Abstract
Aims/hypothesis
Lipotoxicity constitutes the major driving force for type 2 diabetes. Circular RNAs (circRNAs) play important roles in regulating beta cell function and exosomes are essential mediators of intercellular communication. The role of exosomal circRNAs in type 2 diabetes remains largely unknown. We aimed to examine whether lipotoxicity induces dysregulation of circRNAs in beta cell-derived exosomes and to determine the contribution of exosomal circRNAs to the development of type 2 diabetes.
Methods
Exosomes were extracted from MIN6 cells treated with palmitate or BSA, and RNA sequencing was performed. CircGlis3 (Gli-similar 3) expression level was validated by qPCR. The impact of circGlis3 on beta cell function and the deleterious effects of exosomal circGlis3 on islet endothelial cells (islet ECs) were investigated in vitro and in vivo in human and mouse models by gain or loss of function assays. The molecular mechanism of circGlis3 was explored by RNA pull-down and immunoprecipitation assays.
Results
Beta cell-derived exosomal circGlis3 was significantly upregulated under lipotoxic conditions, and exosomal circGlis3 levels were also elevated in the serum of mouse models of diabetes and participants with type 2 diabetes. CircGlis3 participated in lipotoxicity-induced beta cell dysfunction in vitro and in vivo. Moreover, beta cell-derived exosomal circGlis3 could be transferred to islet ECs and reduce the cell viability, cell migration and angiogenesis of islet ECs. Mechanistically, circGlis3 promoted the degradation of glucocorticoid modulatory element-binding protein 1 (GMEB1) by facilitating the interaction between GMEB1 and mindbomb E3 ubiquitin protein ligase 2 (MIB2), thus suppressing the phosphorylation of heat shock protein 27 (HSP27).
Conclusions/interpretation
Our study points to the involvement of circGlis3 in diabetes development, and exosomal circGlis3 transfer as a communication mode between beta cells and islet ECs, suggesting that circGlis3 might be a potential biomarker and therapeutic target for type 2 diabetes.
Data availability
The RNA-sequencing data have been deposited in the NCBI Sequence Read Archive (SRA) database, with accession number PRJNA689673. Mass spectrometry data are available via ProteomeXchange with identifier PXD024693.
Graphical abstract

Similar content being viewed by others

Introduction
Type 2 diabetes is caused by the failure of beta cells to cope with systemic insulin resistance elicited by genetic and environmental factors [1]. The increasing rise in type 2 diabetes prevalence is closely related to obesity epidemics, as obesity plays an important role in insulin resistance [2, 3]. Elevated circulating NEFA, known as lipotoxicity, constitutes the major driving force for insulin resistance and beta cell apoptosis [4, 5]. Additionally, communication between beta cells and intra-islet microvasculature is critical for the maintenance of glucose homeostasis. Islet endothelial cells (islet ECs) produce factors that govern proliferation and insulin secretion of beta cells, and factors associated with diabetes, including palmitate (PA) and glucose, induce damage of islet ECs, which may contribute to beta cell loss [6]. Islet ECs are also important for the rapid and adequate revascularisation of islet grafts [7]. Meanwhile, beta cells promote intra-islet blood flow and the survival and migration of islet ECs [7, 8]. Therefore, exploring the crosstalk between beta cells and islet ECs under lipotoxic conditions is beneficial for understanding the pathogenesis of type 2 diabetes and improving islet revascularisation after transplantation.
Studies have demonstrated the involvement of circular RNAs (circRNAs) in beta cell failure and diabetes complications [9, 10]. CircRNAs, a class of endogenous non-coding RNAs produced by back-splicing, are resistant to RNA exonucleases [11]. CircRNAs participate in diverse pathophysiological processes and act as robust potential biomarkers due to their stability and evolutionary conservation [12, 13]. Moreover, circRNAs are enriched and stable in exosomes, which are small extracellular vesicles of 50–150 nm in diameter and secreted by most cells [14, 15]. Exosomes circulate in bodily fluids and harbour multiple bioactive molecules including RNAs and proteins, which can be transferred to recipient cells and mediate the crosstalk between cells. These features enable exosomes to be utilised for clinical diagnostics and therapeutic development [15, 16]. Increasing evidence suggests that beta cells could transduce signals by secreting exosomes, paving the way for seeking novel therapeutic targets for type 2 diabetes. For example, transplantation of beta cell-derived exosomes improved glucose metabolism and promoted islet angiogenesis in a streptozotocin-induced mouse model of diabetes [17]. In addition, beta cell-derived exosomal microRNAs participated in the development of type 2 diabetes by regulating peripheral insulin sensitivity and systemic inflammation [18, 19]. Notably, exosomal circRNAs have emerged as an additional mechanism for intercellular communication [20, 21]. However, the role of exosomal circRNAs in type 2 diabetes, especially in the crosstalk between beta cells and islet ECs, remains largely unknown.
In this study, we analysed the expression profile of circRNAs in beta cell-derived exosomes and examined the role of exosomal circRNAs in the crosstalk between beta cells and islet ECs and the development of type 2 diabetes.
Methods
For detailed methods, please refer to the electronic supplementary material (ESM).
Ethics statement
All animal studies were approved by the Institutional Animal Care and Use Committee of The First Affiliated Hospital of Sun Yat-sen University. All human studies were performed according to the principles of the Declaration of Helsinki and approved by the Ethics Committee of The First Affiliated Hospital of Sun Yat-sen University. Written informed consent was obtained from all participants.
Clinical specimens
After providing written informed consent, samples of fasting venous blood were obtained from 11 healthy individuals and 11 patients aged between 18 and 60 with type 2 diabetes. Human serum was collected and then serum exosomes were isolated. See ESM Methods for details.
Animal studies
Five-week-old male C57BL/6 J mice (Model Animal Research Centre of Nanjing University, China) underwent 24-week feeding with a normal diet (ND) (D12450B; Research Diets, USA) or a high-fat diet (HFD) (60% fat [wt/wt], D12492; Research Diets). To specifically knockdown circGlis3 (Gli-similar 3) levels in beta cells, HFD-fed mice were injected intraductally with recombinant adeno‐associated virus serotype 8 vectors carrying shRNA (AAV8-shcircGlis3) or negative control (AAV8-NC) under a rat insulin promoter. For in vivo exosome studies, exosomes were extracted from the medium of MIN6 cells overexpressing circGlis3 (circGlis3-Exo) or empty vector (Vec-Exo). Six-week-old male C57BL/6 mice were randomly assigned to receive an ND diet or HFD diet, and mice were injected with PBS, Vec-Exo or circGlis3-Exo once a week for 4 weeks. See ESM Methods for further details.
Islet isolation
Islets were isolated from 10-week-old male db/db mice (BKS.Cg-Dock7m+/+Leprdb/Nju) and age-matched male littermate db/m mice as well as the C57BL/6 J mice in the above-mentioned animal studies. See ESM Methods for details.
CircRNA sequencing and identification
Total RNA was isolated using TRIzol reagent (Invitrogen, USA) from exosomes derived from PA-treated or BSA-treated MIN6 cells. Ribosomal RNA was removed by treating RNA with a RiboMinus Eukaryote Kit (Qiagen, Valencia, CA, USA) and linear RNA was digested by RNase R, followed by the construction of cDNA libraries. The libraries were quality controlled with an Agilent 2200 system and sequenced with an Illumina HiSeq 3000 (Illumina, CA, USA). The clean reads were then aligned to the mouse genome (GRCm38) using HISAT2 (https://daehwankimlab.github.io/hisat2/, v2.1.0), and the unmapped reads were collected for circRNA identification. 20mers from both ends of the unmapped reads were extracted and aligned to the reference genome to find unique anchor positions within the splice site. Anchor reads that aligned in the reversed orientation (head-to-tail) indicated circRNA splicing and then were subjected to circRNA identification tools CircRNA Identifier (CIRI, https://sourceforge.net/projects/ciri/, v1.2) [22], CIRI2 (https://sourceforge.net/projects/ciri/files/CIRI2/, v2.0.6) and find_circ (https://github.com/marvin-jens/find_circ, v1.2) to identify circRNAs [23, 24]. The anchor alignments were extended such that the complete read aligns and the breakpoints were flanked by GU/AG splice sites. A candidate circRNA was called if it was supported by at least two unique back-spliced reads in at least one sample. The identified circRNAs were subjected to statistical analysis of type and length distribution. The differentially expressed circRNAs were filtered by the following criteria: |log2 fold change (PA/BSA)|> 1 and p value < 0.05. CircRNA sequencing was carried out at the Forevergen laboratory (Guangzhou, China). The RNA-sequencing data have been deposited in the NCBI Sequence Read Archive (SRA) database, with accession number PRJNA689673.
