Abstract
Oysters are major transmission vectors of noroviruses (NoVs) in the environment. Outbreaks of NoVs are often associated with the consumption of NoV-contaminated oysters. Laboratory confirmation of suspected oyster samples is a critical step in the surveillance and control of NoVs. Because of non-specific amplification, false-positive results are frequently obtained by semi-nested RT-PCR with the presently widely used primer set (G2SKF/G2SKR). Here, a novel universal PCR primer set N (NG2OF/NG2OR) specific for genogroup II (GII) NoVs was designed based on all GII NoV sequences available in public databases. Specific products were obtained with the primer set N when the NoV-positive oysters, spiked with each of five representative genotypes of GII NoVs (GII.17, GII.13, GII.4, GII.3, and GII.12), were subjected to analyzing. No products were detected with the primer set N for the NoV-negative oysters, while the primer set C gave various non-specific bands. Twenty-three out of 156 fresh oyster samples were NoV-positive with both the primer set N and the classic primer set, while eight were NoV-positive solely with the primer set N. Compared with the classic primer set, the newly designed primer set N had a higher detection rate and improved specificity for GII NoVs in oyster samples. These results show that the novel PCR primer pair is specific and applicable for the detection of GII NoVs in oysters.
Similar content being viewed by others
Introduction
Noroviruses (NoVs) are one of the main causes of human non-bacterial acute gastroenteritis (AGE) (de Graaf et al., 2016). In children less than five years of age, NoVs account for about one-fifth of all AGE cases globally (Ahmed et al., 2014). NoVs belong to the family Caliciviridae (de Graaf et al., 2016; Robilotti et al., 2015). The NoV genome includes a single-stranded, positive-sense RNA, which contains three open reading frames (ORFs): ORF1 encodes six non-structural proteins, including RNA-dependent RNA polymerase (RdRP); ORF2 encodes the major capsid protein (VP1); and ORF3 encodes the minor structural protein (VP2) (de Graaf et al., 2016; Vinjé, 2015). Currently, NoVs are divided into 10 genogroups (GI-GX). Five genogroups (GI, GII, GIV, GVIII, and GIX) infect humans (Chhabra et al., 2019). Among these, GII is the most prevalent human NoV worldwide (Mattison et al., 2018), including China (Qin et al., 2017; Yu et al., 2014).
Transmission of NoVs can be by diverse routes, including fecal–oral (through contaminated food or water), and direct or indirect contact with contaminated surfaces (de Graaf et al., 2016). Oysters are recognized as common transmission vectors of NoVs, and outbreaks are often associated with the consumption of NoV-contaminated oysters (Bitler et al., 2013; Mathijs et al., 2012). As filter feeders, oysters can accumulate NoVs from the surrounding water at concentrations as high as 103–107 genome copies per gram in digestive tissue (Sarmento et al., 2020; Tan et al., 2018). More than 80% of human NoV genotypes have been detected in oyster samples or oyster-related NoV outbreaks (Yu et al., 2015). Eating raw or insufficiently heated oysters presents a potentially high risk for NoV infection (Alfano-Sobsey et al., 2012; Cheng et al., 2005). Oyster-related AGE has been documented worldwide (Iritani et al., 2014; McIntyre et al., 2012; Meghnath et al., 2019; Nenonen et al., 2009; Pu et al., 2018). Therefore, effective and accurate monitoring of NoV contamination in oyster samples is of great importance.
Laboratory confirmation of suspected oyster samples is a critical step in the surveillance and control NoV contamination. Among the molecular detection methods available for NoVs, real-time RT-PCR is relatively mature and highly sensitive for human samples (Jothikumar et al., 2005; Zhou et al., 2017); however, the resultant amplification products are too short (Anonymous, 2013; Kageyama et al., 2003) to be used for subsequent sequencing and genotyping, making investigation of NoV genotypes in oysters difficult. In addition, false-positives are common. With the development of next-generation sequencing, amplicon and transcriptomic sequencing have been increasingly used in oyster NoV research (Desdouits et al., 2020; Imamura et al., 2016a, 2016b; Strubbia et al., 2020). Metagenomic sequencing is time-consuming and costly, making it unsuitable for routine monitoring of NoVs in oysters. Nested RT-PCR, which amplifies target fragments over two rounds of PCR, each using different primers pairs, can improve the detection specificity for samples with low template concentrations (Medici et al., 2005). Two pairs of genogroup-specific primers were initially designed for the detection of GII noroviruses in human stool samples by using RT-PCR (G2SKF/G2SKR) (Kojima et al., 2002) and real-time RT-PCR (COG2F/COG2R) (Kageyama et al., 2003), respectively. Soon after, three of these four primers were combined in semi-nested RT-PCR (COG2F/G2SKR for RT-PCR and G2SKF/G2SKR for semi-nested PCR) to increase the detection sensitivity for oyster samples (Nishida et al., 2003), which were named as set C (C, the abbreviation for classic) in this study. Presently, these primer pairs have been widely used for routine detection of GII NoVs (Ji et al., 2020; Parrón et al., 2019; Zhang et al., 2020), including by semi-nested PCR methods (Bhavanam et al., 2020; Iritani et al., 2010; Ueki et al., 2005).
