Abstract
It is the centenary of the discovery of oligodendrocytes and we are increasingly aware of their importance in the functioning of the brain in development, adult learning, normal ageing and in disease across the life course, even in those diseases classically thought of as neuronal. This has sparked more interest in oligodendroglia for potential therapeutics for many neurodegenerative/neurodevelopmental diseases due to their more tractable nature as a renewable cell in the central nervous system. However, oligodendroglia are not all the same. Even from the first description, differences in morphology were described between the cells. With advancing techniques to describe these differences in human tissue, the complexity of oligodendroglia is being discovered, indicating apparent functional differences which may be of critical importance in determining vulnerability and response to disease, and targeting of potential therapeutics. It is timely to review the progress we have made in discovering and understanding oligodendroglial heterogeneity in health and neuropathology.
Similar content being viewed by others
Introduction
To state the obvious, humans are not the same as mice, rats, zebrafish or other preclinical model animals. Although all our cells are different, the reason that we are human is due to our brain structure, and so it could be argued that understanding the physiological heterogeneity, pattern and biology of human brain cells may be even more important than with other cells. This difference and insufficient understanding of human brain cells may be reflected in the almost complete lack of effective treatments for neurodegenerative diseases. While initially research on the human central nervous system (CNS) has focussed for decades on neurons, more recent developments in the field show the important role of glia—that were first only seen as sticky substance that held the neurons in place [115]—in health and disease. Here, we will focus on oligodendroglia, the most prevalent cell lineage within the white matter, which include oligodendrocyte precursor cells (OPCs), immature pre-myelinating oligodendrocytes and mature myelinating oligodendrocytes. Mature oligodendrocytes form membrane extensions which wrap around nerve axons, forming the myelin sheath. The myelin sheath allows fast conduction velocity of electrical impulses along axons and provides metabolic support from the oligodendrocyte cell body to the underlying axon [34, 68]. Until relatively recently, we have tended to consider oligodendrocytes as one cell type, as they all form myelin, however, there is increasing evidence that oligodendroglia are morphologically [87] and functionally heterogeneous [55] depending on their developmental origin [17, 21, 100], differentiation stage, regional [87, 114] and anatomical location [30, 45, 74, 87], sex [19, 117], age [25, 69, 96, 112, 117] and transcriptome [55] and that this pattern of heterogeneity changes in disease [55].
This year is the centenary of the publication of the first description of oligodendroglia, beautifully drawn by Del Río Hortega, after using his newly developed silver carbonate procedure to stain cells in 1921 [94]. He named these cells oligodendroglia as glial cells with few processes (Greek: oligos = “few”, dendron = “tree”, or more freely “process”) and he described finding them throughout the CNS in grey and white matter [94]. The existence of these cells was doubted initially, but accepted in 1924, with support also from the American-Canadian neurosurgeon Wilder Penfield [86]. In 1928, Del Rio Hortega published a comprehensive overview of oligodendroglia, describing their heterogeneity in terms of their morphology and he speculated that they were involved in the formation of myelin, similarly to the Schwann cells of the peripheral nervous system [95]. One hundred years on, it seems timely to review where we have reached in terms of understanding different types of oligodendroglial heterogeneity, how it occurs, and what it means for CNS health and disease, across the human life course. We will focus on humans where possible, as rodents, used in preclinical models have relatively little white matter in comparison to brain volume [106, 127], and so may not well reflect human physiology. When considering disease, we will also focus mostly on oligodendrocyte heterogeneity in the classical CNS demyelinating disease of adulthood—multiple sclerosis (MS). Understanding human oligodendroglial functional heterogeneity may help us understand CNS health and disease, leading to improved therapeutic strategies.
What is heterogeneity?
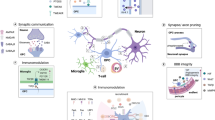
The word heterogeneity is derived from the Greek ‘heteros’ meaning different and ‘genos’ meaning ‘kind’ and so can be defined by diversity in state or content, which we will interpret here as any differences in distinguishing oligodendroglia from each other. According to this definition, oligodendroglia are heterogeneous in their developmental origin [17, 21, 100], morphology [87] and capacity to myelinate [21] depending on their regional niche (grey matter vs. white matter) [87, 114], anatomical site [30, 45, 74, 87], donor sex [19, 117] and donor age [25, 69, 96, 112, 117] (Fig. 1). These differences may be intrinsic [7] and/or extrinsic with cues from the tissue environment [36, 71, 83, 114], including the calibre of the axon being myelinated [7, 52, 80]. We will address these in turn.
Oligodendroglial heterogeneity as defined by developmental source
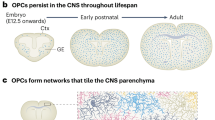
In development, OPCs of the cerebral hemispheres, cerebellum and spinal cord are derived from neural stem cells (NSCs) at different locations. In mouse forebrain development, OPCs are derived from NSCs in three waves. The first wave of OPCs originate from the subventricular zone (SVZ) from the medial ganglionic eminence and the anterior entopeduncular area at embryonic day E12.5 [61]. However, in normality, these are completely replaced by a second wave of OPCs from the lateral and caudal ganglionic eminences by the age of E15.5 and a third wave from the dorsal SVZ into the cortex [61, 101]. Cerebellar OPCs have been shown to mostly originate from extracerebellar brain regions [39], migrating from the metencephalic ventral rhombomere 1 region arising at E11.5 and arriving E16.5–E18.5, proliferating further and being added to by OPCs from the cerebellar ventricular zone [42] and after birth by OPCs from the fourth ventricle [92, 128]. In the spinal cord, the first OPCs arise after E12.5 from the ventrally located premotor neuron domain [88, 93, 118] and later (E15.5) from more dorsal precursor domains of the spinal cord [17, 31], but by birth, these dorsally derived OPCs make up only 10–20% of the OPCs [31, 113]. These developmental OPC generation routes appear conserved in humans [57, 58, 78, 90], but there may also be an additional route as primates show an enlarged outer ventricular zone during development compared to rodents [29, 111, 125] and progenitor cells in this zone produce not only neurons [9, 70] but also OPCs [49]. Furthermore, a recent study has shown the presence of OPCs in human embryos at post conception week 8–11 which is earlier than previously assumed and which supports the notion that human oligodendroglia develop in several waves in parallel to what is known in mice [14].
It is not fully understood if this regional difference in origin translates into biological differences in later oligodendroglial function, and if these oligodendroglia are intrinsically different or influenced by the local environment. The transcription factors expressed by the three waves of OPCs in the mouse forebrain are different, with ventral, medial and dorsal wave OPCs expressing Nkx2.1, Gsh32 and Emx1, respectively [61], suggesting intrinsic differences that may lead to different functions. Furthermore, dorsally derived mouse spinal cord OPCs show enhanced proliferation, recruitment and differentiation into remyelinating oligodendrocytes in response to demyelinating injury in vivo compared to ventrally derived OPCs, although this difference declines with age [21, 130]. The oligodendrocytes subsequently differentiating from these spinal cord OPCs also appear intrinsically different as cultured rodent spinal cord oligodendrocytes produce myelin sheaths that are longer and thicker than forebrain-derived oligodendrocytes even when wrapping the same diameter polymer fibres [7]. However, oligodendrocytes from the same origin but residing in white or grey matter also behave differently [26, 105], suggesting that the local environment where oligodendroglia reside also changes their function. We will return to heterogeneity related to location below, but this evidence suggests that developmental oligodendroglial heterogeneity depends on a combination of regional intrinsic and environmental influences, and that there may be cellular expression markers to define this.