Quantitative real-time PCR and droplet digital PCR
Total RNA of MIN6 cells, islet microvascular endothelial cells (MS-1 cells), or exosomes was extracted. Quantitative real-time PCR (qPCR) analysis was used to determine the relative expression levels of circRNAs or mRNAs, and droplet digital PCR (ddPCR) was performed to analyse the levels of MIN6 cell-derived exosomal circRNAs. The primers used in this study are listed in ESM Table 1. See ESM Methods for details.
Western blot
Total protein was extracted from MIN6 cells, MS-1 cells or exosomes. Western blotting was performed as described previously [25]. The antibodies used here are listed in ESM Table 2.
Cell transfection and lentiviral vector transduction
CircGlis3 overexpression plasmid or siRNAs were transfected into MIN6 cells or MS-1 cells with Lipofectamine 3000 (Invitrogen, USA) following the manufacturer’s instructions. For lentiviral vector transduction, pLKO.1-based shRNA plasmid or pLO5-ciR-circGlis3 plasmid was cotransfected with packaging plasmids psPAX2 and pMD2G (Addgene, USA) into HEK293T cells. Infectious lentiviruses were collected and added into the plate to infect MIN6 cells or MS-1 cells. See ESM Methods for details.
RNA-binding protein immunoprecipitation assay
RNA-binding protein immunoprecipitation (RIP) assay in MS-1 cells was carried out by Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore, USA) as previously described [25]. See ESM Methods for details.
Cell viability analysis
Cell viability of MS-1 cells or MIN6 cells was determined by Cell Counting Kit-8 (CCK-8) (Dojindo, Japan) following the manufacturer’s protocol. See ESM Methods for details.
EdU assay
Cell proliferation of MS-1 cells or MIN6 cells was assessed using the Cell Light EdU DNA imaging kit (RiboBio, China) according to the manufacturer’s instructions. See ESM Methods for details.
TUNEL staining assay
TUNEL staining was performed in MS-1 cells using the One Step TUNEL apoptosis assay kit (Beyotime, China) according to the manufacturer’s instructions.
Flow cytometric analysis
Cell apoptosis of MS-1 cells or MIN6 cells was analysed by flow cytometry (BD FACSCalibur, USA) with Annexin V/propidium iodide staining. See ESM Methods for details.
Wound healing and cell migration assay
For the wound healing assay, MS-1 cells were seeded in 6-well plates and a scratch was created using a 200 μl pipette tip. Scratch wounds were photographed in the same position at 0 h and 48 h, respectively. For the cell migration assay, MS-1 cells were seeded onto the upper insert of the chamber, and allowed to migrate for 48 h. Inserts were stained with 0.1% crystal violet and migrated cells were photographed. See ESM Methods for details.
Tube formation assay
Matrigel (Corning, USA) was added to 96-well plates and incubated for 30 min at 37℃. Then, MS-1 cells were seeded and the tube structures were photographed at 6 h. See ESM Methods for details.
Immunofluorescence
Immunofluorescence microscopy was performed on frozen sections of mouse pancreas and MS-1 cells with indicated primary antibodies. See ESM Methods for details.
Immunohistochemistry
Immunohistochemistry was performed on paraffin sections of mouse pancreas using antibodies against Ki67, Insulin or CD31. See ESM Methods for details.
Co-immunoprecipitation assay
MS-1 cells were lysed and a co-immunoprecipitation assay was performed using antibodies against glucocorticoid modulatory element-binding protein 1 (GMEB1) or HA. See ESM Methods for details.
RNase R treatment
Total RNA from MIN6 cells or MS-1 cells was treated with RNase R (Geneseed Biotech, China) to evaluate the stability of circGlis3. See ESM Methods for details.
Inhibitors
MIN6 cells were treated with GW4869 (Sigma-Aldrich, USA) to inhibit exosome release, and MS-1 cells were treated with MG132 (Sigma-Aldrich) to inhibit the proteolytic activity of the proteasome complex. See ESM Methods for details.
Glucose-stimulated insulin secretion assay
The insulin secretion in MIN6 cells was determined by glucose-stimulated insulin secretion (GSIS) assay. See ESM Methods for details.
RNA fluorescence in situ hybridisation
Fluorescence in situ hybridisation (FISH) assay was performed with Cy3-labelled circGlis3 probe to detect the location of circGlis3 in MIN6 cells. See ESM Methods for details.
Prediction of circGlis3-binding proteins
The software Find Individual Motif Occurrences (FIMO, https://meme-suite.org/meme/tools/fimo, v5.4.1) was applied to predict circGlis3-binding proteins according to the instructions [26].
RNA pull-down and mass spectrometry analysis
Pierce Magnetic RNA–Protein Pull-Down Kit (Thermo Fisher Scientific, USA) was applied to perform RNA pull-down assay as described in a previous study [27]. RNA-interacting proteins were detected by mass spectrometry (Fitgene Biotech, China) or western blot. Mass spectrometry data are available via ProteomeXchange with identifier PXD024693. The details of the RNA pull-down assay are shown in the ESM Methods.
Statistical analysis
All data are presented as mean ± SD from at least three independent experiments. Two-tailed Student’s t test (unpaired) for two groups and ANOVA (followed by Tukey’s post hoc test) for multiple groups were used to analyse the significance of the data. All statistical analyses were performed using SPSS v.22.0 (SPSS, USA) or GraphPad Prism 7.0 software. A p value < 0.05 was considered statistically significant. Human and mouse serum exosomal circGlis3 quantification and histological analyses were blinded and performed by a technical assistant. Otherwise, blinding was not carried out. Data from mice that died in accidents were excluded.
Results
Beta cell-derived exosomal circGlis3 is upregulated under lipotoxic conditions
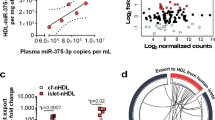
Exosomes were isolated from MIN6 cells incubated with PA or BSA (PA-Exo and BSA-Exo), and identified by transmission electron microscopy (TEM), NanoSight and western blot (Fig. 1a-c). To explore lipotoxicity-induced changes in expression of beta cell-derived exosomal circRNAs, we performed RNA-seq analysis. A total of 1102 circRNAs were detected by CIRI, 49.0% of which consisted of protein coding exons (ESM Fig. 1a). The length of most circRNAs was less than 1000 nucleotides (ESM Fig. 1b). We also used find_circ and CIRI2 to predict circRNAs. A total of 1155 (find_circ) and 416 (CIRI2) circRNAs were identified. Of these, 910 and 31 circRNAs were newly identified by find_circ and CIRI2, respectively. An overlap of 198 circRNAs was observed between all three algorithms (ESM Fig. 1c, ESM Table 3). Among 293 differentially expressed circRNAs obtained by CIRI, 121 were upregulated and 172 were downregulated in PA-Exo (|log2(fold change) |> 1; p value < 0.05) and 23 circRNAs were found to be conserved with human circRNAs (ESM Fig. 1d, ESM Table 4, ESM Table 5).The heatmap displays 20 significantly differentially expressed circRNAs that have been annotated in circBase [28] (Fig. 1d, ESM Table 6). Given the high conservation between humans and mice, four upregulated circRNAs were chosen to verify the RNA-seq results (ESM Fig. 1e, f and Fig. 1e). CircRNA-Glis3 (mmu_circ_0000943, named as circGlis3) was selected for further analysis due to its abundance and significant upregulation in PA-treated MIN6 cells and their exosomes (Fig. 1e, ESM Fig. 1g). Besides, lipotoxicity elevated circGlis3 levels in islets and exosomes (Fig. 1f, ESM Fig. 1h). We further detected circGlis3 levels in serum exosomes. As expected, serum exosomal circGlis3 was more abundant in an HFD-induced mouse model of diabetes and in db/db mice, compared with control mice (Fig. 1g, ESM Fig. 1i). Moreover, human circGLIS3 (hsa_circ_0006370, conserved with mmu_circ_0000943) levels in serum exosomes were higher in participants with type 2 diabetes than in healthy control participants (Fig. 1h). These results suggest that beta cell-derived exosomal circGlis3 was increased and could be delivered by exosomes into the circulation under lipotoxic conditions, making it a potential biomarker of type 2 diabetes.