However, in our study, detection of NoV contamination in oysters by using the primer set C resulted in many non-specific PCR products (Figure S1), which were amplified from the DNAs of oysters and their intestinal bacteria (Table S1). Apparently, the specificity of the primer set C is interfered with by digestive tissue and intestinal bacterial community of oysters.
In this study, we sought to improve specificity for the detection of GII NoVs in oyster samples by designing a novel semi-nested RT-PCR primer set. Comparing with the classic primer set C, our newly designed primer set N (N, the abbreviation for new) can dramatically improve the detection specificity and detection rate for GII NoVs in commercial oyster samples, and represents an alternative to better detect the contamination level of the GII NoVs in oysters.
Materials and Methods
Primer Design and In Silico Analysis
All NoV sequences available online were downloaded from NCBI nucleotide database (https://www.ncbi.nlm.nih.gov/nucleotide/) on December 14, 2020 and then genotyped by using the Norovirus Typing Tool v.2.0 (https://www.rivm.nl/mpf/typingtool/norovirus). Sequences containing ORF1 only or having fewer than 1000 nucleotides (nt) were removed from analysis (Guo et al., 2018; Jia et al., 2020). The most abundant genotype was GII.4, with a sequence abundance of 52.7%. Conserved regions of GII.4 NoV sequences were identified using ClusterW (https://www.geneious.com). Primer candidates were designed based on these conserved regions (Fig. 1). Primer specificity was verified in silico using Primer-BLAST against the GenBank nucleotide database. Mismatches in conserved regions were evaluated using the program of Map primers (Geneious Primer version 2019.0.3) (Fig. 1).
Schematic of design and in silico evaluation of the semi-nested PCR primers for GII noroviruses. Removal of short sequences yielded 7385 sequences ranging from 1000 to 7618 nucleotides, which belong to GII.1 (n = 67), GII.2 (n = 1068), GII.3 (n = 474), GII.4 (n = 3893), GII.5 (n = 46), GII.6 (n = 238), GII.7 (n = 42), GII.8 (n = 49), GII.9 (n = 25), GII.10 (n = 8), GII.11 (n = 8), GII.12 (n = 66), GII.13 (n = 105), GII.14 (n = 53), GII.16 (n = 20), GII.17 (n = 1086), GII.18 (n = 3), GII.19 (n = 2), GII.20 (n = 5), GII.21 (n = 77), GII.22 (n = 2), GII.23 (n = 6), GII.24 (n = 5), GII.25 (n = 7), GII.26 (n = 19), GII.27 (n = 8), GII.NA1 (n = 1), and GII.NA2 (n = 2) based on capsid genotyping. a The GII.4 sequences were used for multiple sequence alignment analysis to find conserved regions. b The conserved regions of GII.4 were mapped to the reference sequences of each genotype to select the best primers. c Mismatches between the primer set N and the reference sequences of each genotype were identified and are in red bold text
Oyster Samples
Commercial oyster samples (Crassostrea gigas, body length of 9–12 cm, n = 156) were purchased fresh from the Luchaogang aquatic products market in Shanghai, China, from October 2020 to January 2021. The oyster samples were transported on ice, and dissected on the day of sampling.
RNA Extraction
The entire digestive gland tissue (approximately 400 mg) of each oyster was dissected and homogenized by using a FastPrep-24 (MP Biomedicals, USA) at a speed of 6 m/s for 30 s. Approximately, 50 mg of the homogenized digestive tissue was subjected to total RNA extraction using the Animal Tissue Total RNA Extraction Kit (Generay Biotech, Shanghai) according to the manufacturer’s instructions. Extracted RNA was stored at − 80 °C until use.