Differentiation markers of oligodendroglial heterogeneity
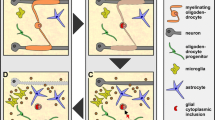
The advent of immunohistochemistry has allowed us to use protein molecular markers to distinguish oligodendroglia from the different developmental origins and also different stages in the differentiation of oligodendroglia in development and adulthood (Fig. 2). This has enabled us to understand stages in the continuum of oligodendroglial differentiation from oligodendrocyte precursor cells (OPCs) to immature oligodendrocytes and then into mature myelinating oligodendrocytes. The markers commonly used to distinguish these are illustrated in Figs. 2 and 3, and will just be stated here as they have been reviewed extensively [64, 77]. These are perhaps more described and used in the in vitro cell culture setting and in development rather than in tissue in adult human CNS, except in the context of a response to pathology where the developmental oligodendroglial differentiation pathway is thought to be at least partly recapitulated in the regenerative response. In preclinical models, or even human stem cell-derived oligodendrocytes cultured in vitro, it is possible to add reporters to reflect the different stages along the differentiation pathway in real time and live. This is particularly effective in the embryonic zebrafish where it is relatively straightforward to use multiple different fluorescent reporters in live fish imaged over time to see the differentiation process in vivo. Live imaging of cortical oligodendroglia with fluorescent reporters is also possible through skull windows in mice, but imaging deeper white matter oligodendroglia is more difficult. These two methods have revealed a heterogeneity of oligodendroglial responses to damage, with remyelinating oligodendrocytes able to be derived from OPCs (as in development) but also surviving oligodendrocytes can extend new processes for myelin repair [6], though perhaps less successfully [82]. There is evidence suggesting this may be similar in humans, as radionucleotide labelling in humans post atomic bomb testing shows that remyelinated lesions in MS brain contain oligodendrocytes which still contain high levels of the radionucleotide, suggesting they are old, rather than from proliferative OPCs which will have diluted out this label [123]. Further evidence from human biopsies of acute and inflammatory MS lesions shows that, unexpectedly, there is not an increase in OPCs or immature oligodendroglia in such lesions, suggesting that these may remyelinate using the oligodendrocytes that are already there [44]. Finally, the mistargeting of myelin to neuronal cell bodies that is seen in zebrafish oligodendrocytes that survive and remyelinate is also found in human MS tissue in the peri-demyelinated lesion areas [82]. These live-imaging data from other species have provided us with information that it is impossible to obtain from humans and means that we need to think again about the heterogeneity of responses of human oligodendrocytes to pathology.
Examples of molecular markers of oligodendroglia on human postmortem brain tissue. a Colorimetric immunohistochemistry using an antibody against OLIG2. b Colorimetric immunohistochemistry using an antibody against NOGOA. c In situ hybridisation (BaseScope Duplex) using probes against RBFOX1 and OLIG2 demonstrating heterogeneity. The pink arrows mark OLIG2-positive oligodendroglia that are RBFOX1 negative, the blue arrow indicates a cell that is RBFOX1 positive, but OLIG2 negative (not oligodendroglial) and the black arrow a double positive cell. d Immunofluorescence using antibodies against OLIG2, RBFOX1 and the nuclear marker Hoechst, again demonstrating heterogeneity. The white arrow marks a cell with only a OLIG2 marker, the red arrow a cell with only a RBFOX1 marker (not oligodendroglial) and the pink arrow an OLIG2+ cell also expressing RBFOX1
As well as using protein markers, we can use RNA in situ hybridisation probes to detect RNA markers of oligodendroglia. Improved technology using multiple short probes to different neighbouring parts of the same RNA, instead of full-length probes, with highly efficient signal amplification, has allowed specific and sensitive detection of single RNA molecules in human tissue—both fresh–frozen and paraffin [27] (Fig. 3). This technology also now allows detection of different splice variants, single base mutations and microRNAs or long non-coding (lnc) RNAs. The latter is of particular importance to oligodendroglial heterogeneity as some are selectively expressed in subsets of oligodendrocytes [55], and there is increasing interest in lncRNAs in driving oligodendroglial differentiation in health and regeneration in disease [43, 60, 119].
These molecular markers alter over differentiation in development and disease, along with well-described morphological changes from bipolar OPCs to highly branched complex oligodendrocytes.
Morphological heterogeneity
In 1928, Del Rio Hortega published his comprehensive review of oligodendrocytes in the CNS, using his new Golgi–Hortega technique initially describing three types of cells on the basis of where they were found: interfascicular (cells aligned in rows along axonal tracts); perineuronal (next to neuronal cell bodies) and perivascular (around blood vessels). In addition, he then classified them on their morphology, dividing them into 4 types: Type I with small rounded cell bodies and many fine processes emerging radially towards thinly myelinated axons, Type II with polygonal cell bodies, fewer and thicker processes directed to axons longitudinally, Type III with a large soma and one to four processes directed towards axons and Type IV with elongated cell bodies with one or two processes [77, 84]. This allowed him to speculate that oligodendrocytes were involved in the formation of myelin, similarly to the Schwann cells of the peripheral nervous system and emphasises how detailed descriptions of anatomy pave the way to more understanding. Although these categories are no longer used, in favour of molecular markers, again live-imaging work from the embryonic zebrafish using oligodendroglial reporters shows heterogeneous morphology in OPCs with functional differences by transcriptomic analysis [73].
Subregional location determining heterogeneity
Del Rio Hortega also observed that his four morphological types were differentially distributed across CNS tissue, suggesting that grey and white matter OPCs may give rise to oligodendrocytes of different morphology, e.g. Type II oligodendrocytes were described as white matter specific [87, 94]. Although there is a greater oligodendroglia density in the white matter, a proportion of them also resides in the grey matter and their behaviour at these different locations differs, at least in the mouse. Mouse white matter OPCs have been shown to proliferate more readily when stimulated [46] and produce more mature oligodendrocytes [114] than mouse grey matter OPCs. This difference is due to both intrinsic as well as extrinsic cues, because when white matter OPCs were transplanted into grey matter, they divided and differentiated more efficiently than grey matter OPCs, but not as efficiently as in the white matter environment [114]. As these white matter and grey matter OPCs were derived from the same source of progenitor cells, the observed difference may be due to environmental cues at key steps of the differentiation with a prolonged effect on the cells.
Anatomical location determining heterogeneity
As well as grey/white matter differences, Del Rio Hortega described morphological oligodendrocyte differences with anatomical region, for example describing oligodendrocyte type IV as only present in the brain stem and in the spinal cord [87, 95]. It has been long known that spinal cord oligodendrocytes produce longer and thicker myelin sheaths than those of other CNS regions [45], but it remained unclear if this was due to intrinsic differences related to their different developmental origin, or secondary to environmental cues such as axonal thickness. Axons in the spinal cord have larger diameters on average than those in the brain and axonal thickness positively correlates with myelin thickness [7, 52, 80]. One study dissected the intrinsic and extrinsic components of this finding by culturing murine oligodendrocytes differentiated from spinal cord or cortical OPCs on synthetic polymer fibres [7]. This study showed that myelin sheath length was associated with both the origin of the cells and the diameter of the fibres. This indicated that oligodendroglia of different anatomical sites are intrinsically different but they can still recognise and adapt to different environmental situations. These functional differences may be reflected in the transcriptional signatures of oligodendroglia as in mice different subtypes of oligodendrocytes as defined by distinct marker gene expressions were found in brain compared to spinal cord, but also within different regions of the spinal cord. However, using lineage tracing, these oligodendrocyte subtypes did not seem to derive from OPCs from the different developmental origins [30], suggesting that even apparent intrinsic differences may not be due to the developmental origin but to early environmental cues during differentiation, at least in mice. This multiplicity of OPC developmental types, but convergence of these into all differentiating into the same subtypes of oligodendrocytes begs the question of why this system has evolved. One can speculate that the OPCs have other functions e.g. in axonal pruning [15, 122], unrelated to the oligodendrocyte differentiation, or that we still do not understand the intricacies of oligodendrocyte heterogeneity at high enough resolution.
Heterogeneity related to sex
As well as regional differences in oligodendroglia, are there fundamental differences between female and male oligodendroglia? These may be determined by their sex chromosome content, or by influences of the sex steroid hormones. Oligodendroglia are influenced by estrogens, progestogens and androgens. Estrogens have been implicated in directly promoting neural stem cell and OPC proliferation, differentiation and myelination from in vitro experiments, as well as ameliorating responses to neuroinflammatory conditions such as Experimental Allergic Encephalomyelitis (EAE) in preclinical models and MS in humans [4, 72] (reviewed in [62]). However, the latter effect directly on oligodendroglia in vivo is difficult to discern from an indirect effect on the immune response and on other cells. Progesterone has similarly been positively implicated in enhancing oligodendrocyte formation and myelination [35, 37]. Testosterone has also been shown to enhance remyelination in male mice [51], especially compared to castrated male mice [10], at least in part due to a direct effect on oligodendroglia. This androgen response may also act through signalling through the androgen receptor on microglia and astrocytes with contribution from the hedgehog pathway, and with a secondary effect on remyelination [66]. In developmental myelination, oligodendroglia will be exposed to maternal sex hormones, and oligodendroglia involved in myelin maintenance and remyelination are exposed to the sex hormones of the adult. In MS, although fewer males are diagnosed with the disease, their prognosis is worse, and remyelination may be more extensive in early MS postmortem lesions in females than in males [63]. There is also emerging exciting evidence that gonosomal genes may directly influence brain cell function in neurodegenerative diseases [16, 23, 67] rather than simply through sex hormone differences. In a genetically modified Alzheimer’s disease mouse model, the expression of the X-linked gene Kdm6a, one of the few genes still expressed on the otherwise inactivated X-chromosome, was associated with longer survival in both sexes [23]. In addition, the inhibition of the Y-chromosomal gene SRY in a murine Parkinson’s disease model reduced clinical signs and improved dopaminergic neuron survival [67]. However, the effect of gonosomal genes on oligodendroglia function remains to be explored and little is known specifically about sex-related differences in oligodendroglia in humans and the contribution to function in health and disease.
