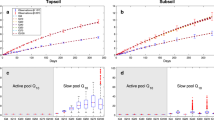
Abstract
Coupled biogeochemical cycles drive ecosystem ecology by influencing individual-to-community scale behaviors; yet the development of process-based models that accurately capture these dynamics remains elusive. Soil organic matter (SOM) decomposition in particular is influenced by resource stoichiometry that dictates microbial nutrient acquisition (‘ecological stoichiometry’). Despite its basis in biogeochemical modeling, ecological stoichiometry is only implicitly considered in high-resolution microbial investigations and the metabolic models they inform. State-of-science SOM decomposition models in both fields have advanced largely separately, but they agree on a need to move beyond seminal pool-based models. This presents an opportunity and a challenge to maximize the strengths of various models across different scales and environmental contexts. To address this challenge, we contend that ecological stoichiometry provides a framework for merging biogeochemical and microbiological models, as both explicitly consider substrate chemistries that are the basis of ecological stoichiometry as applied to SOM decomposition. We highlight two gaps that limit our understanding of SOM decomposition: (1) understanding how individual microorganisms alter metabolic strategies in response to substrate stoichiometry and (2) translating this knowledge to the scale of biogeochemical models. We suggest iterative information exchange to refine the objectives of high-resolution investigations and to specify limited dynamics for representation in large-scale models, resulting in a new class of omics-enabled biogeochemical models. Assimilating theoretical and modelling frameworks from different scientific domains is the next frontier in SOM decomposition modelling; advancing technologies in the context of stoichiometric theory provides a consistent framework for interpreting molecular data, and further distilling this information into tractable SOM decomposition models.
Similar content being viewed by others
Introduction
The world’s soils contain a pool of carbon (C) that is larger than vegetation and atmospheric stocks combined, with 1500 Gt C stored in the top one meter alone (Batjes 2016). A large proportion of belowground C is contained in soil organic matter (SOM) whose destabilization has the potential to dramatically alter future climates (Crowther et al. 2019). SOM decomposition is regulated by physical, chemical, hydrologic, and biological controls (collectively called biophysical controls) on resource availability that determine microbial energy generation and C and nutrient acquisition (Robertson et al. 2019; Wang and Houlton 2009; Zhang et al. 2014). These processes are notoriously difficult to measure and predict beyond the scale of experimental plots (Bond-Lamberty et al. 2016; Naylor et al. 2020). Because of this, SOM decomposition at scales most relevant to climate change continues to be predicted mainly through generalized environmental proxies such as moisture, temperature, minerology, and total soil C or nutrient pool sizes (Bailey et al. 2018)—parameters that leave a substantial amount of uncertainty surrounding model predictions (Todd-Brown et al. 2013, 2014). Given the increase in disturbances and novel environments created by changes in our global climate, the shortcomings of predicting SOM decomposition beyond localized scales point to a need for scalable spatial and biogeochemical processes in SOM decomposition models (Todd-Brown et al. 2013).
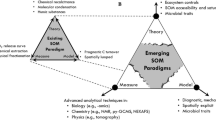
This topic has received considerable attention from multiple scientific domains over the past few decades, resulting in parallel advances between biogeochemical and microbial research (Fig. 1) and calls for greater diversity of measurements used within biogeochemical models (Billings et al. 2021). Collectively, we have made outstanding progress in our understanding of global C cycles; however, a key opportunity remains in integrating theoretical and modelling frameworks from different scientific domains. Existing classes of models each have strengths at different scales and environmental contexts—where one model fails, another excels. Recent work has improved conceptual, empirical, and numerical representations of mineral protection and hydrologic mechanisms that regulate microbial access to SOM (Blankinship et al. 2018; Waring et al. 2020; Woolf and Lehmann 2019). In contrast, detailed representations of the microbial metabolisms that directly convert SOM into carbon dioxide are still missing from predictive models at the ecosystem-scale (Stone et al. 2021). Soil organic matter decomposition (particularly unprotected SOM) is often rate-limited by microbial metabolism (Dwivedi et al. 2019; Hunter et al. 1998; Lehmann and Kleber 2015), which is controlled by microbial access to substrates (Schimel 2021). In turn, microbial decomposition is coupled to necromass stabilization that is now thought to be the primary mechanism of belowground C storage (Cotrufo et al. 2013; Kallenbach et al. 2015, 2016; Liang et al. 2019; Robertson et al. 2019; Zhang et al. 2021; Fig. 2). Microbial processes and soil physical properties are therefore central considerations in SOM modelling, but they are still coarsely represented in cutting-edge SOM decomposition models. Due to recent technological achievements, we now have the ability to measure soil biophysical properties with high molecular resolution (i.e. genes, transcripts, proteins, and metabolites, and soil mineralogy, porosity, and chemistry). Our current challenge lies in interpreting and scaling these vast molecular data types into information useful for biogeochemical models (Hall et al. 2018; Malik et al. 2020). Next-generation models that are able to link across molecular to ecosystem scales should enable more robust predictions of SOM decomposition (Chowdhury et al. 2019), with follow-on improvement in global C cycling predictions.