Sensitivity and Specificity Evaluation of Primer Sets
Oyster digestive tissue was confirmed as NoV-negative by semi-nested RT-PCR with primer set C. These oysters were then used for primer sensitivity and specificity assessment. Firstly, digestive tissues were spiked, respectively, with NoV using NoV-positive stool samples containing five GII genotypes (GII.17, GII.13, GII.4, GII.3, and GII.12) frequently detected in oysters. Digestive tissue was contaminated at a rate of 105 genome copies of NoVs per 50 mg of tissue. Secondly, stool samples containing rotavirus, sapovirus, astrovirus, or GI.4 NoV, were added to NoV-negative oyster digestive tissues as controls.
Semi-nested RT-PCR
RT-PCR (first-round) was performed in a reaction volume of 25 μL using a One Step RT-PCR Kit (Vazyme Biotech, Nanjing) according to the manufacturer’s protocol (Table 1). Reaction conditions were as follows: 50 °C for 30 min; 94 °C for 2 min; 35 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s. The RT-PCR products were diluted ten times and used as templates for semi-nested PCR (Table 1). The second round was also carried out in a volume of 25 μL using a 2 × Taq Master Mix Kit (Novoprotein, Shanghai) according to the manufacturer’s manual. Reaction conditions were as follows: 94 °C for 5 min; 35 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s; hold at 72 °C for 15 min. The amplified products were analyzed by electrophoresis on a 2% agarose gel. The expected sizes of PCR products of the primer set C and primer set N were 344 and 337 bp, respectively.
Results
Primer Design and In Silico Analysis
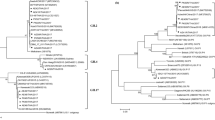
The new primer set N (RT-PCR primers COG2F/NG2OR, semi-nested PCR primers NG2OF/NG2OR) was designed based on conserved regions of the GII.4 NoV genome. The primer sets C and N are shown in Table 1. When the primers were aligned with the 28 GII reference sequences, 13 and 19 mismatches were observed for the primer set N and primer set C, respectively (Fig. 2). These results indicate that both primer sets are specific for GII NoVs in silico.
Sensitivity Evaluation
As shown in Fig. 3B, both primer sets gave the expected products in semi-nested RT-PCR of oyster tissue artificially contaminated with NoV genotypes GII.17, GII.13, GII.4, GII.3, and GII.12. Sequencing results confirmed that all the specific products were consistent with their corresponding genotypes, indicating similarity in the sensitivity of these two primer sets. However, the bands of GII.4 and GII.3 produced by the primer set N are weaker than that by the primer set C, suggesting that the amplification efficiency of the primer set N in detecting GII.4 and GII.3 may be lower.
Experimental evaluation of the performance of semi-nested RT-PCR primer sets. A Detection of norovirus-negative oyster samples. 1–10: Norovirus-negative oyster samples. B Sensitivity assessment based on five representative GII noroviruses. C Specificity assessment based on other viruses. RoV rotavirus, SaV sapovirus, AsV astrovirus, GI.4 norovirus, M 100 bp DNA ladder, N negative control
Specificity Evaluation
Semi-nested RT-PCR using the primer set N gave no obvious bands with any of the control samples (rotavirus, sapovirus, astrovirus, or GI.4 NoV) and NoV-negative oyster samples, and the background of the gel was relatively clean (Fig. 3A, C). In contrast, the primer set C gave various non-specific bands (100–1000 bp), including one near the expected size (344 bp). The non-specific amplicons (false-positive bands) were recovered, cloned, and sequenced; sequencing results based on BLASTN confirmed that these amplicons originated from the genome of oysters or their intestinal bacteria (Figure S1, Table S1). Taken together, these results indicate that the primer set N is highly specific, while the primer set C yields non-specific false-positive amplifications.
Detection in Oyster Samples
A total of 156 oyster samples were detected by semi-nested RT-PCR using the primer set C and set N. The detection rate of NoV-positive sample was higher with the primer set N (19.87%, 31/156; Table 2) than with the primer set C (14.74%, 23/156; Table 2). Twenty-three samples were NoV-positive with two primer sets, while eight were NoV-positive solely with the primer set N. Further sequencing confirmed that all eight samples were NoV-positive with three genotypes including GII.2 (n = 2), GII.3 (n = 4), and GII.17 (n = 2) (Table S2). These results suggest the higher detection efficiency of the primer set N compared with that of the primer set C.