Beta cell-derived exosomal circGlis3 is upregulated under lipotoxic conditions. Exosomes were extracted from the supernatants of MIN6 cells incubated with 0.3 mmol/l PA (PA-Exo) or 0.3% BSA (BSA-Exo) for 24 h, and RNA-seq analysis was performed. (a) Representative TEM pictures of exosomes derived from MIN6 cells. Scale bar, 200 nm. (b) The size distribution of exosomes was detected by nanoparticle tracking (NAT) analysis. (c) Western blot analysis of exosome-specific markers CD9 and TSG101, and the endoplasmic reticulum marker calnexin. (d) Clustered heatmap of the differentially expressed circRNAs in BSA-Exo and PA-Exo. The heatmap was plotted using log10-transformed normalised back-spliced read counts. (e) Absolute quantification of circGlis3 in BSA-Exo and PA-Exo in MIN6 cells by ddPCR. (f) qPCR detected the expression level of exosomal circGlis3 derived from islets treated with 0.3 mmol/l PA or 0.3% BSA for 24 h. (g) qPCR analysis of circGlis3 levels in serum exosomes isolated from mice fed an ND or HFD for 28 weeks (n = 5/group). (h) CircGLIS3 levels in serum exosomes obtained from participants with type 2 diabetes (T2D) and healthy control participants were determined by qPCR (n = 11/group). Data are represented as mean ± SD of 3 independent experiments. *p < 0.05, **p < 0.01 and ***p < 0.001, by two-tailed Student’s t test. TSG101, tumour susceptibility gene 101
Characterisation of circGlis3 in beta cells
CircGlis3 is back-spliced from exon 4 of the Glis3 gene with a length of 1114 bp, and the splice junction was verified by Sanger sequencing (Fig. 2a). The genomic regions of Glis3 exon 4 contain flanking Alu repeats and long introns, which are highly reverse-complementary (ESM Fig. 2a, b). In addition, circGlis3 was extensively expressed in various mouse tissues, particularly enriched in kidney and islets (ESM Fig. 2c). Moreover, circGlis3 could only be amplified using divergent primers in cDNA (Fig. 2b). In addition, circGlis3 was resistant to RNase R digestion and was much more stable than Glis3 linear mRNA (Fig. 2c,d). As revealed by reverse transcription experiments, circGlis3 levels were markedly decreased when using the oligo (dT) primers, compared with random hexamer primers (Fig. 2e). qPCR and FISH assays suggest that circGlis3 was mainly located in the cytoplasm (Fig. 2f,g). Taken together, these data indicate that circGlis3 is a circular and stable transcript.
Characterisation of circGlis3 in beta cells. (a) Scheme illustrating the formation of circGlis3. The splice junction was Sanger sequenced using cDNA samples from PCR. (b) PCR analysis of the existence of circGlis3 with divergent primers and convergent primers in cDNA and gDNA. The inward and outward facing arrowheads indicate convergent and divergent primers, respectively. GAPDH was used as a control for a linear RNA transcript. (c) qPCR analysis of circGlis3 and Glis3 linear mRNA with or without RNase R treatment. Data were normalised to the mock group. (d) Relative RNA levels of circGlis3 and Glis3 mRNA were analysed by qPCR in MIN6 cells treated with actinomycin D at the indicated time points. (e) CircGlis3 expression level after reverse transcription with random or oligo-dT primers using total RNAs from MIN6 cells. (f) The level of circGlis3 in cytosolic and nucleoplasmic fractions of MIN6 cells was assessed by qPCR. β-Actin and U6 were used as cytosol marker and nucleus marker respectively. (g) RNA-FISH for circGlis3 in MIN6 cells. The circGlis3 probe was labelled with Cy3 (red) and nuclei were stained with DAPI (blue). Scale bar, 20 μm. Results are expressed as mean ± SD of 3 independent experiments. **p < 0.01 and ***p < 0.001, by two-tailed Student’s t test. gDNA, genomic DNA
CircGlis3 regulates beta cell function in vitro and in vivo
Next, we investigated the biological functions of circGlis3 in beta cells. Overexpression of circGlis3 reduced cell viability and proliferation, increased cell apoptosis, and impaired insulin secretion (Fig. 3a-f). Additionally, inhibiting circGlis3 did not affect cell viability under basal conditions, but alleviated the decreased cell viability induced by PA (ESM Fig. 3a, b). Then, we explored the role of circGlis3 in vivo. First, we established HFD-induced diabetic mouse models. After 24 weeks of diet intervention, HFD-fed mice displayed elevated fasting blood glucose (FBG) levels and body weight, as well as impaired glucose tolerance and insulin sensitivity (Fig. 3g, ESM Fig. 3c-e), suggesting that the mice had developed insulin resistance and early type 2 diabetes. Second, we constructed AAV8-shcircGlis3, which encoded GFP and was driven by the rat insulin promoter to specifically silence circGlis3 in beta cells. Four weeks after injection, GFP expression was restricted to beta cells and not detected in alpha cells (Fig. 3h, ESM Fig. 3f). CircGlis3, predominantly expressed in the islets, was upregulated in the islets of HFD-induced diabetic mice and was silenced by AAV8-shcircGlis3 (Fig. 3i, ESM Fig. 3g). Similarly, circGlis3 levels were elevated in the islets of db/db mice compared with wild-type mice (ESM Fig. 3h). Furthermore, although there was no statistical difference, AAV8-shcircGlis3 apparently reduced the levels of serum exosomal circGlis3 in HFD-fed mice (p = 0.0735), suggesting that a considerable portion of circulating exosomal circGlis3 came from the pancreatic islets (ESM Fig. 3i). Inhibiting circGlis3 in diabetic mice not only mitigated hyperglycaemia and hyperinsulinaemia, but also improved glucose tolerance and insulin sensitivity (Fig. 3g,j and ESM Fig. 3j, k). Nevertheless, the elevated serum total cholesterol and LDL-cholesterol levels in diabetic mice did not markedly decline after silencing circGlis3 (ESM Fig. 3l). Furthermore, knockdown of circGlis3 ameliorated the decreased beta cell proliferation and increased beta cell apoptosis in diabetic mice (Fig. 3k,l). Collectively, these findings demonstrated that circGlis3 contributed to beta cell dysfunction in vitro and in vivo.