Heterogeneity related to age
Over the human lifespan, oligodendroglia can form and maintain myelin. The first OPCs emerge in the developing human brain at 10 weeks’ gestation, followed by an expansion of the population (15–20 weeks) [57, 58, 97] and myelination commences at 30 weeks’ gestation, but most occurs postnatally [50, 53, 106]. This progresses in a rostral to caudal manner, with spinal cord and cerebellum myelination before the cerebral hemispheres, finishing with the prefrontal cortex [53, 57, 58]. C14 Carbon dating in humans suggests that oligodendrocyte numbers are stable after 5 years of age [124], with slow myelin turnover, although there is evidence from rodents [103, 110, 121] and humans [104], that neuronal activity can drive new myelination, for example by exercise and learning new tasks. With ageing, there is reduced myelin integrity [8], occurring initially in areas of the brain that were myelinated first, as seen on micro-structural magnetic resonance imaging (MRI) [40], and at a pathological level, there is detachment of the paranodal loops at the ends of the myelin sheaths [48, 107], suggesting poorer oligodendroglial function.
This heterogeneity in oligodendroglia/myelin over the normal life course may be important in understanding their vulnerability and response to diseases that occur at different ages. The developmental timing of OPC emergence and myelination make oligodendroglia vulnerable to perinatal hypoxic and inflammatory injury, for example due to premature birth. This reduces the maturation of immature oligodendrocytes into myelinating oligodendrocytes, leading to white matter injury and can result in cognitive and/or physical developmental defects [116]. Genetic diseases causing oligodendrocyte defects can cause dysmyelination (such as in Pelizaeus–Merzbacher disease [84]) which usually leads to early death.
In early to mid-adulthood, the commonest cause of non-traumatic neurological disability in the ‘first world’ is MS, where focal areas of demyelination occur in the CNS with oligodendrocyte loss, and attempts at remyelination (reviewed in [32]). Furthermore, in older age, cerebral small vessel disease is very common [108] and is characterised by white matter changes on magnetic resonance brain scans which correlate with clinical cognitive impairment [24] and endothelial cell, oligodendroglial and myelin changes by pathology [89]. Oligodendrocyte pathology has also been increasingly implicated in other neurodegenerative diseases usually associated with ageing that were previously thought to be classically neuronal, such as progressive supranuclear palsy [2, 99], Huntington’s Disease [109] and Parkinson’s disease [1], as well as psychiatric diseases such as major depressive disorder [81].
These changes suggest that human oligodendroglia change with age, but it is uncertain whether different subsets of these are more prominent in development, adult myelin maintenance, and ageing, whether these confer differential susceptibility to pathology or therapeutic responses to, for example, pro-remyelination drugs. To try and understand these different nuances in oligodendroglial behaviour over age and in pathology, researchers have turned to assessing transcriptional heterogeneity, particularly at a single cell level.
Transcriptional heterogeneity
Recent technology has allowed us to examine heterogeneity within oligodendroglia at an unprecedented level by examining transcripts from single cell RNA sequencing (scRNAseq) or single nuclear RNA sequencing (snRNAseq), allowing phenotyping cells using multiple gene transcripts as a readout of their function. This is now possible at scale, allowing classification of oligodendroglia using similarities (and differences) between these transcripts, for tens to hundreds of thousands of cells, limited currently only by both cost and computational power for the analysis, albeit with, at least with droplet technology, capture of considerably less than half of the expressed transcripts [129]. There has been an explosion of information using these tools about many different cells from many different tissues of a variety of species, all showing that the broad groupings of cells that we have worked with for years contain disparate cells. This is being at least in part driven by an international initiative to sequence all of the different cells of the human body and have the data as open access within the Human Cell Atlas [91] (www.humancellatlas.org).
First scRNAseq studies in adult and juvenile mouse CNS samples found distinct oligodendroglia states and markers for differentiation stages [74, 126]. The oligodendroglia states varied with tissue region indicating regional heterogeneity. At the same time, a proof of principle scRNAseq study showed that similar investigations are possible in adult and foetal human CNS [22]. As the technology improved, snRNAseq allowed use of archived human post-mortem fresh–frozen tissue, significantly increasing the breadth of tissue analysed [41, 65]. Both mouse and human studies show that all expected cell types can be identified using canonical marker genes and that those cell types can typically be further subclustered into groups of cells that express similar genes and differ from other subclusters in their gene expression which also indicates a distinction of differentiation stages and cellular function [55].
In humans, axons are mostly myelinated post-partum and therefore human foetuses lack proper myelinated white matter. However, scRNAseq has shown that human foetal oligodendroglia express EGFR from 20 gestational weeks onwards and include pre-OPCs, OPCs, and SPARCL1-positive OPCs that have transcriptional similarities with the astrocyte lineage [33]. Different to murine OPCs, where typically one daughter cell continues to divide and produce more OPCs whereas the other daughter cell differentiates, in human foetuses, OPCs divide symmetrically several times before they start differentiating [49]. This may be one of the mechanisms that leads to a ~ 3000 times greater relative white matter in humans compared to rodent animal models. The expression of PCDH15 in OPCs leads to daughter cell repulsion which is likely associated with new OPCs infiltrating other brain regions [49], but also allows further proliferation which is inhibited by high OPC density [49].
Although the myelin transcriptome/proteome of mice and humans overlap, these developmental differences, and RNAseq and proteomic studies have shown that there are distinct components present only in one or the other species [54]. In addition, animals are incomplete preclinical models for MS, with a lack of description of MS in any animal species including non-human primates except perhaps in a population of Japanese macaques with an endemic herpesvirus infection, where a low percentage of animals each year develops an MS-like disease [5, 38]. This underlines the importance to better understand human oligodendroglia using human tissue and cells.
Sc/snRNAseq studies in human adults mostly focus on postmortem tissue, simply due to the pragmatics of obtaining samples, comparing oligodendroglial subgroups between mouse and human, and between human health and different neurological diseases.
Using human postmortem brain tissue from the MRC Edinburgh Sudden Death Brain Bank, with pathologically normal human brain, and from the MS Society UK MS tissue bank with pathologically proven MS, we performed snRNAseq using 10 × technology on 20 different white matter brain samples from 4 MS patients and 5 controls [55]. Using this dataset, we subdivided our oligodendroglia into 8 subclusters, including OPCs, committed oligodendrocyte precursors, and 6 types of oligodendrocytes all expressing myelin protein transcripts such as MBP and MOG, but with other transcriptional differences [55]. These subclusters are identifiable by single or groups of transcript markers, some of which are defined in Fig. 4b. We identified that in MS brain, there are fewer OPCs and a shift in the proportions of oligodendrocytes in these subclusters, showing that MS pathology leads to changes in oligodendrocyte functional heterogeneity with some clusters over-represented and some under-represented (Fig. 4a). But what do these differences in subclusters mean functionally, and how does the shift in oligodendroglial composition affect the biology/pathology in MS? We are not yet certain, as we inevitably only have one timepoint in postmortem human data, but examination of the gene transcripts allows us to speculate, generate hypotheses and in the future test these in other systems. Comparison of the RNA transcripts using gene ontology analysis suggests that over half of the transcripts in Oligo4 cluster cells relate to myelination—more than with any of the other subclusters, and suggesting that these cells are been driven to try to actively myelinate/remyelinate. This cluster is under-represented in MS tissue, allowing the hypotheses that either these are selectively vulnerable in MS, or that these cells fail to regenerate after global loss of oligodendrocytes in MS. It is tempting to speculate that these oligodendrocytes are the ones that might be the most useful to generate for successful remyelination in MS. Conversely, Oligo1 subcluster cells have relatively few RNAs that correspond to myelin genes, and instead express transcripts more related to signalling and metabolism. This leads to an interpretation that these cells have completed active myelin formation and are simply maintaining their sheaths, and instead are focussing on signalling and metabolic support to the underlying axon—an essential role for maintenance of myelin throughout life. These are also under-represented in MS tissue, suggesting that few oligodendrocytes even if they achieve remyelination, manage to then provide the same support as in normality. Instead, the subclusters that are over-represented in MS are ones that we believe to be more immature, with markers such as OPALIN, and perhaps represent the block in oligodendrocyte formation which has been much described in the literature [32]. Another cluster over-represented in MS brain, called ‘immune-oligos,’ expresses major histocompatibility complex II (MHC-II) genes, confirmed at the protein level, similarly to in EAE, a mouse T cell-driven model of MS [28]. This may suggest that in MS, some oligodendrocytes are capable of presenting antigen, which may contribute to the immune-driven MS pathology. Whether the oligodendroglia in these different clusters are fixed in these states in adult tissue, or can transition to and from other functional states is also not known, and will only be resolved by functional modelling of live cells, as RNAseq simply gives us a snapshot of the function of each cell at the time of freezing the tissue. If fixed, then clusters represent co-existing distinct oligodendrocyte morphologies, with static presumed functional and regional differences, perhaps suggesting a division of labour for these cells, or that certain oligodendrocytes myelinate certain neuronal axonal types, etc. However, if, in adulthood, they are dynamic and oligodendrocytes can change gene expression signatures, this suggests possibilities for adaptation to the tissue environment over time and potentially to pathology. Bioinformatically, trajectory inference methods have been developed to predict how cells differentiate from immature to mature cells, ordering them based on their transcriptional similarity. However, these methods are not designed to identify transitions in adult tissue at one timepoint, where transitions may be more stochastic and in multiple directions between clusters.