Discussion
NoVs are genetically diverse RNA viruses, and thousands of NoV sequences have emerged in recent years (Chhabra et al., 2019; de Graaf et al., 2016; Jia et al., 2020). In 2013, 8951 NoV sequences were available from the NCBI database. At that time, NoVs were divided into at least six genogroups (GI–GVI) using phylogenetic clustering of the complete VP1 amino acid sequences (Kroneman et al., 2013). Soon afterward, the number of NoV sequences has increased dramatically due to the rapid evolution of the viral genome by genetic mutation and recombination (de Graaf et al., 2016). By 2019, the number of NoV sequences reached 45,542, of which, over 68.8% (n = 31,343) belonged to GII (available data from NCBI). The classification of NoVs was then updated into 10 genogroups (GI-GX), which were further divided into at least 49 capsid genotypes and 60 P-types, based on the diversity of VP1 and RdRP, respectively (Chhabra et al., 2019). Therefore, it is very urgent to update primers for the specific detection of NoVs in oysters.
In our previous study, we re-designed a new universal primer set of nested RT-PCR for the specific detection of GI NoVs in oyster samples (Guo et al., 2018). Our results indicate that the new nested primers (NGIOF/NGIOR) are much higher specific than the classic primers (GISKF/GISKR) (Guo et al., 2018; Kojima et al., 2002).
In this study, non-specific amplification was also observed when oyster samples were analyzed for GII NoV contamination with the presently widely used primer set C (Figure S1, Table S1). Accordingly, the new primer set N for the specific detection of GII NoVs was designed based on the conserved regions of all GII.4 sequences available in public databases (Fig. 1). Using in silico analysis, the new primer set N was found to specifically bind with all 28 GII genotypes. Compared with the classic primer set C, the primer set N possesses fewer mismatches with the reference sequences (Fig. 2). Moreover, the primer set N had minimal degeneracy and no degenerate sites near the 3′ terminus of either primer, suggesting low amplification bias. Furthermore, unlike the primer set C, the primer set N gave no obvious non-specific amplicons with oyster samples (Fig. 3A). Several differences are notable between the two primer sets (Table 1). The primer set N has five degenerate bases versus seven in the primer set C. The reverse primer NG2OR (20 nt) is shorter than G2SKR (23 nt), uses the degenerate base “V” instead of “H” in G2SKR at the same position, and avoids the third degenerate base “R” at the 5′ end of G2SKR (Table 1). In addition, the forward primer NG2OF avoids the second degenerate base “N” at the 5′ end of G2SKF. These differences may be responsible for the observed higher binding specificity of the primer set N for GII NoVs, especially under a low viral load. When GII NoV templates were present in abundance, both primer sets were observed to preferentially bind with NoV cDNA and specifically amplify the target sequences, with no obvious non-specific amplifications (Fig. 3B).
For the artificially contaminated oyster samples, the amplification efficiency of the primer set N in detecting GII.4 and GII.3 NoVs may be lower than that of the primer set C (Fig. 3B). However, comprehensive assessment showed that the primer set N had a higher detection rate for genotypes GII.2, GII.3, and GII.17 than the primer set C (Table 2). Coincidently, novel variants of two of these genotypes have recently emerged (GII.P17-GII.17 during the 2014–2015 season, and GII.P16-GII.2 during the 2016–2017 season), causing notable increases in AGE outbreaks worldwide (Ao et al., 2017; Gao et al., 2019; Lu et al., 2016; Rasmussen et al., 2016). This suggests that, as novel variants of NoVs continue to emerge, primers had to be updated regularly to ensure specific and sensitive detection.
Interestingly, in this study, the detection rate of NoVs is 19.87% (31/156; Table 2) for the 156 oysters collected from October 2020 to January 2021, which is slightly lower than that of the samples checked at other times. This probably results from the stringent polices regarding to the prevention and control of the epidemic of COVID-19 in China. Limitation of direct or indirect contact resulted in the significant decline of COVID-19 and simultaneously hampered the spreading of NoVs as well (Ahn et al., 2021; Eigner et al., 2021; Farah et al., 2021; Lennon et al., 2020).
