geltinger C, Murata T, Matsumura M, Itakura K, Kanazawa I, Sun K, Yokoyama KK (1999) Structural organization and expression of the mouse gene for Pur-1, a highly conserved homolog of the human MAZ gene. Eur J Biochem 259:676–683. https://doi.org/10.1046/j.1432-1327.1999.00081.x
Sumová A, Bendová Z, Sládek M, El-Hennamy R, Matejů K, Polidarová L, Sosniyenko S, Illnerová H (2008) Circadian molecular clocks tick along ontogenesis. Physiol Res 57:S139–S148. https://doi.org/10.33549/physiolres.931458
Takahashi JS (2017) Transcriptional architecture of the mammalian circadian clock. Nat Rev Genet 18:164–179. https://doi.org/10.1038/nrg.2016.150
Umemura Y, Koike N, Ohashi M, Tsuchiya Y, Meng QJ, Minami Y, Hara M, Hisatomi M, Yagita K (2017) Involvement of posttranscriptional regulation of Clock in the emergence of circadian clock oscillation during mouse development. Proc Natl Acad Sci USA 114:E7479–E7488. https://doi.org/10.1073/pnas.1703170114
Xiao T, Li X, Felsenfeld G (2021) The Myc-associated zinc finger protein (MAZ) works together with CTCF to control cohesin positioning and genome organization. Proc Natl Acad Sci U S A 118:e2023127118. https://doi.org/10.1073/pnas.2023127118
Zhang S, Londhe P, Zhang M, Davie JK (2011) Transcriptional analysis of the titin cap gene. Mol Genet Genomics 285:261–272. https://doi.org/10.1007/s00438-011-0603-6
Funding
The work was supported by Natural Science Foundation of Shanxi Province, China (Grant Nos. 201901D111185, 2014021028-1), Science and Technology Innovation Fund of Shanxi Medical University (Grant No. 01201401).
Author information
Authors and Affiliations
Contributions
All authors participated in the design, interpretation of the studies and analysis of the data and review of the manuscript. YY supplied critical reagents. XC: conception and design of the study, data analysis and interpretation, obtaining of funding, drafting the article. All authors approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Cao, X., Yan, Y., Luo, X. et al. Analyses of the circadian clock genes expression in whole embryos and maternal major tissues of mice. J Mol Histol 53, 473–482 (2022). https://doi.org/10.1007/s10735-022-10065-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10735-022-10065-x