Oligodendrocyte heterogeneity at the RNA level. a Simplified standard workflow of an snRNAseq experiment. Shown data are adapted from Jäkel & Agirre et al. [55] and shows postmortem human brain affected/unaffected by multiple sclerosis. b A selection of marker genes visualises transcriptional differences between oligodendrocyte states. c Gene ontology analysis of differentially expressed genes in Oligo4 and Oligo1 indicates functional differences in oligodendroglia states.
The next question is whether this skewed oligodendroglial heterogeneity in MS is MS-specific or a more general response to pathology. This is not yet fully resolved, but in a similar study, using tissue from donors with major depressive disease, by direct comparison of the datasets, the authors identified some similar oligodendroglial subclusters [81]. Many further sc/snRNAseq studies have identified altered human oligodendroglia heterogeneity with different diseases including further work in multiple sclerosis [102], Alzheimer’s disease [75, 79], Huntington’s disease [3], Parkinson’s disease [1], neuropsychiatric disorders [12, 13, 81], and glioblastoma [20]. In these studies, inevitably, various different numbers of oligodendrocyte and OPC clusters were identified, likely depending on probabilities and decisions about data pre-processing and resolutions in the analysis, as well as potential regional differences and differences in the proportions of grey and white matter. These latter possible differences have not yet been well explored in human tissue with these techniques. However, despite this, parallels across datasets are clear: OPALIN and RBFOX1 are clearly reported as markers for two different oligodendroglia states [55, 79]. In addition, both of those studies [55, 79] detected “Immuno-oligos” that express major histocompatibility complex class II genes suggesting antigen presentation capabilities, and these were enriched in both MS and Alzheimer’s disease compared to control groups [55, 79]. As the datasets increase in number and size, we will need a concerted response to integrate relevant datasets from the human white and grey matter, in health and across a variety of diseases, with a subsequent agreed nomenclature to allow us to improve our understanding of normal and pathological transcriptional distinctions among the oligodendroglia population.
These findings have increased interest in the oligodendroglial influence in classical neurodegenerative diseases, partly as therapeutic options are wider with these cells, which, unlike neurons can regenerate. This leads us back to our section on using markers to identify oligodendroglia, as this transcriptomic analysis gives us new markers of oligodendroglia, both at the broad level, e.g. OPCs and mature oligodendrocytes and at the subcluster level, allowing us to validate the presence/number of these oligodendroglia in the CNS in different regions, diseases, etc. without needing to perform RNAseq. Furthermore, there are now new and improved techniques to spatially identify several or multiple of these markers in combination with either RNA probes or antibodies on human postmortem tissue, both in frozen and formalin-fixed paraffin sections. For example, previous studies including ours [55, 79] have shown that a subset of oligodendrocytes expresses the RNA Binding Fox-1 Homolog 1 (RBFOX1) which can be visualised and quantified using in situ hybridisation or antibody-staining techniques (Fig. 3). To this, we can add the even more recent development of on-tissue RNA-sequencing, giving spatial information, with some techniques at single cell resolution covering all genes, single cell epigenomics (e.g. ATACseq), lipidomics and proteomics and it is a very fast-moving field (reviewed in [56]). One can see that the knowledge of (and data from) oligodendroglial heterogeneity will only expand, with the limit perhaps being the ability to analyse such data and cost.
Can we reconcile these sources of heterogeneity?
Above we discussed eight different sources of oligodendrocyte heterogeneity. For clarity and convenience, each source was discussed separately, but these clearly correlate, interact or even perhaps cause one another. For example, in development, OPCs change in morphology from a bipolar cell to one with many processes wrapping axons and this is accompanied by a transcriptional switch with the expression of myelin genes, allowing the formation of the myelin sheath. Similarly, the different developmental origin for example of spinal cord OPCs may influence their regional heterogeneity, their transcriptional signatures and this may determine (at least in part) their ability to myelinate axons with longer and thicker myelin sheaths than brain ones [7].
The current development of spatial transcriptomics technologies to measure multiplexed gene expression in situ [47, 76, 98] will allow us to better correlate transcriptional variation, regionality, morphology and the interactions between heterogeneous oligodendroglia and other CNS cells, not just restricted to whether different oligodendrocyte ensheath different axons, but in their interactions with astrocytes, microglia, vascular cells, immune cells, each other and indeed with different subtypes of these. This is clearly essential for understanding normal and pathological brain function, and will involve complex data analysis in three dimensions, preferably over time. We are likely to increase the sources and complexity of oligodendroglia (and other cell) heterogeneity by including measures of chromatin accessibility, DNA methylation, spatial gene expression, single cell proteomics, and lipidomics and integration of these data modalities is going to be challenging but hopefully exciting. The first step here (with a focus on neurons) has very recently been published through the Brain Research through Advancing Innovative Neurotechnologies (BRAIN) Initiative Cell Census Network using multimodal and multispecies investigations of the primary motor cortex [18].
Challenges and future in studying human oligodendroglia heterogeneity
The challenges of understanding oligodendroglial heterogeneity are large, but importantly, although -Omics predict function, we do not really know yet what impact these changes have on the actual biological function of the cell. For example, do some subclusters represent oligodendrocytes which are primed to remyelinate, others which are dying and yet others which are designed to provide metabolic support to underlying axons? There are hints of this from the bioinformatics, but this needs to be tested experimentally. In the context of MS, are there some individuals who have more of the oligodendrocytes ‘good’ at remyelination, with less chance of disease progression and vice versa, perhaps matching the ‘good’ and ‘bad’ remyelinators detected by Positron emission tomography (PET) scanning in MS [11]. This leaves us excited for the future of using such information to identify disease or disease progression/prognosis if we can convert these findings into markers measurable in the living person. Is there a generic ‘pathological’ oligodendrocyte and/or a ‘regenerative’ one across all diseases, or some diseases? Can we use drugs to transition oligodendrocytes from such a ‘pathological’ state to a ‘regenerative’ one as a new therapeutic strategy? These are not easy questions to answer, as human oligodendroglia reflecting adult in vivo biology are difficult to grow in culture. However, progress is being made with human stem cell-derived cells used to form cultured organoids [59, 85] and injected into immunocompromised mouse brain [120] showing more fidelity to adult human oligodendroglia in vivo.
Conclusion
One hundred years after the first description of oligodendrocytes, we are now in the exciting position that we have discovered that oligodendroglia vary in their function, at least in part depending on origin, location, age, sex and pathology, suggesting that this may be driven by a combination of regional intrinsic and environmental influences. Oligodendroglia are also seeing a revived popularity, with oligodendroglial components now implicated in many neurodegenerative diseases, and due to their potential tractability to therapeutics. We predict that in the future, we will increasingly use markers of these different subtypes of oligodendroglia to predict disease course and stratify patients for therapies, but we have some way to go!