In summary, the established semi-nested RT-PCR with updated primer sets bearing higher specificity and sensitivity is highly desirable and satisfies the urgent needs of surveying and genotyping of GII NoV contamination in oysters.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Ahmed, S. M., Hall, A. J., Robinson, A. E., Verhoef, L., Premkumar, P., Parashar, U. D., Koopmans, M., & Lopman, B. A. (2014). Global prevalence of norovirus in cases of gastroenteritis: A systematic review and meta-analysis. The Lancet Infectious Diseases, 14(8), 725–730. https://doi.org/10.1016/S1473-3099(14)70767-4
Ahn, S. Y., Park, J. Y., Lim, I. S., Chae, S. A., Yun, S. W., Lee, N. M., Kim, S. Y., Choi, B. S., & Yi, D. Y. (2021). Changes in the occurrence of gastrointestinal infections after COVID-19 in Korea. Journal of Korean Medical Science, 36(24), e180. https://doi.org/10.3346/jkms.2021.36.e180
Alfano-Sobsey, E., Sweat, D., Hall, A., Breedlove, F., Rodriguez, R., Greene, S., Pierce, A., Sobsey, M., Davies, M., & Ledford, S. L. (2012). Norovirus outbreak associated with undercooked oysters and secondary household transmission. Epidemiology and Infection, 140(2), 276–282. https://doi.org/10.1017/S0950268811000665
Anonymous, P. (2013). Microbiology of food and animal feed: Horizontal method for detection of hepatitis A virus and norovirus in food using real-time RT-PCR: Part 2: Method for qualitative detection (ISO/TS 15216-2:2013).
Ao, Y., Wang, J., Ling, H., He, Y., Dong, X., Wang, X., Peng, J., Zhang, H., Jin, M., & Duan, Z. (2017). Norovirus GII.P16/GII.2-associated gastroenteritis, China, 2016. Emerging Infectious Diseases, 23(7), 1172–1175. https://doi.org/10.3201/eid2307.170034
Bhavanam, S., Freedman, S. B., Lee, B. E., Zhuo, R., Qiu, Y., Chui, L., Xie, J., Ali, S., Vanderkooi, O. G., Pang, X. L., The Alberta Provincial Pediatric Enteric Infection Team A. (2020). Differences in illness severity among circulating norovirus genotypes in a large pediatric cohort with acute gastroenteritis. Microorganisms, 8(12), 1873. https://doi.org/10.3390/microorganisms8121873
Bitler, E. J., Matthews, J. E., Dickey, B. W., Eisenberg, J. N. S., & Leon, J. S. (2013). Norovirus outbreaks: A systematic review of commonly implicated transmission routes and vehicles. Epidemiology and Infection, 141(8), 1563–1571. https://doi.org/10.1017/S095026881300006X
Cheng, P. K. C., Wong, D. K. K., Chung, T. W. H., & Lim, W. W. L. (2005). Norovirus contamination found in oysters worldwide. Journal of Medical Virology, 76(4), 593–597.
Chhabra, P., de Graaf, M., Parra, G. I., Chan, M. C., Green, K., Martella, V., Wang, Q., White, P. A., Katayama, K., Vennema, H., Koopmans, M. P. G., & Vinje, J. (2019). Updated classification of norovirus genogroups and genotypes. Journal of General Virology, 100(10), 1393–1406. https://doi.org/10.1099/jgv.0.001318
de Graaf, M., van Beek, J., & Koopmans, M. P. G. (2016). Human norovirus transmission and evolution in a changing world. Nature Reviews Microbiology, 14(7), 421–433. https://doi.org/10.1038/nrmicro.2016.48
Desdouits, M., Wacrenier, C., Ollivier, J., Schaeffer, J., & Guyader, F. S. L. (2020). A targeted metagenomics approach to study the diversity of norovirus GII in shellfish implicated in outbreaks. Viruses, 12(9), 978. https://doi.org/10.3390/v12090978
Eigner, U., Verstraeten, T., & Weil, J. (2021). Decrease in norovirus infections in Germany following COVID-19 containment measures. The Journal of Infection, 82(6), 276–316. https://doi.org/10.1016/j.jinf.2021.02.012
Farah, Y., Syed Hasan, A., & Irfan, U. (2021). Norovirus outbreak amid COVID-19 in the United Kingdom; priorities for achieving control. Journal of Medical Virology. https://doi.org/10.1002/jmv.27426
Gao, Z., Liu, B., Yan, H., Li, W., Jia, L., Tian, Y., Chen, Y., Wang, Q., & Pang, X. (2019). Norovirus outbreaks in Beijing, China, from 2014 to 2017. The Journal of Infection, 79(2), 159–166. https://doi.org/10.1016/j.jinf.2019.05.019
Guo, P., Yu, Y., Pan, Y., Yan, S., & Wang, Y. (2018). Design and evaluation of nested PCR primers for specific detection of genogroup I noroviruses in oysters. Molecular and Cellular Probes, 40, 40–43. https://doi.org/10.1016/j.mcp.2018.06.002
Imamura, S., Haruna, M., Goshima, T., Kanezashi, H., Okada, T., & Akimoto, K. (2016a). Application of next-generation sequencing to evaluate the profile of noroviruses in pre- and post-depurated oysters. Foodborne Pathogens and Disease, 13(10), 559–565.