CircGlis3 regulates beta cell function in vitro and in vivo. (a–f) MIN6 cells were stably transfected with circGlis3 overexpression plasmid (circGlis3) or empty vector (Vector). qPCR analysed the levels of circGlis3 and Glis3 mRNA (a). Cell viability of MIN6 cells was measured by CCK-8 assay (b). Cell proliferation was detected via EdU staining. Scale bar, 100 μm (c). Cell apoptosis was evaluated by flow cytometry (d). Western blot detected the expression levels of proteins (PCNA, Bcl-2, cleaved caspase-3). β-Tubulin served as loading control, and the numbers between each row in the western blots are the quantification levels, relative to the Vector group set to 1.00 (e). Insulin secretion was analysed by GSIS assay (f). (g–l) Mice were fed an ND or an HFD for 24 weeks, then HFD-fed mice were injected intraductally with AAV8-NC or AAV8-shcircGlis3. Both expression cassettes encoded the GFP reporter gene and were driven by rat insulin promotor. Animals were analysed 4 weeks after injection. FBG levels were monitored every 2 weeks (g). Immunofluorescence analysis of insulin and GFP expression in islets. Scale bar, 100 μm (h). The level of circGlis3 in islets was measured by qPCR (n = 6/group) (i). IPGTTs were conducted 4 weeks after AAV-8 injection and the AUC was calculated (n = 8/group) (j). Immunohistochemistry analysis of insulin and Ki67 in mouse pancreatic sections. Arrows indicate Ki67-positive beta cells. Scale bar, 50 μm (k). Beta cell apoptosis was assessed by TUNEL (red) and insulin/GFP (green) coimmunostaining. Yellow arrows indicate the DAPI/TUNEL/insulin or DAPI/TUNEL/GFP copositive beta cells. Scale bar, 100 μm (l). Data are represented as mean ± SD of 3 independent experiments. *p < 0.05, **p < 0.01 and ***p < 0.001. †p < 0.001 HFD group vs ND group. ‡p < 0.001 HFD + AAV8-shcircGlis3 group vs HFD + AAV8-NC group. Two-tailed Student’s t tests were applied in (a–d, f, g). One-way ANOVA (with Tukey post hoc test) was used for analysing the difference in (i–l). PI, propidium iodide; Bcl-2, B cell lymphoma 2; GSIS, glucose-stimulated insulin secretion; PCNA, proliferating cell nuclear antigen
Beta cell-derived exosomal circGlis3 promotes islet EC dysfunction
Interestingly, islet-derived exosomal circGlis3 levels were upregulated in HFD-induced diabetic mice, but downregulated by AAV8-shcircGlis3 (Fig. 4a), implying that circGlis3 may be delivered by exosomes as a form of cell-to-cell communication. Moreover, islet capillary density was reduced in diabetic mice, whereas it was improved by inhibiting circGlis3 in beta cells (Fig. 4b). On the basis of structurally and functionally unique capillary network in pancreatic islets [8], we speculated that beta cell-derived exosomal circGlis3 may act as a paracrine regulator of islet EC function. We first investigated the effect of lipotoxic beta cell-derived exosomes on islet ECs. PKH67-labelled exosomes from MIN6 cells were internalised by MS-1 cells (ESM Fig. 4a). Compared with PBS or BSA-Exo, PA-Exo reduced MS-1 cell viability (ESM Fig. 4b). Of note, the supernatants recovered after ultracentrifugation of the exosome preparation did not affect cell viability (ESM Fig. 4c), suggesting that the deleterious effect was not mediated by PA or other soluble factors. Besides, PA-Exo inhibited nitric oxide (NO) production, cell migration and angiogenesis, and promoted apoptosis of MS-1 cells (ESM Fig. 4d-g). Moreover, PA-Exo increased circGlis3 levels in MS-1 cells and prior addition of the exosome secretion inhibitor GW4869 to MIN6 cells blocked the delivery of circGlis3 from MIN6 cells into MS-1 cells (Fig. 4c), indicating that MIN6 cells secreted extracellular circGlis3 predominantly in an exosome-dependent manner. Lipotoxicity failed to alter circGlis3 levels in MS-1 cells (ESM Fig. 4h). The elevation of circGlis3 in recipient cells was not affected by actinomycin D, excluding the involvement of endogenous induction (Fig. 4d). Additionally, we overexpressed circGlis3 in MIN6 cells and exosomes (Fig. 4e). CircGlis3-Exo elevated circGlis3 levels in MS-1 cells and reduced cell viability and proliferation, while increasing cell apoptosis (Fig. 4f-j). CircGlis3-Exo also inhibited cell migration and tube formation, without significant effect on NO production (Fig. 4k-m, ESM Fig. 4i). We further verified the function of circGlis3 by modulating circGlis3 levels in MS-1 cells (Fig. 5a, ESM Fig. 5a). As expected, overexpression of circGlis3 induced dysfunction of MS-1 cells (Fig. 5b-h). Moreover, circGlis3 knockdown had little impact on cell proliferation, cell migration and tube formation, but reduced lipotoxicity-induced cell apoptosis (ESM Fig. 5b-f). Apoptosis markers were upregulated by overexpressing circGlis3, and downregulated by silencing circGlis3 (Fig. 5i). Neither NO production nor p-eNOS protein levels were affected by altering circGlis3 levels (ESM Fig. 5g, h). Of note, silencing circGlis3 in MS-1 cells rescued the upregulated apoptosis markers and antagonised the decreased cell viability induced by PA-Exo (Fig. 5j, ESM Fig. 5i). Overall, these data showed that beta cell-derived exosomal circGlis3 promoted islet EC dysfunction.
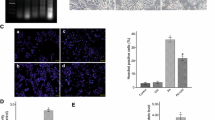
Beta cell-derived exosomal circGlis3 promotes islet EC dysfunction. (a, b) Animals (ND, HFD, HFD + AAV8-NC, HFD + AAV8-shcircGlis3) were treated as described in Fig. 3. Islets were isolated and cultured for 48 h, then exosomes were extracted from the supernatants and the level of exosomal circGlis3 was detected by qPCR (a). Immunohistochemistry was used to analyse the islet capillary density after intervention. Scale bar, 50 μm. Dashed lines indicate the area of islets (b). (c) MIN6 cells were pre-treated with GW4869 (10 μmol/l) or not, and exosomes (BSA-Exo or PA-Exo) were extracted. CircGlis3 levels in MS-1 cells incubated with PBS or exosomes were determined by qPCR. (d) MS-1 cells were treated with actinomycin D (1 μg/ml), followed by PBS or the indicated exosome treatment. The expression level of circGlis3 in MS-1 cells was detected. (e) MIN6 cells were stably transfected with circGlis3-overexpressing plasmid (ov-circGlis3) or empty vector (Control). qPCR confirmed the overexpression of circGlis3 in MIN6 cells and exosomes. (f–m) Exosomes overexpressing circGlis3 (circGlis3-Exo) or vector control (Vec-Exo) were isolated, and MS-1 cells were cocultured with PBS or indicated exosomes for 48 h. The level of circGlis3 in MS-1 cells was determined by qPCR (f). Cell viability of MS-1 cells was measured by CCK-8 assay (g). Cell proliferation was evaluated via EdU staining and data were analysed by Image J. Scale bar, 100 μm (h). Cell apoptosis was detected by flow cytometry (i). Proteins (PCNA, Bax, and cleaved caspase-3) were analysed by western blot. The numbers between each row in the western blots are the quantification levels, relative to the PBS group set to 1.00 (j). Migration capacity of MS-1 cells was evaluated by scratch wound healing assays. Dashed lines indicate the wound edges of the scratch. Scale bar, 200 μm (k). Transwell assays (l) and tube formation assays (m) were conducted. Scale bar, 100 μm. Data are expressed as mean ± SD of 3 independent experiments. *p < 0.05, **p < 0.01 and ***p < 0.001, NS indicates no statistical significance, by two-tailed Student’s t test in (e), and one-way ANOVA (with Tukey post hoc test) in (a–d, f–i, k–m). Bax, Bcl-2-associated X protein; PCNA, proliferating cell nuclear antigen; PI, propidium iodide
CircGlis3 damages the function of islet ECs in vitro. (a–h) MS-1 cells were transfected with circGlis3 overexpression plasmid or empty vector for 48 h. qPCR detected the overexpression of circGlis3 in MS-1 cells with or without RNase R treatment (a). Cell viability was measured by CCK-8 assay (b). Cell proliferation was evaluated via EdU staining and data were analysed by Image J. Scale bar, 100 μm (c). Cell apoptosis was analysed by flow cytometry (d). Migration capacity of MS-1 cells was evaluated by scratch wound healing assays. Dashed lines indicate the wound edges of the scratch. Scale bar, 200 μm (e). Transwell assays (f) and tube formation assays (g) were conducted. Scale bar, 100 μm. Western blot analysis of proliferation markers (PCNA, Cyclin D1) after overexpressing circGlis3 (h). (i) circGlis3 levels were overexpressed or silenced by transfecting circGlis3 overexpression plasmid or siRNAs targeting circGlis3 (siRNA-1 and siRNA-2) into MS-1 cells. After 24 h of siRNA transfection, MS-1 cells were treated with or without 0.3 mmol/l PA for another 24 h. Apoptosis markers (Bax and cleaved caspase-3) were detected by western blot. (j) After 24 h of siRNA transfection, MS-1 cells were incubated with exosomes derived from MIN6 cells (BSA-Exo or PA-Exo) for 48 h. Proteins (Bax and cleaved caspase-3) were measured by western blot. The numbers between each row in the western blots are the quantification levels, relative to the left-hand lane set to 1.00 (h–j). Values are represented as mean ± SD of 3 independent experiments. *p < 0.05, **p < 0.01 and ***p < 0.001, by two-tailed Student’s t test in (a–g). PI, propidium iodide; Bax, Bcl-2-associated X protein; PCNA, proliferating cell nuclear antigen
CircGlis3 directly interacts with GMEB1 and promotes its degradation in MS-1 cells
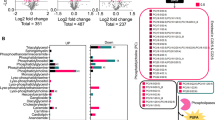
Given that circGlis3 was mainly located in the cytoplasm of MS-1 cells (ESM Fig. 6a), we first examined whether circGlis3 could bind to microRNAs. However, circGlis3 did not bind to argonaute 2 (AGO2), a component of the RNA-induced silencing complex (ESM Fig. 6b). Increasingly, studies have revealed the binding capability of circRNAs on proteins [27, 29]. Therefore, RNA pull-down assays were performed to identify potential circGlis3-binding proteins. The circular and linear RNA probes were validated by RT-PCR (ESM Fig. 6c). Mass spectrometry assay identified 29 differential proteins between circGlis3 and linear transcript pull-down groups (ESM Table 7). These proteins were overlapped with circGlis3-binding proteins predicted by FIMO software (ESM Table 8), and two proteins (GMEB1 and hepatocyte nuclear factor 3β [HNF3β]) were left (Fig. 6a, ESM Fig. 6d). GMEB1, a member of the family of KDWK proteins that inhibits cell apoptosis by binding to proteins [30], was present in the circGlis3 probe pull-down samples, while HNF3β was not detected by western blot (Fig. 6b). Meanwhile, RIP assays showed that circGlis3 directly interacted with GMEB1, but not HNF3β (Fig. 6c, ESM Fig. 6e). The colocalisation of circGlis3 and GMEB1 in MS-1 cells was further verified (Fig. 6d). Moreover, GMEB1 protein levels were reduced by overexpressing circGlis3 and increased by silencing circGlis3, while modifying circGlis3 levels had no effect on Gmeb1 mRNA levels (Fig. 6e, ESM Fig. 6f, g). Reduced GMEB1 protein levels were observed in MS-1 cells with circGlis3 overexpression, and a more significant decrease was detected in the cytoplasm (Fig. 6f,g).