References
Agarwal D, Sandor C, Volpato V, Caffrey TM, Monzón-Sandoval J, Bowden R et al (2020) A single-cell atlas of the human substantia nigra reveals cell-specific pathways associated with neurological disorders. Nat Commun 11:1–11. https://doi.org/10.1038/s41467-020-17876-0
Ahmed Z, Asi YT, Lees AJ, Revesz T, Holton JL (2013) Identification and quantification of oligodendrocyte precursor cells in multiple system atrophy, progressive supranuclear palsy and Parkinson’s disease. Brain Pathol 23:263–273. https://doi.org/10.1111/j.1750-3639.2012.00637.x
Al-Dalahmah O, Sosunov AA, Shaik A, Ofori K, Liu Y, Vonsattel JP et al (2020) Single-nucleus RNA-seq identifies Huntington disease astrocyte states. Acta Neuropathol Commun 8:19. https://doi.org/10.1186/s40478-020-0880-6
Arvanitis DN, Wang H, Bagshaw RD, Callahan JW, Boggs JM (2004) Membrane-associated estrogen receptor and caveolin-1 are present in central nervous system myelin and oligodendrocyte plasma membranes. J Neurosci Res 75:603–613. https://doi.org/10.1002/jnr.20017
Axthelm MK, Bourdette DN, Marracci GH, Su W, Mullaney ET, Manoharan M et al (2011) Japanese macaque encephalomyelitis: a spontaneous multiple sclerosis-like disease in a nonhuman primate. Ann Neurol 70:362–373. https://doi.org/10.1002/ana.22449
Bacmeister CM, Barr HJ, McClain CR, Thornton MA, Nettles D, Welle CG et al (2020) Motor learning promotes remyelination via new and surviving oligodendrocytes. Nat Neurosci 23:819–831. https://doi.org/10.1038/s41593-020-0637-3
Bechler ME, Byrne L, Bechler ME, Byrne L (2015) CNS Myelin Sheath Lengths Are an Intrinsic Property of Oligodendrocytes. Curr Biol 25:2411–2416. https://doi.org/10.1016/j.cub.2015.07.056
Bennett IJ, Madden DJ, Vaidya CJ, Howard DV, Howard JH (2010) Age-related differences in multiple measures of white matter integrity: a diffusion tensor imaging study of healthy aging. Hum Brain Mapp 31:378–390. https://doi.org/10.1002/hbm.20872
Betizeau M, Cortay V, Patti D, Pfister S, Gautier E, Bellemin-Ménard A et al (2013) Precursor diversity and complexity of lineage relationships in the outer subventricular zone of the primate. Neuron 80:442–457. https://doi.org/10.1016/j.neuron.2013.09.032
Bielecki B, Mattern C, Ghoumari AM, Javaid S, Smietanka K, Ghanem CA et al (2016) Unexpected central role of the androgen receptor in the spontaneous regeneration of myelin. Proc Natl Acad Sci U S A 113:14829–14834. https://doi.org/10.1073/pnas.1614826113
Bodini B, Veronese M, García-Lorenzo D, Battaglini M, Poirion E, Chardain A et al (2016) Dynamic imaging of individual remyelination profiles in multiple sclerosis. Ann Neurol 79:726–738. https://doi.org/10.1002/ana.24620
Bøstrand SMK, Williams A (2021) Oligodendroglial heterogeneity in neuropsychiatric disease. Life 11:125. https://doi.org/10.3390/life11020125
Brenner E, Tiwari GR, Kapoor M, Liu Y, Brock A, Mayfield RD (2020) Single cell transcriptome profiling of the human alcohol-dependent brain. Hum Mol Genet 29:1144–1153. https://doi.org/10.1093/HMG/DDAA038
van Bruggen D, Pohl F, Langseth CM, Kukanja P, Lee H, Kabbe M et al (2021) Developmental landscape of human forebrain at a single-cell level unveils early waves of oligodendrogenesis. bioRxiv. https://doi.org/10.1101/2021.07.22.453317
Buchanan J, Elabbady L, Collman F, Jorstad NL, Bakken TE, Ott C, et al (2021) Oligodendrocyte precursor cells prune axons in the mouse neocortex. bioRxiv 2021.05.29.446047
Caceres A, Jene A, Esko T, Perez-Jurado LA, Gonzalez JR (2020) Extreme downregulation of chromosome Y and Alzheimer’s disease in men. Neurobiol Aging 90:150.e1-150.e4. https://doi.org/10.1016/j.neurobiolaging.2020.02.003
Cai J, Qi Y, Hu X, Tan M, Liu Z, Zhang J et al (2005) Generation of oligodendrocyte precursor cells from mouse dorsal spinal cord independent of Nkx6 regulation and Shh signaling. Neuron 45:41–53. https://doi.org/10.1016/j.neuron.2004.12.028
Callaway EM, Dong H-W, Ecker JR, Hawrylycz MJ, Huang ZJ, Lein ES et al (2021) A multimodal cell census and atlas of the mammalian primary motor cortex. Nature 598:86–102. https://doi.org/10.1038/s41586-021-03950-0
Chowen JA, Garcia-Segura LM (2021) Role of glial cells in the generation of sex differences in neurodegenerative diseases and brain aging. Mech Ageing Dev 196:111473. https://doi.org/10.1016/j.mad.2021.111473
Couturier CP, Ayyadhury S, Le PU, Nadaf J, Monlong J, Riva G et al (2020) Single-cell RNA-seq reveals that glioblastoma recapitulates a normal neurodevelopmental hierarchy. Nat Commun. https://doi.org/10.1038/s41467-020-17186-5
Crawford AH, Tripathi RB, Richardson WD, Franklin RJM (2016) Developmental origin of oligodendrocyte lineage cells determines response to demyelination and susceptibility to age-associated functional decline. Cell Rep 15:761–773. https://doi.org/10.1016/j.celrep.2016.03.069
Darmanis S, Sloan SA, Zhang Y, Enge M, Caneda C, Shuer LM et al (2015) A survey of human brain transcriptome diversity at the single cell level. Proc Natl Acad Sci U S A 112:7285–7290. https://doi.org/10.1073/pnas.1507125112
Davis EJ, Broestl L, Williams G, Garay BI, Lobach I, Devidze N et al (2020) A second X chromosome contributes to resilience in a mouse model of Alzheimer’s disease. Sci Transl Med. https://doi.org/10.1126/SCITRANSLMED.AAZ5677
Debette S, Markus HS (2010) The clinical importance of white matter hyperintensities on brain magnetic resonance imaging: Systematic review and meta-analysis. BMJ 341:288. https://doi.org/10.1136/bmj.c3666
Defrancesco M, Egger K, Marksteiner J, Esterhammer R, Hinterhuber H, Deisenhammer EA et al (2014) Changes in white matter integrity before conversion from mild cognitive impairment to Alzheimer’s disease. PLoS ONE. https://doi.org/10.1371/journal.pone.0106062
Dimou L, Simon C, Kirchhoff F, Takebayashi H, Götz M (2008) Progeny of Olig2-expressing progenitors in the gray and white matter of the adult mouse cerebral cortex. J Neurosci 28:10434–10442. https://doi.org/10.1523/JNEUROSCI.2831-08.2008
Erben L, Buonanno A (2019) Detection and quantification of multiple RNA sequences using emerging ultrasensitive fluorescent in situ hybridization techniques. Curr Protoc Neurosci 87:e63. https://doi.org/10.1002/cpns.63
Falcão AM, van Bruggen D, Marques S, Meijer M, Jäkel S, Agirre E et al (2018) Disease-specific oligodendrocyte lineage cells arise in multiple sclerosis. Nat Med 24:1837–1844. https://doi.org/10.1038/s41591-018-0236-y
Fish JL, Dehay C, Kennedy H, Huttner WB (2008) Making bigger brains–The evolution of neural-progenitor-cell division. J Cell Sci 121:2783–2793. https://doi.org/10.1242/jcs.023465
Floriddia EM, Lourenço T, Zhang S, van Bruggen D, Hilscher MM, Kukanja P et al (2020) Distinct oligodendrocyte populations have spatial preference and different responses to spinal cord injury. Nat Commun 11:1–15. https://doi.org/10.1038/s41467-020-19453-x
Fogarty M, Richardson WD, Kessaris N (2005) A subset of oligodendrocytes generated from radial glia in the dorsal spinal cord. Development 132:1951–1959. https://doi.org/10.1242/dev.01777
Franklin RJM, Ffrench-Constant C (2017) Regenerating CNS myelin—From mechanisms to experimental medicines. Nat Rev Neurosci 18:753–769. https://doi.org/10.1038/nrn.2017.136