Imamura, S., Haruna, M., Goshima, T., Kanezashi, H., Okada, T., & Akimoto, K. (2016b). Application of next-generation sequencing to investigation of norovirus diversity in shellfish collected from two coastal sites in Japan from 2013 to 2014. The Japanese Journal of Veterinary Research, 64(2), 113–122.
Iritani, N., Kaida, A., Abe, N., Kubo, H., Sekiguchi, J.-I., Yamamoto, S. P., Goto, K., Tanaka, T., & Noda, M. (2014). Detection and genetic characterization of human enteric viruses in oyster-associated gastroenteritis outbreaks between 2001 and 2012 in Osaka City, Japan. Journal of Medical Virology, 86(12), 2019–2025. https://doi.org/10.1002/jmv.23883
Iritani, N., Kaida, A., Kubo, H., Abe, N., Goto, K., Ogura, H., & Seto, Y. (2010). Molecular epidemiology of noroviruses detected in seasonal outbreaks of acute nonbacterial gastroenteritis in Osaka City, Japan, from 1996–1997 to 2008–2009. Journal of Medical Virology, 82(12), 2097–2105. https://doi.org/10.1002/jmv.21915
Ji, L., Hu, G., Xu, D., Wu, X., Fu, Y., & Chen, L. (2020). Molecular epidemiology and changes in genotype diversity of norovirus infections in acute gastroenteritis patients in Huzhou, China, 2018. Journal of Medical Virology. https://doi.org/10.1002/jmv.26247
Jia, T., Yu, Y., & Wang, Y. (2020). A recombinase polymerase amplification-based lateral flow strip assay for rapid detection of genogroup II noroviruses in the field. Archives of Virology, 165(12), 2767–2776. https://doi.org/10.1007/s00705-020-04798-x
Jothikumar, N., Lowther, J. A., Henshilwood, K., Lees, D. N., Hill, V. R., & Vinjé, J. (2005). Rapid and sensitive detection of noroviruses by using TaqMan-based one-step reverse transcription-PCR assays and application to naturally contaminated shellfish samples. Applied and Environmental Microbiology, 71(4), 1870–1875.
Kageyama, T., Kojima, S., Shinohara, M., Uchida, K., Fukushi, S., Hoshino, F. B., Takeda, N., & Katayama, K. (2003). Broadly reactive and highly sensitive assay for norwalk-like viruses based on real-time quantitative reverse transcription-PCR. Journal of Clinical Microbiology, 41(4), 1548–1557. https://doi.org/10.1128/jcm.41.4.1548-1557.2003
Kojima, S., Kageyama, T., Fukushi, S., Hoshino, F. B., Shinohara, M., Uchida, K., Natori, K., Takeda, N., & Katayama, K. (2002). Genogroup-specific PCR primers for detection of Norwalk-like viruses. Journal of Virological Methods, 100(1–2), 107–114.