CircGlis3 interacts with GMEB1 and promotes its degradation in MS-1 cells. (a) Venn diagram shows the proteins pulled down by biotin-labelled circGlis3 in MS-1 cells and their overlapping analysis with predicted circGlis3-binding proteins (RBPs). (b) Western blot validated the proteins pulled down by circular (C-Glis3) or linear (L-Glis3) RNA probe from the lysates of MS-1 cells. (c) RIP assay was performed using anti-GMEB1 antibody, followed by qPCR and RT-PCR to detect circGlis3 enrichment. (d) Dual RNA-FISH and immunofluorescence showed the colocalisation of circGlis3 (red) and GMEB1 (green) in MS-1 cells. Scale bar, 10 μm. (e) Western blot detected the protein levels of GMEB1 after overexpressing or silencing circGlis3 in MS-1 cells. (f–i) MS-1 cells were stably transfected with empty vector (Vec-control) or circGlis3 overexpression plasmid (ov-circGlis3). FISH and immunofluorescence were used to analyse the protein levels of GMEB1 after overexpressing circGlis3 in MS-1 cells. Scale bar, 20 μm (f). Western blot detected subcellular levels of GMEB1 in nuclear and cytoplasmic protein extractions. GAPDH and histone H3 were used as cytoplasmic and nuclear protein control, respectively (g). MS-1 cells were treated with the protein synthesis inhibitor cycloheximide (CHX, 50 μg/ml) for the indicated time. GMEB1 protein levels were analysed by western blot (h), and the speed of protein degradation was calculated (i). (j) MS-1 cells with circGlis3 overexpression were treated with MG132 (10 μmol/l) for 12 h and GMEB1 protein levels were detected. (k) Cell lysates from stable circGlis3-overexpressing (ov-circGlis3) or stable circGlis3-knockdown (sh-circGlis3) MS-1 cells treated with MG132 for 12 h were immunoprecipitated with GMEB1 antibody or IgG control, and then immunoblotted with ubiquitin and GMEB1 antibodies. (l) Western blot validated the MIB2 protein pulled down by circular (C-Glis3) RNA probe from the lysates of MS-1 cells. (m–o) HA-tagged Mib2 plasmid was transfected into MS-1 cells. RIP assay was performed using anti-HA antibody, followed by qPCR and RT-PCR to detect circGlis3 enrichment (m). Coimmunoprecipitation was performed using anti-GMEB1 antibody (n) or anti-HA antibody (o) in MS-1 cells. (p) Immunoprecipitation analysed the association between GMEB1 and MIB2 in MG132-treated MS-1 cells bearing Vec-control or ov-circGlis3. The numbers between each row in the western blots were the quantification levels, relative to the left-hand lane set to 1.00 (e, g, h, j). Data are represented as mean ± SD of 3 independent experiments. *p < 0.05, **p < 0.01 and ***p < 0.001, by two-tailed Student’s t test. HA, haemagglutinin
Then, we determined whether circGlis3 affected the degradation of GMEB1. As expected, overexpressing circGlis3 significantly reduced GMEB1 protein stability, which could be abolished by the proteasome inhibitor MG132 (Fig. 6h-j). The ubiquitination of GMEB1 was also increased by overexpressing circGlis3 and decreased by inhibiting circGlis3 (Fig. 6k). Additionally, we tried to identify the domain where GMEB1 binds to circGlis3. Flag-tagged recombinant Gmeb1 plasmids with full-length (FL) or different forms of truncation were constructed and transfected into MS-1 cells (ESM Fig. 6h, i). In vitro binding assays indicated that the RNA-binding domain [1–157 amino acids (aa)] of GMEB1 was crucial for its interaction with circGlis3 (ESM Fig. 6j). Furthermore, we noticed that mindbomb E3 ubiquitin protein ligase 2 (MIB2) was detected in circGlis3 pull-down samples (ESM Table 7, ESM Fig. 6k). Western blot verified the existence of MIB2 and RIP assay showed the endogenous enrichment of circGlis3 in RNA coprecipitated by MIB2 (Fig. 6l,m). Coimmunoprecipitation assays indicated that GMEB1 and MIB2 bound to each other in MS-1 cells (Fig. 6n,o). Furthermore, circGlis3 overexpression enhanced the interaction between GMEB1 and MIB2 (Fig. 6p). Altogether, these findings indicate that circGlis3 directly binds to GMEB1 and promotes its degradation by facilitating the interaction between GMEB1 and MIB2.
CircGlis3 induces MS-1 cell dysfunction via the GMEB1/HSP27 signalling pathway
To further elucidate the functional roles of GMEB1, we knocked down Gmeb1 levels using siRNAs (Fig. 7a, ESM Fig. 7a). Inhibition of GMEB1 resulted in notably decreased MS-1 cell viability, cell proliferation, migration and angiogenesis, and increased cell apoptosis (Fig. 7b-e, ESM Fig. 7b, c). Then, the downstream targets of GMEB1 were explored. GMEB1 participates in functional regulation by interacting with proteins including heat shock protein 27 (HSP27) and pro-caspase 8 [30, 31]. Given that HSP27 is involved in cell migration, cell survival and other important biological functions [32, 33], we focused on exploring the interplay between GMEB1 and HSP27. Immunoprecipitation assay revealed that GMEB1 interacted directly with HSP27 in MS-1 cells (Fig. 7f). We also overexpressed GMEB1 in MS-1 cells (ESM Fig. 7d). The level of p-HSP27 was decreased by silencing GMEB1 and increased by overexpressing GMEB1, whereas the total HSP27 remained unaffected (ESM Fig. 7e). Consistently, inhibiting GMEB1 reduced proliferation marker and elevated pro-apoptotic protein levels (Fig. 7g). We next determined whether the detrimental impacts of circGlis3 on MS-1 cells were mediated by the GMEB1/HSP27 axis. As anticipated, overexpressing circGlis3 decreased cell viability, cell migration and angiogenesis, and increased cell apoptosis, which were rescued by elevating GMEB1 levels (Fig. 7h-k). Meanwhile, overexpressing GMEB1 ameliorated the dysregulated protein levels induced by overexpression of circGlis3 (Fig. 7l). Moreover, lipotoxic beta cell-derived exosomes or exosomes overexpressing circGlis3 decreased the protein levels of GMEB1 and p-HSP27 in MS-1 cells (ESM Fig. 7f). Overall, these data indicated that circGlis3 exerted its effects on MS-1 cells, at least partly, through the GMEB1/HSP27 pathway.