Parallel development of processed-based biogeochemical and microbial models across scales of molecular resolution. A short description of a representative class of models from biogeochemical (left) vs. microbiological (right) research domains is listed within each box. Model types that bridge between microbial and biogeochemical research domains are in the center column. High molecular resolution models are on the top row and low molecular resolution models are on the bottom row. Modelling frameworks that leverage ecological stoichiometry are depicted in red, and opportunities to use these principles to guide multiscale model development are depicted in blue. We present opportunities for stoichiometry to serve as an underlying framework to integrate state-of-science SOM decomposition models across ecological hierarchies as well as across biogeochemical and microbial disciplines – from individual microorganisms to bulk pools and fluxes. Ecological stoichiometry is implicitly used to guide both biogeochemical and microbiological models and therefore presents a natural linkage point for assimilating and scaling models from both research domains
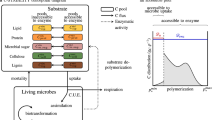
Overview of Factors Regulating Soil Carbon Cycles. A Microbial metabolism is the proximate control over bioavailable C cycling. These controls dominate in unprotected SOM pools in which accessibility is not limited and provide an avenue for stoichiometry to improve model predictions. B Mineral-association and hydrologic variables are also key components of SOM cycling, particularly in deeper soil horizons with high mineral content. C These conceptualizations are consolidated within the MEMS framework. In MEMS, SOM is more readily bioavailable in the organic horizon where growth of microbial biomass and to a lesser extent dissolved organic matter support C flow to and stabilization within mineral horizons. Panels B and C are reproduced from Jilling et al. (2018) and Robertson et al. (2019), as permitted under the creative commons license
Research opportunities for integrating microbial and biogeochemical modelling approaches
The importance of various controls on decomposition rates differs with the state of SOM at a given point in time and space, the portion of the soil system that is being represented in a given model, and with the predictive goal of the model. Biophysical attributes that regulate SOM decomposition collectively vary with soil type and horizon (Patel et al. 2021; Rumpel and Kögel-Knabner 2011); and SOM pool size and composition is dynamic within a single environment. SOM is continuously added to soils through plant litter production, root turnover, and rhizodeposition, while simultaneously being metabolized through microbial decomposition that leads to secondary metabolites and biomass synthesis, all of which have multiple possible routes through soil C cycles (Cotrufo et al. 2013; Liang et al. 2019; Sulman et al. 2014; Wieder et al. 2014). Small changes in the balance of various SOM inputs and subsequent decomposition processes impact the overall fate of C within the Earth system. Embedded in this is an understanding of rate limitation, as soil biology interacts with physiochemistry to constitute a holistic soil ecosystem. In some cases, SOM decomposition may be regulated by abiotic interactions with soil biology—relationships between SOM substrates and microbial decomposition are perhaps most important in the soil organic horizon and within the rhizosphere in mineral layers, where decomposition ultimately regulates C flow to persistent SOM pools.
These ideas have been formalized into a biogeochemical model that centers on mineral stabilization while also highlighting the controls of microbial metabolic efficiency on rates of SOM formation and accumulation (Robertson et al. 2019), Fig. 2. While there may be physical limits on the amount of SOM that can accumulate in mineral layers (the C-saturation hypothesis, Six et al. 2002; Stewart et al. 2007), Craig et al. (2021) recently used a simulation model to show that microbial community dynamics which impact microbial biomass C flow are also strong regulators of mineral-stabilized SOM. As such, microorganisms impact SOM cycling across all soil horizons. Understanding the strength and mechanisms of this regulation across different portions of the soil system (e.g. soil fractions and horizons) will further help constrain SOM model development but is beyond the scope of this paper.
Key foci of biogeochemical modelling efforts to improve estimates of SOM decomposition and/or accumulation include mineral-SOM interactions, soil hydrologic properties, chemical speciation of SOM, and microbial bimass and/or microbial use efficiency (Dungait et al. 2012; Robertson et al. 2019; Sierra and Müller 2015; Wieder et al. 2014; Zhang et al. 2021). Physicochemical protection (e.g. mineral-associated ‘MAOM’ or ‘protected pool(s)’ vs. ‘unprotected pool(s)’ including rhizosphere and particulate organic matter) and SOM chemistry in particular are pervasive across most classes of biogeochemical models (Robertson et al. 2019; Sulman et al. 2014; Zhang et al. 2021). For instance, the PROMISE framework centers on SOM accessibility as a key determinant of decomposition, encompassing the known roles of clay mineral composition and heterogeneity in residence times of identical molecules (Müller and Höper 2004; Waring et al. 2020). State-of-science biogeochemical frameworks, even those with an emphasis on physical processes, acknowledge chemical diversity within all SOM pools by using chemical parameters to constrain the fate of various classes SOM and/or generate a distribution of decomposition rates for a given pool (Azizi-Rad et al. 2021; Robertson et al. 2019; Waring et al. 2020; Zhang et al. 2021).
While the rate-limiting biophysical factor(s) fluctuates with environmental context, microbial metabolism is the proximate control of bioavailable SOM decomposition in all cases. Because the connectivity between protected and unprotected SOM is already encompassed by state-of-science biogeochemical models, we largely consider processes occurring within unprotected SOM pools when considering avenues for consolidating microbial and biogeochemical models. While we recognize that pore-scale spatial data is critical to understanding SOM decomposition, these data types remain difficult to obtain and are not yet widely used. In contrast, omics data are becoming widespread, and information on microbial communities reflect their underlying physical and chemical habitat. Process-based models arising from microbiology generally represent much smaller spatial and temporal scales than biogeochemical models, with inputs comprised of higher molecular resolution data. They tend to focus on genomic pathways or microbial traits that lead to decompostion (Borer et al. 2019; Borer and Or 2021; Heinken et al. 2021; Malik et al. 2020; Wang and Allison 2021).
Each class of microbial and biogeochemical models provides valuable information on SOM decomposition while also acknowledging considerable error and seeking opportunities for improvement. Therefore, a pressing question remains: how do we merge disperate model structures and underlying theoretical assumptions such that we maximize our predictive power of SOM decomposition, mineralization, and stabilization?