Kroneman, A., Vega, E., Vennema, H., Vinjé, J., White, P. A., Hansman, G., Green, K., Martella, V., Katayama, K., & Koopmans, M. (2013). Proposal for a unified norovirus nomenclature and genotyping. Archives of Virology, 158(10), 2059–2068. https://doi.org/10.1007/s00705-013-1708-5
Lennon, R. P., Griffin, C., Miller, E. L., Dong, H., Rabago, D., & Zgierska, A. E. (2020). Norovirus infections drop 49% in the United States with strict COVID-19 public health interventions. Acta Medica Academica, 49(3), 278–280. https://doi.org/10.5644/ama2006-124.317
Lu, J., Fang, L., Zheng, H., Lao, J., Yang, F., Sun, L., Xiao, J., Lin, J., Song, T., Ni, T., Raghwani, J., Ke, C., Faria, N. R., Bowden, T. A., Pybus, O. G., & Li, H. (2016). The evolution and transmission of epidemic GII.17 Noroviruses. The Journal of Infectious Diseases, 214(4), 556–564. https://doi.org/10.1093/infdis/jiw208
Mathijs, E., Stals, A., Baert, L., Botteldoorn, N., Denayer, S., Mauroy, A., Scipioni, A., Daube, G., Dierick, K., Herman, L., Van Coillie, E., Uyttendaele, M., & Thiry, E. (2012). A review of known and hypothetical transmission routes for noroviruses. Food and Environmental Virology, 4(4), 131–152. https://doi.org/10.1007/s12560-012-9091-z
Mattison, C. P., Cardemil, C. V., & Hall, A. J. (2018). Progress on norovirus vaccine research: Public health considerations and future directions. Expert Review of Vaccines, 17(9), 773–784. https://doi.org/10.1080/14760584.2018.1510327
McIntyre, L., Galanis, E., Mattison, K., Mykytczuk, O., Buenaventura, E., Wong, J., Prystajecky, N., Ritson, M., Stone, J., Moreau, D., & Youssef, A. (2012). Multiple clusters of norovirus among shellfish consumers linked to symptomatic oyster harvesters. Journal of Food Protection, 75(9), 1715–1720. https://doi.org/10.4315/0362-028X.JFP-12-113
Medici, M. C., Martinelli, M., Ruggeri, F. M., Abelli, L. A., Bosco, S., Arcangeletti, M. C., Pinardi, F., De Conto, F., Calderaro, A., Chezzi, C., & Dettori, G. (2005). Broadly reactive nested reverse transcription-PCR using an internal RNA standard control for detection of noroviruses in stool samples. Journal of Clinical Microbiology, 43(8), 3772–3778.
Meghnath, K., Hasselback, P., McCormick, R., Prystajecky, N., Taylor, M., McIntyre, L., Man, S., Whitfield, Y., Warshawsky, B., McKinley, M., Bitzikos, O., Hexemer, A., & Galanis, E. (2019). Outbreaks of norovirus and acute gastroenteritis associated with British Columbia oysters, 2016–2017. Food and Environmental Virology, 11(2), 138–148. https://doi.org/10.1007/s12560-019-09374-4
Nenonen, N. P., Hannoun, C., Olsson, M. B., & Bergström, T. (2009). Molecular analysis of an oyster-related norovirus outbreak. Journal of Clinical Virology, 45(2), 105–108. https://doi.org/10.1016/j.jcv.2009.04.011
Nishida, T., Kimura, H., Saitoh, M., Shinohara, M., Kato, M., Fukuda, S., Munemura, T., Mikami, T., Kawamoto, A., Akiyama, M., Kato, Y., Nishi, K., Kozawa, K., & Nishio, O. (2003). Detection, quantitation, and phylogenetic analysis of noroviruses in Japanese oysters. Applied and Environment Microbiology, 69(10), 5782–5786. https://doi.org/10.1128/aem.69.10.5782-5786.2003
Parrón, I., Álvarez, J., Jané, M., Cornejo Sánchez, T., Razquin, E., Guix, S., Camps, G., Pérez, C., & Domínguez, À. (2019). A foodborne norovirus outbreak in a nursing home and spread to staff and their household contacts. Epidemiology and Infection, 147, e225. https://doi.org/10.1017/S0950268819001146
Pu, J., Miura, T., Kazama, S., Konta, Y., Azraini, N. D., Ito, E., Ito, H., Omura, T., & Watanabe, T. (2018). Weekly variations in norovirus genogroup II genotypes in Japanese oysters. International Journal of Food Microbiology, 284, 48–55. https://doi.org/10.1016/j.ijfoodmicro.2018.06.027
Qin, S. W., Chan, T. C., Cai, J., Zhao, N., Miao, Z. P., Chen, Y. J., & Liu, S. L. (2017). Genotypic and epidemiological trends of acute gastroenteritis associated with noroviruses in China from 2006 to 2016. International Journal of Environmental Research and Public Health, 14(11), 1341. https://doi.org/10.3390/ijerph14111341