CircGlis3 regulates MS-1 cell function through the GMEB1/HSP27 signalling pathway. (a–e) Scramble (Scr) siRNA or siRNAs targeting Gmeb1 (siGmeb1-1 and siGmeb1-2) were transfected into MS-1 cells for 48 h. The protein levels of GMEB1 were evaluated by western blot (a). Cell proliferation was detected via EdU staining and data were analysed by Image J. Scale bar, 100 μm (b). Cell apoptosis was analysed by flow cytometry (c). Migration capacity was evaluated by transwell assays and images were obtained. Scale bar, 100 μm (d). Tube formation assays were performed and images were captured. Scale bar, 100 μm (e). (f) Immunoprecipitation analysed the association between GMEB1 and HSP27 in MS-1 cells. (g) Western blot detected the proteins (GMEB1, HSP27, p-HSP27, PCNA, Bcl-2 and cleaved caspase-3) expression after silencing Gmeb1. (h–l) MS-1 cells with circGlis3 overexpression were transfected with pcDNA-Gmeb1 plasmid (ov-Gmeb1) or empty vector (vector) for rescue assays. For apoptosis assessment, TUNEL staining was performed and the level of TUNEL-positive cells was calculated. Scale bar, 100 μm (h). Cell viability was detected by CCK-8 assay (i). Images of transwell assays (j) and tube formation assays (k) were photographed. Scale bar, 100 μm. Western blot analysed the levels of proteins (GMEB1, HSP27, p-HSP27, PCNA, Bcl-2 and cleaved caspase-3) in MS-1 cells (l). The numbers between each row in the western blots are the quantification levels, relative to the left-hand lane set to 1.00 (a, g, i). Data are expressed as mean ± SD of 3 independent experiments. *p < 0.05, **p < 0.01 and ***p < 0.001, by one-way ANOVA with Tukey post hoc test. Bcl-2, B cell lymphoma 2; PCNA, proliferating cell nuclear antigen; PI, propidium iodide
Beta cell-derived exosomal circGlis3 induces islet EC dysfunction in vivo
We then examined the effects of beta cell-derived exosomal circGlis3 on islet ECs in vivo. Mice were injected with MIN6 cell-derived exosomes (circGlis3-Exo, Vec-Exo) or PBS as illustrated in Fig. 8a. The uptake of exosomes by islet ECs was confirmed by immunostaining (ESM Fig. 8a). Here, CD31, a marker for endothelial cells, was applied to evaluate the vascular density in islets. CircGlis3-Exo reduced the density of islet ECs in HFD-fed mice, whereas no significant changes were observed in ND-fed mice (Fig. 8b). Moreover, CD31-positive islet ECs were separated by flow cytometry (Fig. 8c). CircGlis3-Exo increased the levels of circGlis3 and Bax, while reducing Bcl2 mRNA levels and the protein levels of GMEB1 and p-HSP27 in islet ECs of HFD-fed mice (Fig. 8d-f). Additionally, circGlis3-Exo elevated FBG levels and impaired glucose tolerance of HFD-fed mice, whereas no obvious impacts were found in ND-fed mice (ESM Fig. 8b-d). We think that the elevated FBG levels and impaired glucose tolerance may be ascribed to the islet EC dysfunction and the uptake of circGlis3-Exo by beta cells.
Beta cell-derived exosomal circGlis3 induces islet EC dysfunction in vivo. Exosomes were extracted from the medium of MIN6 cells overexpressing circGlis3 (circGlis3-Exo) or empty vector (Vec-Exo). C57BL/6 J mice fed with ND or HFD were randomly subdivided to receive intraductal injection of PBS, Vec-Exo or circGlis3-Exo. Subsequently, PBS or exosomes were injected into mice through the tail vein once a week for 4 weeks. Animals were analysed at day 31. (a) Scheme showing the timeline for exosome injection. (b) Mouse pancreatic sections were stained for CD31 (green) and DAPI (blue). Islet capillary density was quantified as the ratio of CD31-positive cells to islet area. Scale bar, 100 μm. Dashed lines indicate the area of islets. (c) At the end of the intervention, islets were isolated from three mice, pooled, and dissociated. Islet cells were stained with CD31-PE and islet ECs were sorted by flow cytometry. Arrows indicate CD31-positive islet ECs. (d, e) The levels of circGlis3 (d), Bcl2 and Bax (e) in separated islet ECs were determined by qPCR. (f) The protein levels of GMEB1 and p-HSP27 in islet ECs of HFD-fed mice were detected by immunofluorescence staining of pancreatic sections. Scale bar, 100 μm. Dashed lines indicate the area of islets. Results are expressed as mean ± SD of 3 independent experiments. *p < 0.05 and **p < 0.01, NS indicates no statistical significance, by one-way ANOVA with Tukey post hoc test. PE, phycoerythrin
Discussion
In this study, we demonstrate that circGlis3 participates in the development of type 2 diabetes in two different ways. CircGlis3 facilitated beta cell dysfunction by inhibiting cell survival and insulin secretion. Moreover, beta cell-derived exosomal circGlis3 impaired the function of islet ECs by regulating GMEB1 protein degradation and the downstream HSP27 phosphorylation. These findings not only reveal the role of circGlis3 in the pathophysiology of diabetes, but also suggest the feasibility of circGlis3 as a potential biomarker and therapeutic target for type 2 diabetes.
CircRNAs are critical regulators of diverse cellular processes and are implicated in regulating beta cell function [10, 34]. However, few studies have elucidated the impact of circRNAs on intercellular communication in type 2 diabetes. Our study identified a set of differentially expressed circRNAs in beta cell-secreted exosomes under lipotoxic conditions. Here, we focused on circGlis3, which is derived from exon 4 of the GLIS3 gene, a candidate gene for type 2 diabetes. GLIS3 encodes a transcription factor responsible for beta cell differentiation and insulin secretion [35]. We illustrated how circGlis3 caused damage to beta cells, suggesting that circGlis3 and its linear transcripts exert antithetical roles in diabetes. In addition, circularisation of exons can compete with canonical pre-mRNA splicing, indicating that circRNA biogenesis might be an important regulator of mRNA production [36]. We found that lipotoxicity upregulated circGlis3 levels, while reducing Glis3 mRNA levels in beta cells (data not shown). We speculate that lipotoxicity may affect the alternative splicing of Glis3 pre-mRNA, leading to the transition from canonical splicing to circularisation, which requires further investigation. Moreover, circGLIS3 (originating from exons 5, 6, 7 and 8 of the human GLIS3 gene) was upregulated in non-small cell lung cancer (NSCLC) and promoted NSCLC progression [37]. The inconsistent functions between circGlis3 and circGLIS3 may be attributed to different origination of circRNAs and different cell types.
CircRNAs exert functions in multiple ways, such as acting as miRNA sponges, interacting with proteins, or encoding functional peptides [27, 29, 38]. In this study, we uncovered that circGlis3 directly bound to GMEB1 protein and promoted its degradation by facilitating the interaction between GMEB1 and MIB2. It is reported that GMEB1 inhibits cell apoptosis by binding to procaspases, CASP8 and FADD like apoptosis regulator (CFLARL) and TNF receptor associated factor 3 (TRAF3) [39,40,41]. Here, we showed that GMEB1 interacted with HSP27 and promoted the phosphorylation of HSP27, which is associated with actin cytoskeleton organisation in vascular endothelial growth factor (VEGF)-stimulated endothelial cell migration [42], thereby favouring the survival and migration of islet ECs. Besides, HSP27 is phosphorylated by protein kinases including mitogen-activated protein kinase-activated protein kinase 2 (MK2), mitogen-activated protein kinase-activated protein kinase 3 (MK3), protein kinase B (PKB) and protein kinase C (PKC), and dephosphorylated by protein phosphatase 2A (PP2A) [43]. Therefore, it is reasonable to assume that GMEB1 may increase p-HSP27 levels by facilitating interaction between HSP27 and the kinases, or inhibiting the dephosphorylation of HSP27, which needs further exploration.