We see a ripe opportunity for harmonizing microbial and biogeochemical SOM models within and across all scales through substrate (plant and microbial) chemistries, which determine SOM composition (from a biogeochemical perspective) as well as the rate and pathway of microbial metabolism (from a microbial perspective). Billings et al. (2021) have identified SOM chemistry as a measurement for the next generation of soil C research, highlighting the increasing usage of molecular advances such as Fourier-Transformed Infrared Spectroscopy (FTIR, Cheng et al. 2006; Keiluweit et al. 2010), mid-infrared spectral libraries (Dangal et al. 2019), and Diffuse Reflectance Fourier-Transformed Infrared (DRIFTS, Kaiser and Ellerbrock 2005; Leue et al. 2010). Substrate chemistry is represented in fine-scale biogeochemical models (e.g. reaction networks and reactive transport models) by influencing reactions with specific SOM molecules, extracellular enzymes, or inorganic nutrients (Porta et al. 2018; Riley et al. 2014; Wilson et al. 2019). Microbiology also considers substrate chemistry at the individual level (e.g. single cell metabolic models), albeit with different terminology, in the framework of metabolomics. Metabolic models consider substrate chemistry in the context of a model organism’s genome to make predictions of growth dynamics or specific metabolomic products (e.g. GEMS, Duarte et al. 2004; Loira et al. 2012; Lu et al. 2019). Biogeochemical models at intermediate scales consider the composition of elemental and SOM pools, often through relatively coarse parameters such as fast vs. slow cycling SOM pools (unprotected vs. protected), mineral SOM, chemical classes or C:N ratios of SOM, and/or organic vs. inorganic nutrient pools. Many recent advances to consider SOM composition have been incorporated in MEND (Wang et al. 2015, 2020a), RESOM (Tang and Riley 2015), CORPSE (Sulman et al. 2014) and MIMICS-CN (Kyker-Snowman et al. 2020), yet none leverage molecular chemical information or stoichiometry of compounds or elements other than C and N. Microbial models at the intermediate-scale simulate interactions between individuals, limited sets of individuals (or as cumulative “mixed-bag” organisms)(Henry et al. 2016), or their characteristic traits to predict metabolite chemistry or changes in the growth of specific organisms or functional guilds (Bouskill et al. 2012; Follows and Dutkiewicz 2011; Malik et al. 2020; Reed et al. 2014). Recent research has also implicitly used the intersection of biogeochemistry and microbiology through substrate chemistry in order to move towards integration by representing fine- (individual microbial metabolisms and metabolites), intermediate- (groups of microorganisms and/or elemental pool composition), and coarse-resolution (multiple interacting microbial groups, “mixed bag” communities, and/or elemental pool sizes) connectivity of microbial drivers and biogeochemical outcomes in predictive models at a particular level of resolution (Figs. 1, 3 and 4). Most of the aforementioned biogeochemical models also include some aspects of mineral-OM interactions and/or soil hydrologic properties.
Representation of state-of-science SOM decomposition models across scales of molecular resolution. At the highest level of molecular resolution (top), genome-enabled models predict the production and consumption of specific metabolites by specific microorganisms or groups of microorganisms. This class of models is rooted in fundamental microbiology. At intermediate scales, community-scale (mixed-bag) metabolic models and reaction networks have been largely separately developed in microbiology and biogeochemistry, respectively. Finally, at coarser levels of molecular resolution, biogeochemical models predict SOM decomposition using nutrient-regulated exchange between lumped SOM pools with specific chemical attributes
Schematic of opportunities for ecological stoichiometry to guide iterative approaches that generate omics-enabled biogeochemical models. Ecological stoichiometry can guide a cycle of top-down and bottom-up approaches for more efficient generation of next generation models. Top-down approaches use nutrient limitations inferred from bulk biogeochemical measurements to guide the determination of rate-limiting pathways in SOM decomposition. In turn, rate-limiting pathways can reveal specific metabolic pathways and/or microorganism for detailed representation in predictive models. Bottom-up approaches use ecological stoichiometry as a framework for guiding the interpretation of high molecular resolution data streams with the help of new machine learning algorithms, for example by focusing on metabolic pathways that tend to be involved in nutrient acquisition. Using these approaches to identify the rate-limiting steps of SOM decomposition can then aid in the parameterization and calibration of larger-scale models by informing the tuning of substrate use efficiencies and/or by revealing pathways for more detailed representation. By using both approaches iteratively, we can identify specific portions of genome-enabled reaction networks that add predictive power with detailed representation in large-scale SOM decomposition models. A schematic of iterative top-down and bottom-up approaches is shown on the left. This leads to the development of a stoichiometry-informed reaction networks (middle) that can be merged into existing state-of-science SOM decomposition models (right). The MEMS model on the righthand side of the figure is reproduced from Robertson et al. (2019) under the creative commons license. The representation of the MEND model is reproduced from Wang et al. (2020a) with permission from the Elsevier publishing company
Despite their coarse representation of microbial processes, biogeochemical models with explicit microbial dynamics represent an improvement over seminal biogeochemical models (Wieder et al. 2013). This has led to soil microbiology research aimed at improving predictions of SOM decomposition through microbial genomics. For instance, Stone et al. (2021) recently demonstrated that the efficiency of microbial SOM decomposition co-varies with taxonomy and soil nutrient status. Other efforts include the prediction of microbial carbon use efficiency from microbial metagenomes (Saifuddin et al. 2019), microbial function under warming (Guo et al. 2020), and SOM decomposition from genome-derived microbial traits (Malik et al. 2020; Wang and Allison 2021)—all of which emerge from decades of work in microbial ecology aiming to understand the role of microbiome composition in SOM decomposition (Graham et al. 2016). State-of-science models in both biogeochemistry and microbiology arise out of a need to move beyond pool-based models towards continuous relationships between microbial metabolism and SOM decomposition and persistence (Tang and Riley 2020; Waring et al. 2020), but they rely on different underlying frameworks such that there are few common concepts, parameters, and equations with which to connect biogeochemical and microbial models.