Fu Y, Yang M, Yu H, Wang Y, Wu X, Yong J et al (2021) Heterogeneity of glial progenitor cells during the neurogenesis-to-gliogenesis switch in the developing human cerebral cortex. Cell Rep 34:108788. https://doi.org/10.1016/j.celrep.2021.108788
Fünfschilling U, Supplie LM, Mahad D, Boretius S, Saab AS, Edgar J et al (2012) Glycolytic oligodendrocytes maintain myelin and long-term axonal integrity. Nature 485:517–521. https://doi.org/10.1038/nature11007
Gago N, Akwa Y, Sananès N, Guennoun R, Baulieu EE, El-Etr M et al (2001) Progesterone and the oligodendroglial lineage: Stage-dependent biosynthesis and metabolism. Glia 36:295–308. https://doi.org/10.1002/glia.1117
Ghorbani S, Yong VW (2021) The extracellular matrix as modifier of neuroinflammation and remyelination in multiple sclerosis. Brain. https://doi.org/10.1093/brain/awab059
Ghoumari AM, Ibanez C, El-Etr M, Leclerc P, Eychenne B, O’Malley BW et al (2003) Progesterone and its metabolites increase myelin basic protein expression in organotypic slice cultures of rat cerebellum. J Neurochem 86:848–859. https://doi.org/10.1046/j.1471-4159.2003.01881.x
Govindan AN, Fitzpatrick KS, Manoharan M, Tagge I, Kohama SG, Ferguson B et al (2021) Myelin-specific T cells in animals with Japanese macaque encephalomyelitis. Ann Clin Transl Neurol 8:456–470. https://doi.org/10.1002/acn3.51303
Grimaldi P, Parras C, Guillemot F, Rossi F, Wassef M (2009) Origins and control of the differentiation of inhibitory interneurons and glia in the cerebellum. Dev Biol 328:422–433. https://doi.org/10.1016/j.ydbio.2009.02.008
Grydeland H, Vértes PE, Váša F, Romero-Garcia R, Whitaker K, Alexander-Bloch AF et al (2019) Waves of maturation and senescence in micro-structural MRI markers of human cortical myelination over the lifespan. Cereb Cortex 29:1369–1381. https://doi.org/10.1093/cercor/bhy330
Habib N, Avraham-Davidi I, Basu A, Burks T, Shekhar K, Hofree M et al (2017) Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat Methods 14:955–958. https://doi.org/10.1038/nmeth.4407
Hashimoto R, Hori K, Owa T, Miyashita S, Dewa K, Masuyama N et al (2016) Origins of oligodendrocytes in the cerebellum, whose development is controlled by the transcription factor, Sox9. Mech Dev 140:25–40. https://doi.org/10.1016/j.mod.2016.02.004
He D, Wang J, Lu Y, Deng Y, Zhao C, Xu L et al (2017) lncRNA Functional networks in oligodendrocytes reveal stage-specific myelination control by an lncOL1/Suz12 complex in the CNS. Neuron 93:362–378. https://doi.org/10.1016/j.neuron.2016.11.044
Heß K, Starost L, Kieran NW, Thomas C, Vincenten MCJ, Antel J et al (2020) Lesion stage-dependent causes for impaired remyelination in MS. Acta Neuropathol 140:359–375. https://doi.org/10.1007/s00401-020-02189-9
Hildebrand C, Remahl S, Persson H, Bjartmar C (1993) Myelinated nerve fibres in the CNS. Prog Neurobiol 40:319–384. https://doi.org/10.1016/0301-0082(93)90015-K
Hill RA, Patel KD, Medved J, Reiss AM, Nishiyama A (2013) NG2 cells in white matter but not gray matter proliferate in response to PDGF. J Neurosci 33:14558–14566. https://doi.org/10.1523/JNEUROSCI.2001-12.2013
Hilscher MM, Langseth CM, Kukanja P, Yokota C, Floriddia E, Nilsson M, et al (2021) Spatial cell type mapping of the oligodendrocyte lineage in the mouse juvenile and adult CNS with in situ sequencing. bioRxiv 2021.06.04.447052
Hinman JD, Peters A, Cabral H, Rosene DL, Hollander W, Rasband MN et al (2006) Age-related molecular reorganization at the node of Ranvier. J Comp Neurol 495:351–362. https://doi.org/10.1002/cne.20886
Huang W, Bhaduri A, Velmeshev D, Wang S, Wang L, Rottkamp CA et al (2020) Origins and proliferative states of human oligodendrocyte precursor cells. Cell 182:594-608.e11. https://doi.org/10.1016/j.cell.2020.06.027
Hüppi PS, Warfield S, Kikinis R, Barnes PD, Zientara GP, Jolesz FA et al (1998) Quantitative magnetic resonance imaging of brain development in premature and mature newborns. Ann Neurol 43:224–235. https://doi.org/10.1002/ana.410430213
Hussain R, Ghoumari AM, Bielecki B, Steibel J, Boehm N, Liere P et al (2013) The neural androgen receptor: A therapeutic target for myelin repair in chronic demyelination. Brain 136:132–146. https://doi.org/10.1093/brain/aws284
Ibrahim M, Butt AM, Berry M (1995) Relationship between myelin sheath diameter and internodal length in axons of the anterior medullary velum of the adult rat. J Neurol Sci 133:119–127. https://doi.org/10.1016/0022-510X(95)00174-Z
Inder TE, Huppi PS (2000) In vivo studies of brain development by magnetic resonance techniques. Ment Retard Dev Disabil Res Rev 6:59–67. https://doi.org/10.1002/(SICI)1098-2779(2000)6:1%3c59::AID-MRDD8%3e3.0.CO;2-E
Ishii A, Dutta R, Wark GM, Il HS, Han DK, Trapp BD et al (2009) Human myelin proteome and comparative analysis with mouse myelin. Proc Natl Acad Sci U S A 106:14605–14610. https://doi.org/10.1073/pnas.0905936106
Jäkel S, Agirre E, Falcão AM, Bruggen D, Lee KW, Knuesel I et al (2019) Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566:543–547. https://doi.org/10.1038/s41586-019-0903-2
Jäkel S, Williams A (2020) What have advances in transcriptomic technologies taught us about human white matter pathologies? Front Cell Neurosci 14:1–14. https://doi.org/10.3389/fncel.2020.00238
Jakovcevski I, Filipovic R, Mo Z, Rakic S, Zecevic N (2009) Oligodendrocyte development and the onset of myelination in the human fetal brain. Front Neuroanat 3:1–15. https://doi.org/10.3389/neuro.05.005.2009
Jakovcevski I, Zecevic N (2005) Sequence of oligodendrocyte development in the human fetal telencephalon. Glia 49:480–491. https://doi.org/10.1002/glia.20134
James OG, Selvaraj BT, Magnani D, Burr K, Connick P, Barton SK et al (2021) iPSC-derived myelinoids to study myelin biology of humans. Dev Cell 56:1346–1358. https://doi.org/10.1016/j.devcel.2021.04.006
Katsel P, Roussos P, Fam P, Khan S, Tan W, Hirose T et al (2019) The expression of long noncoding RNA NEAT1 is reduced in schizophrenia and modulates oligodendrocytes transcription. Npj Schizophr 5:1–9. https://doi.org/10.1038/s41537-019-0071-2
Kessaris N, Fogarty M, Iannarelli P, Grist M, Wegner M, Richardson WD (2006) Competing waves of oligodendrocytes in the forebrain and postnatal elimination of an embryonic lineage. Nat Neurosci 9:173–179. https://doi.org/10.1038/nn1620
Kipp M, Amor S, Krauth R, Beyer C (2012) Multiple sclerosis: Neuroprotective alliance of estrogen-progesterone and gender. Front Neuroendocrinol 33:1–16. https://doi.org/10.1016/j.yfrne.2012.01.001
Kuhlmann T, Goldschmidt T, Antel J, Wegner C, König F, Metz I et al (2009) Gender differences in the histopathology of MS? J Neurol Sci 286:86–91. https://doi.org/10.1016/j.jns.2009.07.014
Kuhn S, Gritti L, Crooks D, Dombrowski Y (2019) Oligodendrocytes in development myelin generation and beyond. Cells 8:1424. https://doi.org/10.3390/cells8111424
Lake BB, Chen S, Sos BC, Fan J, Kaeser GE, Yung YC et al (2018) Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat Biotechnol 36:70–80. https://doi.org/10.1038/nbt.4038