Exosome transport is considered to be an effective way to transmit signals and modulate biological function of recipient cells [15]. Notably, the ability of exosomes to transfer insulin-resistant phenotype and deleterious signals to insulin-sensitive tissues has been regarded as an important mechanism for the development of diabetes [19, 44]. We demonstrate that circGlis3 embedded in exosomes derived from lipotoxic beta cells was transmitted to islet ECs, thus inducing the dysfunction of recipient cells and aggravating the progression of diabetes. Pancreatic islets have a high density of capillary networks, and there is an endothelial–endocrine interaction during beta cell growth and islet angiogenesis [45]. Islet ECs are essential for islet revascularisation after transplantation and supporting beta cell survival [46]. Our in vivo studies show that knockdown of circGlis3 in beta cells not only ameliorated beta cell dysfunction under diabetic conditions, but also reduced islet EC loss. Meanwhile, beta cell-derived circGlis3 accelerated islet EC loss and beta cell dysfunction in diabetic mice. Of note, apart from islet ECs, beta cells are also the recipient cells of exosomes [47]. The uptake of exosomal circGlis3 by beta cells may be implicated in beta cell damage. Additionally, exosomal circGlis3 levels were elevated in the serum of mouse models of diabetes and participants with type 2 diabetes, indicating the potential of circGlis3 as a promising biomarker for type 2 diabetes. Furthermore, we found that circGlis3 overexpression impaired insulin secretion in vitro, whereas plasma insulin levels were increased in HFD-fed mice, which overexpressed circGlis3 in islets. It is widely accepted that hyperinsulinaemia is a compensation for HFD-induced systemic insulin resistance, and there is a vicious cycle between hyperinsulinaemia and insulin resistance [48]. The overloaded beta cells eventually fail to compensate, resulting in overt hyperglycaemia and type 2 diabetes [49]. In our study, despite the apparent hyperinsulinaemia, beta cell mass in HFD-fed mice was already decreased in the early stage of diabetes, indicating that the increased circGlis3 levels triggered by HFD may be involved in the progression of beta cell failure. Hyperinsulinaemia might be followed by hyposinsulinaemia over the time course. Interestingly, we also noticed that inhibiting circGlis3 in beta cells mitigated hyperinsulinaemia and improved insulin sensitivity in diabetic mice, which prompted us to speculate that the decrease in serum insulin levels with circGlis3 inhibition may be an adaptation to the improved peripheral insulin sensitivity. Beta cell-derived circGlis3 may be delivered to peripheral tissues including liver, skeletal muscle and adipose tissue via exosomes, thereby affecting peripheral insulin sensitivity. However, the pattern of beta cell-derived exosomal circGlis3 distribution and their functional roles in recipient organs are worthy of further study.
In summary, our findings reveal that circGlis3 not only contributed to lipotoxicity-induced beta cell dysfunction, but also participated in regulating islet EC function through the exosomal route. Importantly, our study elucidates a novel regulatory mechanism by which beta cells act on islet ECs and illustrates the significance of circGlis3 as a potential biomarker and therapeutic target for type 2 diabetes.
Data availability
The RNA-sequencing data have been deposited in the NCBI Sequence Read Archive (SRA) database (www.ncbi.nlm.nih.gov/sra), with accession number PRJNA689673. Mass spectrometry data are available via ProteomeXchange (www.proteomexchange.org/) with identifier PXD024693. All other data are within the paper and the ESM.
Abbreviations
- AAV8:
-
Adeno-associated virus serotype 8 vector
- AAV8-NC:
-
AAV8-negative control
- AAV8-shcircGlis3:
-
AAV8-circGlis3 shRNA
- circRNA:
-
Circular RNA
- CIRI:
-
CircRNA Identifier
- ddPCR:
-
Droplet digital PCR
- -Exo:
-
Exosomes (incubated with PA, BSA etc.)
- FBG:
-
Fasting blood glucose
- FISH:
-
Fluorescence in situ hybridisation
- Glis3:
-
Gli-similar 3
- GMEB1:
-
Glucocorticoid modulatory element-binding protein 1
- HFD:
-
High-fat diet
- HNF3β:
-
Hepatocyte nuclear factor 3β
- HSP27:
-
Heat shock protein 27
- Islet EC:
-
Islet endothelial cell
- MIB2:
-
Mindbomb E3 ubiquitin protein ligase 2
- ND:
-
Normal diet
- NSCLC:
-
Non-small cell lung cancer
- PA:
-
Palmitate
- qPCR:
-
Quantitative real-time PCR
- RIP:
-
RNA-binding protein immunoprecipitation
- TEM:
-
Transmission electron microscopy
References
Zheng Y, Ley SH, Hu FB (2018) Global aetiology and epidemiology of type 2 diabetes mellitus and its complications. Nat Rev Endocrinol 14:88–98. https://doi.org/10.1038/nrendo.2017.151
Johnson AM, Olefsky JM (2013) The origins and drivers of insulin resistance. Cell 152:673–684. https://doi.org/10.1016/j.cell.2013.01.041
Samuel VT, Shulman GI (2016) The pathogenesis of insulin resistance: integrating signaling pathways and substrate flux. J Clin Invest 126:12–22. https://doi.org/10.1172/JCI77812
Acosta-Montano P, Garcia-Gonzalez V (2018) Effects of dietary fatty acids in pancreatic beta cell metabolism, implications in homeostasis. Nutrients 10:393. https://doi.org/10.3390/nu10040393
Jezek P, Jaburek M, Holendova B, Plecita-Hlavata L (2018) Fatty acid-stimulated insulin secretion vs. lipotoxicity. Molecules 23:1483. https://doi.org/10.3390/molecules23061483
Hogan MF, Liu AW, Peters MJ et al (2017) Markers of islet endothelial dysfunction occur in male B6.BKS(D)-Leprdb/J mice and may contribute to reduced insulin release. Endocrinology 158:293–303. https://doi.org/10.1210/en.2016-1393
Narayanan S, Loganathan G, Dhanasekaran M et al (2017) Intra-islet endothelial cell and beta-cell crosstalk: implication for islet cell transplantation. World J Transplant 7:117–128. https://doi.org/10.5500/wjt.v7.i2.117
Olerud J, Mokhtari D, Johansson M et al (2011) Thrombospondin-1: an islet endothelial cell signal of importance for beta-cell function. Diabetes 60:1946–1954. https://doi.org/10.2337/db10-0277
Jiang Q, Liu C, Li CP et al (2020) Circular RNA-ZNF532 regulates diabetes-induced retinal pericyte degeneration and vascular dysfunction. J Clin Invest 130:3833–3847. https://doi.org/10.1172/JCI123353
Stoll L, Sobel J, Rodriguez-Trejo A et al (2018) Circular RNAs as novel regulators of beta-cell functions in normal and disease conditions. Mol Metab 9:69–83. https://doi.org/10.1016/j.molmet.2018.01.010
Kristensen LS, Andersen MS, Stagsted LVW, Ebbesen KK, Hansen TB, Kjems J (2019) The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet 20:675–691. https://doi.org/10.1038/s41576-019-0158-7
Weng W, Wei Q, Toden S et al (2017) Circular RNA ciRS-7-A promising prognostic biomarker and a potential therapeutic target in colorectal cancer. Clin Cancer Res 23:3918–3928. https://doi.org/10.1158/1078-0432.CCR-16-2541
Zheng Q, Bao C, Guo W et al (2016) Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun 7:11215. https://doi.org/10.1038/ncomms11215
Li Y, Zheng Q, Bao C et al (2015) Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res 25:981–984. https://doi.org/10.1038/cr.2015.82
Wang Y, Liu J, Ma J et al (2019) Exosomal circRNAs: biogenesis, effect and application in human diseases. Mol Cancer 18:116. https://doi.org/10.1186/s12943-019-1041-z
Chang W, Wang J (2019) Exosomes and their noncoding RNA cargo are emerging as new modulators for diabetes mellitus. Cells 8:853. https://doi.org/10.3390/cells8080853
Sun Y, Mao Q, Shen C, Wang C, Jia W (2019) Exosomes from beta-cells alleviated hyperglycemia and enhanced angiogenesis in islets of streptozotocin-induced diabetic mice. Diabetes Metab Syndr Obes 12:2053–2064. https://doi.org/10.2147/DMSO.S213400
Sun Y, Zhou Y, Shi Y et al (2021) Expression of miRNA-29 in pancreatic beta cells promotes inflammation and diabetes via TRAF3. Cell reports 34:108576. https://doi.org/10.1016/j.celrep.2020.108576