Ecological stoichiometry (i.e. the role of elemental composition in regulating ecological interactions, Sterner and Elser 2002; Van de Waal et al. 2018) is a useful framework for cross-scale and cross-domain integration. While many biogeochemical models are built on the principles of ecological stoichiometry, they are largely absent from microbial models. Ecological stoichiometry uses the elemental (e.g. C:H:N:O:P:S) imbalance between organisms and their substrates to predict that nutrient limitations universally control biological activity (Elser et al. 1996, 2000; Sterner 1995). It has been leveraged to decipher overarching patterns in the processes governing SOM decomposition (Billings & Ballantyne IV 2013; Sinsabaugh et al. 2013; Sinsabaugh and Shah 2011) and is the guiding framework for decades of biogeochemical investigations that spawned process-based models.
Biogeochemical modelling research has begun to explore the usage of ecological stoichiometry across a broad range of scales. At the largest scale (pool models), a recent update to the CENTURY model (Parton 1996) demonstrates increased accuracy from a tighter coupling of C and N dynamics (Berardi et al. 2020). At higher molecular resolution, Buchkowski et al. (2019) have improved predictions of SOM decomposition by more explicitly accounting for the stoichiometries of microbial biomass and Fatichi et al. (2019) have proposed direct representation of microbial communities involved in coupled SOM and nutrient cycling, an approach that could not only aid in increasing the accuracy of SOM decomposition predictions but also become a useful tool for empiricists to interpret multidimensional microbial data.
By using an ecological stoichiometry framework to combine new microbiological and computational tools with biogeochemistry, we have an unexploited opportunity to combine predictive models across measurement scales and to better understand global patterns in microbial SOM decomposition. The use of top-down (ecosystem to molecular) and bottom-up (molecular to ecosystem) information exchange guided by ecological stoichiometry can iteratively (1) refine the objectives of high molecular resolution investigations and (2) specify a limited set of dynamics for representation in large-scale models through omics-enabled reaction networks (Fig. 4). Top-down approaches leverage bulk biogeochemical pool sizes to identify nutrient limitation(s) that then guide parameters selection and calibration of reduced complexity models. Bottom-up approaches use stoichiometric principles to guide the analysis and interpretation of large molecular datasets by establishing predictive rules that are translatable across scales. We propose that coupling these methods can specifically improve representations of SOM decomposition within the soil organic layer that ultimately regulate C transport and stabilization in mineral horizons. By using ecological stoichiometry as a conceptual and modelling framework to transfer knowledge across scales and couple existing models, we have the potential to develop a new generation of reduced complexity omics-enabled models.
Ecological stoichiometry as a guiding framework in soil organic matter decomposition
Microbial (i.e. fungal, bacterial, archaeal, and viral) degradation of chemically diverse SOM is reliant on broader organismal and soil stoichiometry (Buchkowski et al. 2015, 2019; van Groenigen et al. 2006). Nutrient availabilities in soil determine the metabolic pathways (and their efficiencies) and substrates that are used during SOM decomposition (Ge et al. 2020; Sinsabaugh et al. 2016; Wei et al. 2020). In turn, microbial decomposition and necromass products fuel a cycle of SOM decomposition, formation, and stabilization. Much belowground nitrogen (N) and sulfur (S) is stored within organic matter that is mineralized by soil microbial communities (Freney 1986; Jenkinson 1990; Kirkby et al. 2011). While phosphorus (P) availability is often associated with mineral weathering, the decomposition of organic matter is a key source of P in many ecosystems (Heuck et al. 2015; Margalef et al. 2017). Microbial decomposition increases the bioavailability of plant inputs and SOM by cleaving polymeric compounds into simple substrates that can be directly assimilated by microorganisms. This process is often catalyzed by extracellular enzymes that release nutrients from organic matter, connecting C, N, S, and P cycling (Allison et al. 2014). Viral predation can also influence these dynamics by lysing specific clades of microorganisms and by influencing organic matter accessibility to microorganisms (Kuzyakov and Mason-Jones 2018; Weinbauer 2004; Wilhelm and Suttle 1999). Global patterns of mechanisms governing SOM formation and cycling and the specific environmental conditions under which certain metabolic pathways are favored by microbial communities are active areas of research.
Identifying parameters that regulate decomposition across different ecosystems is challenging because heterogeneity in nutrient statuses, substrate energetics, mineral and hydrologic properties, and microbial communities can lead to different apparent controls in different soils (Milcu et al. 2011; Sullivan et al. 2014). In some cases, rates of decomposition are linked to microbial adjustments in the acquistion of specific elements in response to differences between microbial biomass and substrate stoichiometries (i.e. nutrient limitations, Billings & Ballantyne IV 2013; Billings and Ziegler 2008; Frost et al. 2005; Manzoni 2017; Manzoni et al. 2008; Spohn 2016; Sterner and Elser 2002). This research is rooted in the paradigm that organisms exhibit stoichiometric homeostasis in which they maintain stable biomass elemental ratios regardless of substrate stoichiometry. It leads to the prediction that nutrient limitations universally regulate biological activity (Redfield 1958; Spohn 2016). Specifically, the relative strength of the relationship between microbial metabolisms involved in N, S, or P acquisition and SOM decomposition should be strongest in soils with the lowest underlying proportion of that element. Indeed, the stoichiometry of microbial biomass features relatively little global variation (60:7:1 C:N:P) (Cleveland and Liptzin 2007) and substrate stoichiometries are highly variable (Frost et al. 2005; Manzoni 2017; Manzoni et al. 2008; Spohn 2016; Sterner and Elser 2002), supporting a dynamic in which microorganisms must adapt to account for soil nutrient concentrations.