Rasmussen, L. D., Schultz, A. C., Uhrbrand, K., Jensen, T., & Fischer, T. K. (2016). Molecular evidence of oysters as vehicle of norovirus GII.P17-GII.17. Emerging Infectious Diseases, 22(11), 2024–2025. https://doi.org/10.3201/eid2211.161171
Robilotti, E., Deresinski, S., & Pinsky, B. A. (2015). Norovirus. Clinical Microbiology Reviews, 28(1), 134–164. https://doi.org/10.1128/CMR.00075-14
Sarmento, S. K., Guerra, C. R., Malta, F. C., Coutinho, R., Miagostovich, M. P., & Fumian, T. M. (2020). Human norovirus detection in bivalve shellfish in Brazil and evaluation of viral infectivity using PMA treatment. Marine Pollution Bulletin, 157, 111315. https://doi.org/10.1016/j.marpolbul.2020.111315
Strubbia, S., Schaeffer, J., Besnard, A., Wacrenier, C., Le Mennec, C., Garry, P., Desdouits, M., & Le Guyader, F. S. (2020). Metagenomic to evaluate norovirus genomic diversity in oysters: Impact on hexamer selection and targeted capture-based enrichment. International Journal of Food Microbiology, 323, 108588. https://doi.org/10.1016/j.ijfoodmicro.2020.108588
Tan, D. M., Lyu, S. L., Liu, W., Zeng, X. Y., Lan, L., Qu, C., Zhuge, S. Y., Zhong, Y. X., Xie, Y. H., & Li, X. G. (2018). Utility of droplet digital PCR assay for quantitative detection of norovirus in shellfish, from production to consumption in Guangxi, China. Biomedical and Environmental Sciences, 31(10), 713–720. https://doi.org/10.3967/bes2018.096
Ueki, Y., Sano, D., Watanabe, T., Akiyama, K., & Omura, T. (2005). Norovirus pathway in water environment estimated by genetic analysis of strains from patients of gastroenteritis, sewage, treated wastewater, river water and oysters. Water Research, 39(18), 4271–4280. https://doi.org/10.1016/j.watres.2005.06.035
Vinjé, J. (2015). Advances in laboratory methods for detection and typing of norovirus. Journal of Clinical Microbiology, 53(2), 373–381. https://doi.org/10.1128/JCM.01535-14
Yu, Y., Cai, H., Hu, L., Lei, R., Pan, Y., Yan, S., & Wang, Y. (2015). Molecular epidemiology of oyster-related human noroviruses and their global genetic diversity and temporal-geographical distribution from 1983 to 2014. Applied and Environmental Microbiology, 81(21), 7615–7624. https://doi.org/10.1128/AEM.01729-15
Yu, Y., Yan, S., Li, B., Pan, Y., & Wang, Y. (2014). Genetic diversity and distribution of human norovirus in China (1999–2011). BioMed Research International, 2014, 1–13. https://doi.org/10.1155/2014/196169
Zhang, X. F., Chen, J. R., Song, C. L., Xie, D. J., Tan, M., Wang, L., Koroma, M. M., Hou, Y. Z., Dong, Z. P., Yu, J. R., Duan, W. T., Zhao, D. D., Du, J. R., Zhu, L., & Dai, Y. C. (2020). Characterization of a hospital-based gastroenteritis outbreak caused by GII6 norovirus in Jinshan, China. Epidemiology and Infection, 148, e289. https://doi.org/10.1017/S0950268820002538
Zhou, Z., Tian, Z., Li, Q., Tian, P., Wu, Q., Wang, D., & Shi, X. (2017). Capture RT-qPCR: A new simple and sensitive method to detect human norovirus in oysters. Frontiers in Microbiology, 8, 554. https://doi.org/10.3389/fmicb.2017.00554
Acknowledgements
The stool samples, containing norovirus, rotavirus, sapovirus, and astrovirus, were kindly provided by the Chinese Center for Disease Control and Prevention (CDC).
Funding
This study was funded by the National Key R&D Program of China (2017YFC1600703) and the National Natural Science Foundation of China (31601570, 41876195).
Author information
Authors and Affiliations
Contributions
YW conceived and designed this study. YY and TJ downloaded the sequences and designed the primers. TJ and LD performed the sampling and experiments. LD and YY wrote the manuscript. YW modified the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dong, L., Jia, T., Yu, Y. et al. Updating a New Semi-nested PCR Primer Pair for the Specific Detection of GII Norovirus in Oysters. Food Environ Virol 14, 149–156 (2022). https://doi.org/10.1007/s12560-022-09511-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12560-022-09511-6