Xu H, Du X, Xu J et al (2020) Pancreatic beta cell microRNA-26a alleviates type 2 diabetes by improving peripheral insulin sensitivity and preserving beta cell function. PLoS Biol 18:e3000603. https://doi.org/10.1371/journal.pbio.3000603
Zhang X, Wang S, Wang H et al (2019) Circular RNA circNRIP1 acts as a microRNA-149-5p sponge to promote gastric cancer progression via the AKT1/mTOR pathway. Mol Cancer 18:20. https://doi.org/10.1186/s12943-018-0935-5
Zhao RT, Zhou J, Dong XL et al (2018) Circular ribonucleic acid expression alteration in exosomes from the brain extracellular space after traumatic brain injury in mice. J Neurotrauma 35:2056–2066. https://doi.org/10.1089/neu.2017.5502
Gao Y, Wang J, Zhao F (2015) CIRI: an efficient and unbiased algorithm for de novo circular RNA identification. Genome Biol 16:4. https://doi.org/10.1186/s13059-014-0571-3
Gao Y, Zhang J, Zhao F (2018) Circular RNA identification based on multiple seed matching. Brief Bioinform 19:803–810. https://doi.org/10.1093/bib/bbx014
Memczak S, Jens M, Elefsinioti A et al (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495:333–338. https://doi.org/10.1038/nature11928
Xiong L, Gong Y, Wu L et al (2020) LncRNA-Malat1 is involved in lipotoxicity-induced ss-cell dysfunction and the therapeutic effect of Exendin-4 via Ptbp1. Endocrinology 161:bqaa065. https://doi.org/10.1210/endocr/bqaa065
Grant CE, Bailey TL, Noble WS (2011) FIMO: scanning for occurrences of a given motif. Bioinformatics 27:1017–1018. https://doi.org/10.1093/bioinformatics/btr064
Wong CH, Lou UK, Li Y et al (2020) CircFOXK2 promotes growth and metastasis of pancreatic ductal adenocarcinoma by complexing with RNA-binding proteins and sponging MiR-942. Cancer Res 80:2138–2149. https://doi.org/10.1158/0008-5472.CAN-19-3268
Glazar P, Papavasileiou P, Rajewsky N (2014) circBase: a database for circular RNAs. RNA 20:1666–1670. https://doi.org/10.1261/rna.043687.113
Yang F, Fang E, Mei H et al (2019) Cis-acting circ-CTNNB1 promotes beta-catenin signaling and cancer progression via DDX3-mediated transactivation of YY1. Cancer Res 79:557–571. https://doi.org/10.1158/0008-5472.CAN-18-1559
Tsuruma K, Nakagawa T, Morimoto N et al (2006) Glucocorticoid modulatory element-binding protein 1 binds to initiator procaspases and inhibits ischemia-induced apoptosis and neuronal injury. J Biol Chem 281:11397–11404. https://doi.org/10.1074/jbc.M510597200
Theriault JR, Charette SJ, Lambert H, Landry J (1999) Cloning and characterization of hGMEB1, a novel glucocorticoid modulatory element binding protein. FEBS Lett 452:170–176. https://doi.org/10.1016/s0014-5793(99)00634-1
Chen H, Lu N, Xu C, Zheng M (2019) The role of heat shock protein 27 phosphorylation in the proliferation and apoptosis of human umbilical vein endothelial cells induced by Visfatin. Microvasc Res 121:30–36. https://doi.org/10.1016/j.mvr.2018.08.003
Shan R, Liu N, Yan Y, Liu B (2020) Apoptosis, autophagy and atherosclerosis: relationships and the role of Hsp27. Pharmacol Res 166:105169. https://doi.org/10.1016/j.phrs.2020.105169
Wu L, Xiong L, Li J et al (2020) Circ-Tulp4 promotes beta-cell adaptation to lipotoxicity by regulating soat1 expression. J Mol Endocrinol 65:149–161. https://doi.org/10.1530/JME-20-0079
Yang Y, Bush SP, Wen X, Cao W, Chan L (2017) Differential gene dosage effects of diabetes-associated gene glis3 in pancreatic beta cell differentiation and function. Endocrinology 158:9–20. https://doi.org/10.1210/en.2016-1541
Ashwal-Fluss R, Meyer M, Pamudurti NR et al (2014) circRNA biogenesis competes with pre-mRNA splicing. Mol Cell 56:55–66. https://doi.org/10.1016/j.molcel.2014.08.019
Wu Z, Jiang H, Fu H, Zhang Y (2021) A circGLIS3/miR-644a/PTBP1 positive feedback loop promotes the malignant biological progressions of non-small cell lung cancer. Am J Cancer Res 11:108–122
Zhang M, Zhao K, Xu X et al (2018) A peptide encoded by circular form of LINC-PINT suppresses oncogenic transcriptional elongation in glioblastoma. Nat Commun 9:4475. https://doi.org/10.1038/s41467-018-06862-2
An W, Yao S, Sun X et al (2019) Glucocorticoid modulatory element-binding protein 1 (GMEB1) interacts with the de-ubiquitinase USP40 to stabilize CFLARL and inhibit apoptosis in human non-small cell lung cancer cells. J Exp Clin Cancer Res 38:181. https://doi.org/10.1186/s13046-019-1182-3
Kotsaris G, Kerselidou D, Koutsoubaris D et al (2020) TRAF3 can interact with GMEB1 and modulate its anti-apoptotic function. J Biol Res (Thessalon) 27:7. https://doi.org/10.1186/s40709-020-00117-2
Nakagawa T, Tsuruma K, Uehara T, Nomura Y (2008) GMEB1, a novel endogenous caspase inhibitor, prevents hypoxia- and oxidative stress-induced neuronal apoptosis. Neurosci Lett 438:34–37. https://doi.org/10.1016/j.neulet.2008.04.023
Sawada J, Li F, Komatsu M (2015) R-ras inhibits VEGF-induced p38MAPK activation and HSP27 phosphorylation in endothelial cells. J Vasc Res 52:347–359. https://doi.org/10.1159/000444526
Kostenko S, Moens U (2009) Heat shock protein 27 phosphorylation: kinases, phosphatases, functions and pathology. Cell Mol Life Sci 66:3289–3307. https://doi.org/10.1007/s00018-009-0086-3
Ying W, Riopel M, Bandyopadhyay G et al (2017) Adipose tissue macrophage-derived exosomal miRNAs can modulate in vivo and in vitro insulin sensitivity. Cell 171:372–384.e12. https://doi.org/10.1016/j.cell.2017.08.035
Peiris H, Bonder CS, Coates PT, Keating DJ, Jessup CF (2014) The beta-cell/EC axis: how do islet cells talk to each other? Diabetes 63:3–11. https://doi.org/10.2337/db13-0617
Narayanan S, Loganathan G, Mokshagundam S, Hughes MG, Williams SK, Balamurugan AN (2018) Endothelial cell regulation through epigenetic mechanisms: depicting parallels and its clinical application within an intra-islet microenvironment. Diabetes Res Clin Pract 143:120–133. https://doi.org/10.1016/j.diabres.2018.06.018
Guay C, Menoud V, Rome S, Regazzi R (2015) Horizontal transfer of exosomal microRNAs transduce apoptotic signals between pancreatic beta-cells. Cell Commun Signal 13:17. https://doi.org/10.1186/s12964-015-0097-7
Inaishi J, Saisho Y (2020) Beta-cell mass in obesity and type 2 diabetes, and its relation to pancreas fat: a mini-review. Nutrients 12:3846. https://doi.org/10.3390/nu12123846
Heydemann A (2016) An overview of murine high fat diet as a model for type 2 diabetes mellitus. J Diabetes Res 2016:2902351. https://doi.org/10.1155/2016/2902351
Acknowledgements
The authors thank Z. Liang (Department of Colorectal Surgery, The Sixth Affiliated Hospital of Sun Yat-sen University) for his help in the RNA pull-down assay.
Funding
This project was supported by the Department of Finance of Guangdong Province (20160902), the National Natural Science Foundation of China (81701367) and Medical Research Foundation of Guangdong Province (A2021122).
Author information
Authors and Affiliations
Contributions
LX, YYG and HPX designed the study, analysed and interpreted data, and wrote and revised the manuscript. LX, LC, LTW, WMH, DBC and ZSP conducted the experiments, acquired and analysed data, and revised the article critically for important intellectual content. JL, XNZ, LS and YBL contributed to the analysis of data and revised the article critically for important intellectual content. All authors gave their approval of the version to be published. YYG and HPX are the guarantors of this work and take full responsibility for the content of the manuscript.
Corresponding authors
Ethics declarations
Authors’ relationships and activities
The authors declare that there are no relationships or activities that might bias, or be perceived to bias, their work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xiong, L., Chen, L., Wu, L. et al. Lipotoxicity-induced circGlis3 impairs beta cell function and is transmitted by exosomes to promote islet endothelial cell dysfunction. Diabetologia 65, 188–205 (2022). https://doi.org/10.1007/s00125-021-05591-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-021-05591-4