The dependency of SOM decomposition on soil nutrients has been investigated most frequently using a suite of biogeochemical techniques in observational or fertilization studies such as CO2 flux, mass loss, bulk chemistry (e.g. C:N), microbial biomass measurements, and/or potential enzyme activity measurements (Sullivan et al. 2014). Each of these approaches provides indirect evidence for nutrient-regulated microbial decomposition using comparatively low molecular resolution. Additionally, the majority of researchers quanitfy total or coarsely-defined lumped C pools with minimal characterization of specific SOM compound classes or their stoichiometry. For instance, the classical pool and flux model focuses on three or four operationally defined C pools that are connected by biological drivers of decomposition, namely extracellular enzymes (Abramoff et al. 2018). As such, current estimates of nutrient limitations on SOM decomposition vary widely (Cleveland et al. 2006; Grandy et al. 2008; Kirkby et al. 2013, 2014; Sullivan et al. 2014; Takriti et al. 2018), and the majority of research has focused on a few elements, generally on linkages between C and N cycling (Cleveland et al. 2006; Grandy et al. 2008; Khan et al. 2016; Kirkby et al. 2013, 2014).
Microbial potential enzyme activity remains the primary tool in biogeochemical research for measuring the decomposition potential of specific compound classes in SOM pools. Extracellular enzymes catalyze the depolymerization of complex SOM compounds into simpler molecules for direct uptake and are ubiqutous in soils (Allison et al. 2010). The standard suite of enzyme assays target organic substrates including proteins, carbohydrates, amino sugars, organic phosphates, and lignins (Allison et al. 2007; Burns 1982; Sinsabaugh and Shah 2011). However, several microbially-mediated steps in SOM decomposition are more under-investigated relative to others. Measurements of phenol oxidases, peroxidases, and sulfatases in particular tend to be constrained by methodological limitations, and published enzyme assay methods are often used improperly across gradients of pH, co-factors, or substrate concentrations (Nannipieri et al. 2018; Schimel 2021). Additionally, while the ratios of enzyme activities have been used to assess nutrient limitations (Grandy et al. 2008; Hill et al. 2014; Jing et al. 2020; Moorhead et al. 2013; Sinsabaugh et al. 2008), these assays can be influenced by pH and substrate availability and only provide potential rates. They are low molecular resolution compared to many omics-based approaches because they target chemical bonds contained by classes of SOM rather than specific molecules, and they can have long turnover times that decouple them from SOM cycling at a specific point in time (Schimel et al. 2017).
Microbial investigations into nutrient-regulated SOM decomposition have traditionally used a complementary set of tools, revealing differences in metabolic strategies to maintain stoichiometric homeostasis using lumped traits (e.g., community-level variation between bacteria and fungi or copiotrophic and oligotrophic metabolisms, Elser et al. 2003; Fierer et al. 2007; Strickland and Rousk 2010). Changes in microbial heterotrophy has also been suggested to vary microbial preferences for C vs. N globally, an inference drawn from primarily biogeochemical data (Taylor and Townsend 2010). Understanding how changes in metabolic strategies of individual microorganisms responding to changes in substrate stoichiometry scale to a microbial community’s elemental use efficiency and capacity for decomposition is a key unknown in process-based modelling.
Omics-based characterization of microbial communities (i.e. metagenomic, metatranscriptomic, metaproteomic, and metabolomic) and other high molecular resolution approaches (e.g. chemical probes) provide direct means to evaluate how stoichiometry regulates the ecology of decomposition. These approaches have been highlighted as emerging tools in ecological stoichiometry but are not yet widely implemented (Van de Waal et al. 2018). Microbial genes and their expression patterns can reveal the genetic potential (genes), expression (transcripts), and translation (proteins) of specific enzymes involved in decomposition and more directly evaluate if biological N, S, and P acquisition vary predictably with SOM stoichiometry. While these modern microbial approaches have their own limitations (e.g. data annotation and discrepancies in turnover time when compared to rates), they provide a more complete representation of biogeochemical cycles than enzyme potential assays.
Several potential analysis targets for stoichiometric regulation of SOM decomposition have been identified in omics pipelines. For example, Finn et al. (2020) and Wilhelm et al. (2019) proposed lists of relevant Kyoto Encyclopedia of Genes and Genomes (KEGG) Orthologies and Carbohydrate-Active enzyme database (CAZy) annotations. These contain enzymes involved in the decomposition of organic C as well as N, S, P, and iron cycles, including commonly measured enzymes. They span many classes of enzymes involved in soil C and nutrient cycling, including those targeting cellulose, hemi-celluloses, lignin, cellobiose, mineral and organic N, mineral and organic P, and mineral and organic S. These provide a tangible and comprehensive set of microbial genes, transcripts, proteins, and metabolites that is consistent with biogeochemical methods used to investigate SOM decomposition. New annotations for molecular markers involved in SOM decomposition continue to be discovered, and our ability for molecular resolution into stoichiometry-based processes, if viewed within a consistent framework to biogeochemical applications, can help illuminate patterns where biogeochemical models lack mechanistic resolution.