Laouarem Y, Kassoussi A, Zahaf A, Hutteau-Hamel T, Mellouk A, Bobé P et al (2021) Functional cooperation of the hedgehog and androgen signaling pathways during developmental and repairing myelination. Glia 69:1369–1392. https://doi.org/10.1002/glia.23967
Lee J, Pinares-Garcia P, Loke H, Ham S, Vilain E, Harley VR (2019) Sex-specific neuroprotection by inhibition of the Y-chromosome gene, SRY, in experimental Parkinson’s disease. Proc Natl Acad Sci U S A 116:16577–16582. https://doi.org/10.1073/pnas.1900406116
Lee Y, Morrison BM, Li Y, Lengacher S, Farah MH, Hoffman PN et al (2012) Oligodendroglia metabolically support axons and contribute to neurodegeneration. Nature 487:443–448. https://doi.org/10.1038/nature11314
Leong SY, Rao VTS, Bin JM, Gris P, Sangaralingam M, Kennedy TE et al (2014) Heterogeneity of oligodendrocyte progenitor cells in adult human brain. Ann Clin Transl Neurol 1:272–283. https://doi.org/10.1002/acn3.55
Lui JH, Hansen DV, Kriegstein AR (2011) Development and evolution of the human neocortex. Cell 146:18–36. https://doi.org/10.1016/j.cell.2011.06.030
Marangon D, Boccazzi M, Lecca D, Fumagalli M (2020) Regulation of oligodendrocyte functions: targeting lipid metabolism and extracellular matrix for myelin repair. J Clin Med 9:470. https://doi.org/10.3390/jcm9020470
Marin-Husstege M, Muggironi M, Raban D, Skoff RP, Casaccia-Bonnefil P (2004) Oligodendrocyte progenitor proliferation and maturation is differentially regulated by male and female sex steroid hormones. Dev Neurosci 26:245–254. https://doi.org/10.1159/000082141
Marisca R, Hoche T, Agirre E, Hoodless LJ, Barkey W, Auer F et al (2020) Functionally distinct subgroups of oligodendrocyte precursor cells integrate neural activity and execute myelin formation. Nat Neurosci 23:363–374. https://doi.org/10.1038/s41593-019-0581-2
Marques S, Zeisel A, Codeluppi S, van Bruggen D, Mendanha Falcao A, Xiao L et al (2016) Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science (80-) 352:1326–1329. https://doi.org/10.1126/science.aaf6463
Mathys H, Davila-Velderrain J, Peng Z, Gao F, Mohammadi S, Young JZ et al (2019) Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570:332–337. https://doi.org/10.1038/s41586-019-1195-2
Maynard KR, Collado-Torres L, Weber LM, Uytingco C, Barry BK, Williams SR et al (2021) Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat Neurosci 24:425–436. https://doi.org/10.1038/s41593-020-00787-0
Miron VE, Kuhlmann T, Antel JP, JP, (2011) Cells of the oligodendroglial lineage, myelination, and remyelination. Biochim Biophys Acta - Mol Basis Dis 1812:184–193. https://doi.org/10.1016/j.bbadis.2010.09.010
Mo Z, Zecevic N (2009) Human fetal radial glia cells generate oligodendrocytes in vitro. Glia 57:490–498. https://doi.org/10.1002/glia.20775
Morabito S, Miyoshi E, Michael N, Shahin S, Martini AC, Head E et al (2021) Single-nucleus chromatin accessibility and transcriptomic characterization of Alzheimer’s disease. Nat Genet 53:1143–1155. https://doi.org/10.1038/s41588-021-00894-z
Murray JA, Blakemore WF (1980) The relationship between internodal length and fibre diameter in the spinal cord of the cat. J Neurol Sci 45:29–41. https://doi.org/10.1016/S0022-510X(80)80004-9
Nagy C, Maitra M, Tanti A, Suderman M, Théroux JF, Davoli MA et al (2020) Single-nucleus transcriptomics of the prefrontal cortex in major depressive disorder implicates oligodendrocyte precursor cells and excitatory neurons. Nat Neurosci 23:771–781. https://doi.org/10.1038/s41593-020-0621-y
Neely S, Williamson J, Klingseisen A, Zoupi L, Early J, Williams A, et al. (2020) New oligodendrocytes exhibit more abundant and accurate myelin regeneration than those that survive demyelination. 1–14. doi: https://doi.org/10.1101/2020.05.22.110551
Ong W, Marinval N, Lin J, Nai MH, Chong YS, Pinese C et al (2020) Biomimicking Fiber Platform with Tunable Stiffness to Study Mechanotransduction Reveals Stiffness Enhances Oligodendrocyte Differentiation but Impedes Myelination through YAP-Dependent Regulation. Small 16:1–13. https://doi.org/10.1002/smll.202003656
Osório MJ, Goldman SA (2018) Neurogenetics of Pelizaeus–Merzbacher disease, 1st ed. Elsevier B.V.
Paşca AM, Sloan SA, Clarke LE, Tian Y, Makinson CD, Huber N et al (2015) Functional cortical neurons and astrocytes from human pluripotent stem cells in 3D culture. Nat Methods 12:671–678. https://doi.org/10.1038/nmeth.3415
Penfield W (1924) Oligodendroglia and its relation to classical neuroglia. Brain 47:430–452. https://doi.org/10.1093/brain/47.4.430
Pérez-Cerdá F, Sánchez-Gómez MV, Matute C (2015) Pío del Río Hortega and the discovery of the oligodendrocytes. Front Neuroanat 9:7–12. https://doi.org/10.3389/fnana.2015.00092
Pringle NP, Richardson WD (1993) A singularity of PDGF alpha-receptor expression in the dorsoventral axis of the neural tube may define the origin of the oligodendrocyte lineage. Development 117:525–533. https://doi.org/10.1242/dev.117.2.525
Rajani RM, Quick S, Ruigrok SR, Graham D, Harris SE, Verhaaren BFJ et al (2018) Reversal of endothelial dysfunction reduces white matter vulnerability in cerebral small vessel disease in rats. Sci Transl Med. https://doi.org/10.1126/scitranslmed.aam9507
Rakic S, Zecevic N (2003) Early oligodendrocyte progenitor cells in the human fetal telencephalon. Glia 41:117–127. https://doi.org/10.1002/glia.10140
Regev A, Teichmann SA, Lander ES, Amit I, Benoist C, Birney E et al (2017) The human cell atlas Elife 6:1–30. https://doi.org/10.7554/eLife.27041
Reynolds R, Wilkin GP (1988) Development of macroglial cells in rat cerebellum II. An in situ immunohistochemical study of oligodendroglial lineage from precursor to mature myelinating cell. Development 102:409–425
Richardson WD, Smith HK, Sun T, Pringle NP, Hall A, Woodruff R (2000) Oligodendrocyte lineage and the motor neuron connection. Glia 29:136–142. https://doi.org/10.1002/(SICI)1098-1136(20000115)29:2%3c136::AID-GLIA6%3e3.0.CO;2-G
Del Rio-Hortega P (1921) La glía de escasas radiaciones (oligodendroglía). Bol Real Soc Esp Hist Nat 63–92
Del Rio-Hortega P (1928) Tercera aportación al conocimiento morfológico e interpretación funcional de la oligodendroglía. Mem Real Soc Esp Hist Nat 5–122
Rivera AD, Pieropan F, Chacon-De-La-Rocha I, Lecca D, Abbracchio MP, Azim K et al (2021) Functional genomic analyses highlight a shift in Gpr17-regulated cellular processes in oligodendrocyte progenitor cells and underlying myelin dysregulation in the aged mouse cerebrum. Aging Cell 20:1–14. https://doi.org/10.1111/acel.13335
Rivkin MJ, Flax J, Mozell R, Osathanondh R, Volpe JJ, Villa-Komaroff L (1995) Oligodendroglial development in human fetal cerebrum. Ann Neurol 38:92–101. https://doi.org/10.1002/ana.410380116
Rodriques SG, Stickels RR, Goeva A, Martin CA, Murray E, Vanderburg CR, et al. (2019) Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution. Science (80- ) 363:1463–1467. doi: https://doi.org/10.1126/science.aaw1219
Rohan Z, Milenkovic I, Lutz MI, Matej R, Kovacs GG (2016) Shared and distinct patterns of oligodendroglial response in α-synucleinopathies and tauopathies. J Neuropathol Exp Neurol 75:1100–1109. https://doi.org/10.1093/jnen/nlw087
Rowitch DH (2004) Glial specification in the vertebrate neural tube. Nat Rev Neurosci 5:409–419. https://doi.org/10.1038/nrn1389