Opportunities for ecological stoichiometry and high-resolution measurements in SOM decomposition modelling
Stoichiometric relationships underlie the kinetics of SOM decomposition (for instance, through nutrient limitations) and therefore are essential to developing and incorporating omics-enabled models into ecosystem scale models. The enormous amount of information about soil microbial and chemical diversity provided by new molecular data types complicates these efforts (Jansson and Hofmockel 2018) such that most current SOM decomposition models cannot yet account for cutting-edge molecular approaches to represent detailed microbial processes. Additionally, there is a need to move beyond models that separate SOM by chemical, physical, and functional properties (e.g., pool-based models) to more fluid relationships between microorganisms and SOM decomposition (Abramoff et al. 2018; Tang and Riley 2020; Waring et al. 2020). Microbial data mining approaches have made incremental advances to this effort, but remaining obstacles underline an opportunity for broader use of ecological frameworks, such as stoichiometric theory, to guide targeted investigations particularly with respect to unprotected pools of SOM (Fig. 2). This approach can help determine which pathways to include in any metabolic model and further scale high molecular resolution predictions to soil ecosystems (Zakem et al. 2020). Below, we highlight opportunities for ecological stoichiometry to inform the development and merging of high-resolution models such as omics-enabled reaction networks with biogeochemical models at larger scales (Fig. 4).
State-of-science models already implicitly use ecological stoichiometry to unify microbiology and biogeochemistry at intermediate scales, as they constrain rates of SOM decomposition with parameters that represent nutrient availability, stoichiometries of organic substrates or microbial biomass, and carbon and nutrient use efficiencies [e.g., MEND (Wang et al. 2015, 2020a), RESOM (Tang and Riley 2015), and MIMICS-CN (Kyker-Snowman et al. 2020)]. These models arise from biogeochemical research and are based on C pools with lumped characteristics. However, they generally include two elements at most, in part due to computational barriers to representing all processes in all elemental cycles as well as their interconnectivity (Abramoff et al. 2018; Moorhead et al. 2012; Sulman et al. 2019; Wang et al. 2015; Wieder et al. 2015).
A small set of coarse parameters is often used to denote microbial processes within state-of-science biogeochemical models, in particular C and nutrient use efficiencies (CUE, NUE) that describe C and nutrient conversion into microbial biomass (Allison et al. 2010; Ballantyne IV & Billings 2018; Cleveland and Liptzin 2007; Geyer et al. 2016; Min et al. 2016; Sihi et al. 2016; Sinsabaugh et al. 2013; Sinsabaugh et al. 2016; Wang et al. 2015; Wieder et al. 2014). CUE, in particular, has been shown to dramatically alter predictions of belowground C flux in Earth system’s models (Ballantyne IV & Billings 2018). CUE and NUE incorporate the principles of ecological stoichiometry into microbial-explicit biogeochemical models by serving as proxies for generalized microbial activity. They are most often estimated from empirical measurements of organic matter stoichiometry and/or microbial biomass, or from the ratios of the potential activity of C- vs. nutrient-acquring enzymes (Manzoni et al. 2012; Sinsabaugh et al. 2016). Under nutrient limitation, microorganisms reduce their C uptake, acquire nutrients, and/or respire or excrete excess C as enzymes or metabolites such that C assimilation into biomass often declines (Anderson et al. 2005; Del Giorgio and Cole 1998; Manzoni 2017; Manzoni et al. 2008, 2017; Middelboe and Søndergaard 1993; Milcu et al. 2011; Mooshammer et al. 2014b; Smith and Prairie 2004; Sterner and Elser 2002). Thus, CUE typically decreases when nutrients are limiting. In contrast, when nutrients exist in excess, microorganisms adjust their uptake or rate of respiration/excretion to decrease NUE (Milcu et al. 2011; Mooshammer et al. 2014a).
The balance of stoichiometric impacts on decomposition is difficult to predict due in part to microbial interactions (i.e., stoichiometry influences individual behavoir rather than community-level behavoir that ultimately regulates decomposition, Manzoni 2017; Manzoni et al. 2012; Sterner and Elser 2002), highlighting an opportunity for improvement through omics-enabled research. There is also large variation in the relationship of microbial CUE and NUE to SOM decomposition due to spatial and temporal changes in resource availability, microbial community structure, and soil physical properties (Frey et al. 2013; Herron et al. 2009; Malik et al. 2018; Manzoni et al. 2012; Qiao et al. 2019; Sinsabaugh et al. 2013, 2016). Biogeochemical models have shown that small changes in CUE can have large impacts on SOM decomposition (Li et al. 2014; Six et al. 2006; Wieder et al. 2013), offering an intriguing possibility for model improvement through detailed representations of nutrient-regulated microbial metabolic pathways to develop scalable omics-enable reaction networks.
At the finest molecular resolution where microbial information is best-suited (Fig. 3), there is a limited set of modelling approaches that currently predict rates of SOM decomposition, largely through by representing genomic and/or metabolic processes. Omics-enabled metabolic models, in particular, infer function from molecular data types by placing annotations within the context of a cell’s biochemical abilities to consume substrates, grow, and produce energy (Cuevas et al. 2016). They can include thousands of genes, metabolites, transcriptomes, proteomes, and associated reactions (Lieven et al. 2020; Seaver et al. 2020). However, they are limited by a set of biochemical reactions selected a priori that define the possible functions of microorganisms and provide a structure in which to root functional annotations (Seaver et al. 2020). In tandem, flux balance analysis is widely used within omics-enabled models to predict fluxes through the reactions in a metabolic network (Cuevas et al. 2016; Orth et al. 2010). These omics-enabled models can also be coupled to other methods of decoding high-resolution molecular data, such as thermodynamic theory (Garayburu-Caruso et al. 2020; Song et al. 2020), deep learning (Zampieri et al. 2019), or network modelling (Kessell et al. 2020; McClure et al. 2020) to further extend their applicability.
We see opportunity for ecology stoichiometry to provide this framework by guiding representations of microbial metabolism in biogeochemical models, particularly where stoichiometric regulation already exists in the unprotected or biologically-available SOM pools of many microbial-explicit biogeochemical models (e.g. MEND-CN, MIMICS-CN), and as a complement to abiotic controls in mineral soil (Fig. 4). Ecological stoichiometry influences the behavior of individual microorganisms, as they act to maintain stoichiometric balance through adjusting growth rate or biomass stoichiometry and/or mining or excreting nutrients. This understanding could be incorporated into individual-scale models by adjusting active metabolic processes based on environmental nutrient context, for instance through metabolic flux analysis. Increasing in scale towards trait-based models (e.g. Malik et al. 2020; Wang and Allison 2021), ecological stoichiometry operates by impacting the growth rates of organisms with particular traits – for example, microorganisms with specific nutrient acquisition pathways, r- and K- life strategies, stress tolerance, or other competitive advantages. Incorporation of stoichiometric principles such as substrate stoichiometry and/or environmental nutrient status at this scale can guide an individual’s expressed metabolisms in the context of other microorganisms, as well as the interactions of their metabolites and extracellular enzymes, and provides a more computationally favorable model structure. In turn, these omics-enabled representations can provide resolution into SOM cycling that currently aggregates microbial activity via lumped SOM pools and complement existing representations of mineral-sorption processes (Fig. 4).
Ecological stoichiometry can also aid in merging models from microbiology and biogeochemistry by limiting the need for detailed representations of SOM decomposition processes to specific metabolic pathways based on environmental context. In a bottom-up approach, ecological stoichiometry provides an overarching hypothesis and sets a priori expections for pathways of interest to assist in the interpretation of omics measurements (Fig. 4). Using either hypothesis-driven experiments or observational data from ecological networks, data mining efforts could focus on nutrient acquisition metabolic pathways (e.g. peptidase-, phosphatase-, and sulfatase-including metabolic pathways) in order of hypothesized nutrient limitations to allow for the compression of multi-omic data down to a manageable size. Data from multiple sources across ecoregions can be leveraged in this way to move towards transferrable principles of SOM cycling. Metabolic pathways with the most predictive power of SOM decomposition (inferred from transcripts, proteins, and metabolites mapping to metabolic pathways) then guide the portions of intermediate-scale models in which greater resolution of these pathways could lead to more accurate predictions. This could result in inclusion and/or calibration of specific portions of SOM pools that are most tightly coupled to decomposition rates (e.g., protein or necromass pools) or of specific metabolic pathways instead of lower resolution measurements such as CUE and NUE. Dynamically activating omics-enabled reaction networks for a specific nutrient status could provide the maximum benefit of high molecular resolution while being computationally tractable. In turn, parameters and rate estimates made at the intermediate scale can then influence the relative importance of variables in low molecular resolution models such as CUE/NUE, bacterial:fungal biomass, and/or biomass C:N:S:P.
In parallel, top-down approaches use coarsest measurement scale (e.g. bulk biogeochemistry) to first identify nutrient concentrations that then guide model development at higher molecular resolutions. Patterns in nutrient concentrations and pool sizes can inform the selection of limited high-resolution parameters for inclusion in reduced complexity models thereby optimizing added value from molecular techniques. For instance, soil nutrient status at a given point in time can be inferred from nutrient amendment experiments and bulk measurements of soil biogeochemistry, minerology, and major compound classes in SOM, and subsequently used to define a set of reactions that should most strongly impact rates of SOM decomposition. Examples might include representing organic N cycles (or amino acid metabolisms therein) for N limitation, microbial biomass recycling (necromass decomposition) for P limitation, and/or organosulfatases for S limitation—or combinations therein in the case of nutrient co-limitation—in omics-enabled reaction networks (Fig. 4). Outcomes from models with enhanced representation of specific SOM decomposition pathways can then further point to specific microbial genes/transcripts/proteins in pathways associated with the rate limiting step in SOM decomposition (if biological). Using this approach, we can narrow down molecular data to specific microbial metabolisms or omics markers of interest to build omics-enabled models including only the most relevant parameters that are scalable to the level of soil systems. This approach reduces the computational cost of high resolution reaction networks by only representing portions of a soil system that most strongly impact SOM decomposition in detail.
When used in combination, the result of these iterative approaches is a new suite of models that represent a limited set of microbial metabolisms, guided by stoichiometric principles and high-resolution molecular measurements, that are computationally feasible and more accurately predict SOM decomposition, termed ‘omics-enabled biogeochemical models’ (Fig. 4). Such models address a major challenge to the next generation of predictive models––deriving omics-enabled reaction networks that can improve assessments of soil C storage and emissions across molecular-to-ecosystem scales.
Opportunities afforded by investments in new molecular and computational technologies
The power of ecological stoichiometry to guide model development across scales of biological complexity can increase even further when leveraged in combination with new advances in molecular technologies and machine learning algorithms. Research that simultaneously investigates microbial roles in C, N, S, and P cycling is rare, and enzyme activity assays are imprecise relative to omics or probe-based assays. Investments in untargeted molecular approaches are generating new SOM cycling analyses by allowing detection of active microbial metabolisms. These are nascent technologies that have the potential to improve our understanding of SOM decomposition but need greater usage and development to surmount remaining challenges.