Rowitch DH, Kriegstein AR (2010) Developmental genetics of vertebrate glial-cell specification. Nature 468:214–222. https://doi.org/10.1038/nature09611
Schirmer L, Velmeshev D, Holmqvist S, Kaufmann M, Werneburg S, Jung D et al (2019) Neuronal vulnerability and multilineage diversity in multiple sclerosis. Nature 573:75–82. https://doi.org/10.1038/s41586-019-1404-z
Schneider S, Gruart A, Grade S, Zhang Y, Kröger S, Kirchhoff F et al (2016) Decrease in newly generated oligodendrocytes leads to motor dysfunctions and changed myelin structures that can be rescued by transplanted cells. Glia 64:2201–2218. https://doi.org/10.1002/glia.23055
Scholz J, Klein MC, Behrens TEJ, Johansen-Berg H (2009) Training induces changes in white-matter architecture. Nat Neurosci 12:1370–1371. https://doi.org/10.1038/nn.2412
Schulz K, Vulpe CD, Harris LZ, David S (2011) Iron efflux from oligodendrocytes is differentially regulated in gray and white matter. J Neurosci 31:13301–13311. https://doi.org/10.1523/JNEUROSCI.2838-11.2011
Semple BD, Blomgren K, Gimlin K, Ferriero DM, Noble-Haeusslein LJ (2013) Brain development in rodents and humans: Identifying benchmarks of maturation and vulnerability to injury across species. Prog Neurobiol 106–107:1–16. https://doi.org/10.1016/j.pneurobio.2013.04.001
Shepherd MN, Pomicter AD, Velazco CS, Henderson SC, Dupree JL (2012) Paranodal reorganization results in the depletion of transverse bands in the aged central nervous system. Neurobiol Aging 33:203.e13-203.e24. https://doi.org/10.1016/j.neurobiolaging.2010.08.001
Shi Y, Wardlaw JM (2016) Update on cerebral small vessel disease: a dynamic whole-brain disease. Stroke Vasc Neurol 1:83–92. https://doi.org/10.1136/svn-2016-000035
Siebzehnrübl FA, Raber KA, Urbach YK, Schulze-Krebs A, Canneva F, Moceri S et al (2018) Early postnatal behavioral, cellular, and molecular changes in models of Huntington disease are reversible by HDAC inhibition. Proc Natl Acad Sci USA 115:E8765–E8774. https://doi.org/10.1073/pnas.1807962115
Simon C, Götz M, Dimou L (2011) Progenitors in the adult cerebral cortex: Cell cycle properties and regulation by physiological stimuli and injury. Glia 59:869–881. https://doi.org/10.1002/glia.21156
Smart IHM, Dehay C, Giroud P, Berland M, Kennedy H (2002) Unique morphological features of the proliferative zones and postmitotic compartments of the neural epithelium giving rise to striate and extrastriate cortex in the monkey. Cereb Cortex 12:37–53. https://doi.org/10.1093/cercor/12.1.37
Spitzer SO, Sitnikov S, Kamen Y, Evans KA, Kronenberg-Versteeg D, Dietmann S et al (2019) Oligodendrocyte progenitor cells become regionally diverse and heterogeneous with age. Neuron 101:459-471.e5. https://doi.org/10.1016/j.neuron.2018.12.020
Vallstedt A, Klos JM, Ericson J (2005) Multiple dorsoventral origins of oligodendrocyte generation in the spinal cord and hindbrain. Neuron 45:55–67. https://doi.org/10.1016/j.neuron.2004.12.026
Viganò F, Möbius W, Götz M, Dimou L (2013) Transplantation reveals regional differences in oligodendrocyte differentiation in the adult brain. Nat Neurosci 16:1370–1372. https://doi.org/10.1038/nn.3503
Virchow R (1856) Gesammelte A bhantllungen zur Wissenschaftlichen Medicin. Hamm, Grote
Volpe JJ, Kinney HC, Jensen FE, Rosenberg PA (2011) The developing oligodendrocyte: key cellular target in brain injury in the premature infant. Int J Dev Neurosci 29:423–440. https://doi.org/10.1016/j.ijdevneu.2011.02.012
Vukusic S, Confavreux C (2003) Prognostic factors for progression of disability in the secondary progressive phase of multiple sclerosis. J Neurol Sci 206:135–137. https://doi.org/10.1016/S0022-510X(02)00426-4
Warf BC, Fok-Seang J, Miller RH (1991) Evidence for the ventral origin of oligodendrocyte precursors in the rat spinal cord. J Neurosci 11:2477–2488. https://doi.org/10.1523/jneurosci.11-08-02477.1991
Wei H, Dong X, You Y, Hai B, Duran RC, Wu X et al (2021) OLIG2 regulates lncRNAs and its own expression during oligodendrocyte lineage formation. BMC Biol 19:132. https://doi.org/10.1186/s12915-021-01057-6
Windrem MS, Schanz SJ, Guo M, Tian G-F, Washco V, Stanwood N et al (2008) Neonatal chimerization with human glial progenitor cells can both remyelinate and rescue the otherwise lethally hypomyelinated Shiverer mouse. Cell Stem Cell 2:553–565. https://doi.org/10.1016/j.stem.2008.03.020
Xiao L, Ohayon D, Mckenzie IA, Sinclair-Wilson A, Wright JL, Fudge AD et al (2016) Rapid production of new oligodendrocytes is required in the earliest stages of motor-skill learning. Nat Neurosci 19:1210–1217. https://doi.org/10.1038/nn.4351
Xiao Y, Hoodless LJ, Petrucco L, Portugues R, Czopka T (2021) Oligodendrocyte precursor cells sculpt the visual system by regulating axonal remodeling. bioRxiv. https://doi.org/10.1101/2021.03.11.434829
Yeung MSY, Djelloul M, Steiner E, Bernard S, Salehpour M, Possnert G et al (2019) Dynamics of oligodendrocyte generation in multiple sclerosis. Nature 566:538–542. https://doi.org/10.1038/s41586-018-0842-3
Yeung MSY, Zdunek S, Bergmann O, Bernard S, Salehpour M, Alkass K et al (2014) Dynamics of oligodendrocyte generation and myelination in the human brain. Cell 159:766–774. https://doi.org/10.1016/j.cell.2014.10.011
Zecevic N, Chen Y, Filipovic R (2005) Contributions of cortical subventricular zone to the development of the human cerebral cortex. J Comp Neurol 491:109–122. https://doi.org/10.1002/cne.20714
Zeisel A, Munoz-Manchado AB, Codeluppi S, Lonnerberg P, La Manno G, Jureus A et al (2015) Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science (80-) 347:1138–1142. https://doi.org/10.1126/science.aaa1934
Zhang K, Sejnowski TJ (2000) A universal scaling law between gray matter and white matter of cerebral cortex. Proc Natl Acad Sci U S A 97:5621–5626. https://doi.org/10.1073/pnas.090504197
Zhang L, Goldman JE (1996) Developmental fates and migratory pathways of dividing progenitors in the postnatal rat cerebellum. J Comp Neurol 370:536–550. https://doi.org/10.1002/(SICI)1096-9861(19960708)370:4%3c536::AID-CNE9%3e3.0.CO;2-5
Zhang X, Li T, Liu F, Chen Y, Yao J, Li Z et al (2019) Comparative analysis of droplet-based ultra-high-throughput single-cell RNA-Seq systems. Mol Cell 73:130-142.e5. https://doi.org/10.1016/j.molcel.2018.10.020
Zhu Q, Whittemore SR, Devries WH, Zhao X, Kuypers NJ, Qiu M (2011) Dorsally-derived oligodendrocytes in the spinal cord contribute to axonal myelination during development and remyelination following focal demyelination. Glia 59:1612–1621. https://doi.org/10.1002/glia.21203
Acknowledgements
Our funding sources include the Multiple Sclerosis Society UK, Medical Research Council Human Cell Atlas, and UK Dementia Research Institute as UK DRI which was funded by the MRC, Alzheimer’s Society and Alzheimer’s Research UK. We would also like to acknowledge that Fig. 4c was kindly provided by Dr. Sarah Jäkel.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Seeker, L.A., Williams, A. Oligodendroglia heterogeneity in the human central nervous system. Acta Neuropathol 143, 143–157 (2022). https://doi.org/10.1007/s00401-021-02390-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-021-02390-4