We especially highlight stable isotope probing (SIP; Bernard et al. 2007; Dumont and Murrell 2005; Pepe-Ranney et al. 2016), new ultrahigh-resolution metabolomics (Tfaily et al. 2017), activity-based protein profiling (ABPP, Killinger et al. 2019), and machine learning as new classes of technologies deserving of further development and broader consideration. Briefly, microbial applications of SIP use substrates enriched in heavy isotopes (e.g. 13 C, 18O, 15 N) amended to microbial communities to reveal isotope-labeled biomarkers of active microbial populations (Bernard et al. 2007; Dumont and Murrell 2005; Pepe-Ranney et al. 2016). While SIP has long been used in biogeochemistry, recent advances are increasing the tractability of SIP within amplicon profiles, metagenomes, metatranscriptomes, and metaproteomes (Barnett and Buckley 2020; Jameson et al. 2017; Pepe-Ranney et al. 2016; Wilhelm et al. 2019; Youngblut et al. 2018a, b). In parallelel, ultrahigh-resolution metabolomics, such as Fourier transform ion cyclotron resonance mass spectrometry (FTICR-MS, Tfaily et al. 2015; Tfaily et al. 2017), can illuminate new aspects of SOM chemistry by enabling the determination of elemental stoichiometry in thousands of SOM molecules simultaneously (Tfaily et al. 2017). Formulas assigned via ultrahigh-resolution metabolomics are a promising tool for evaluating metabolisms influenced by ecological stoichiometry by providing information on molecules with specific stoichiometries (e.g. C:N ratios of SOM molecules in a standing pool) or changes in these molecules during the course of experiments. For instance, Zhao et al. (2020) recently used FTICR-MS to evaluate SOM chemistries across soils with differing mineralogy and nutrient status. Finally, new targeted approaches can improve the accuracy of existing enzyme-based approaches. Activity-based protein profiling (ABPP, Killinger et al. 2019), for example, leverages chemical probes that react irreversibly with protein families to identify enzymes binding to specific molecules, and when paired with mass spectrometry-based approaches, can also quantify rates of enzyme production (Killinger et al. 2019). In contrast to enzyme activity assays that reveal potential enzyme rates, ABPP has the potential to dramatically improve investigations of SOM decomposition by revealing the true expression of decomposition enzymes; however, it has never been used in soils (Sadler and Wright 2015; Whidbey and Wright 2018; Zegeye et al. 2020).
Even with technological advances, the interpretation of untargeted molecular approaches is complicated by the inadequate description of metabolic pathways for soil microorganisms in existing databases and the unsuitability of standard statistical approaches in ecology for combining multidimensional data types. Unsupervised machine learning is emerging as a tool both to decipher hidden patterns in complex data and to eliminate the needs for a priori relationships between genes (e.g. via metabolic pathways or metagenome-assembled genomes) and for gap filling to infer absent data. There is an enormous diversity of machine learning algorithms with varying levels of complexity that are well-suited to discern patterns in data (Rana et al. 2020; Were et al. 2015). Techniques such as K-nearest-neighbor (Peterson 2009), artificial neural network (ANN, Sarle 1994), support vector machines (SVM, Wang 2005), neuro-fuzzy (Nauck et al. 1997), decision tree classifiers (Safavian and Landgrebe 1991), and random forests (Liaw and Wiener 2002) are most commonly employed in microbiology and environmental disciplines (e.g. Cai et al. 2019; Clercq et al. 2019; Dong and Chen 2019; Qdais et al. 2010; Rahimian Boogar et al. 2019; Thompson et al. 2019; Wang et al. 2020b). Still, we lack rigorous machine learning investigations in which sampling and experimental designs move beyond feature identification to translating selected features into meaningful ecological outcomes that acknowledge the environmental complexities of soils. Pairing machine learning with ecological theory for experimental hypothesis testing is needed to push fundamental biology forward (Rana et al. 2020; Were et al. 2015). Using new high-resolution molecular data types in spatially- and temporally-distributed sampling, paired with machine learning, is the gold standard for future research in SOM decomposition and can offer unprecedented untargeted resolution into the pathways by which microorganisms decompose SOM (Manzoni et al. 2017; Mooshammer et al. 2014b).
Conclusions
We have made significant advances in understanding global C cycles and now face the challenge of merging concepts and modelling frameworks from different scientific fields. To facilitate this effort, we identify substrate chemistry as a linkage point between microbiology and biogeochemistry, and we propose using ecological stoichiometry as a unifying framework that can help overcome a major limtation to understanding global SOM decomposition—intrepreting the massive amount of data generated by molecular approaches and translating them to the ecosystem scale. Our hope is that this common foundation can maximize the strengths of models across scales and environmental contexts, and we encourage better communication among empiricists and modelers within and across domains. Investing in and deploying new technologies in the context of stoichiometric theory provides an untapped and promising avenue for interpreting the vast amount of data we can now generate and distilling this information into a new generation of omics-enabled biogeochemical models.
Data availability
Not applicable.
Code availability
Not applicable.
References
Abramoff R, Xu X, Hartman M, O’Brien S, Feng W, Davidson E, Finzi A, Moorhead D, Schimel J, Torn M (2018) The Millennial model: in search of measurable pools and transformations for modeling soil carbon in the new century. Biogeochemistry 137(1–2):51–71
Allison SD, Gartner TB, Holland K, Weintraub M, Sinsabaugh RL (2007) Soil enzymes: linking proteomics and ecological processes. In: Allison SD, Gartner TB, Holland K, Weintraub M, Sinsabaugh RL (eds) Manual of Environmental Microbiology, 3rd edn. American Society of Microbiology, Washington, pp 704–711
Allison SD, Weintraub MN, Gartner TB, Waldrop MP (2010) Evolutionary-economic principles as regulators of soil enzyme production and ecosystem function. In: Allison SD, Weintraub MN, Gartner TB, Waldrop MP (eds) Soil enzymology. Springer, New York, pp 229–243
Allison SD, Chacon SS, German DP (2014) Substrate concentration constraints on microbial decomposition. Soil Biol Biochem 79:43–49
Anderson TR, Hessen DO, Elser JJ, Urabe J (2005) Metabolic stoichiometry and the fate of excess carbon and nutrients in consumers. Am Nat 165(1):1–15
Azizi-Rad M, Chanca I, Herrera‐Ramírez D, Metzler H, Sierra CA (2021) Stochastic and deterministic interpretation of pool models. Glob Change Biol 27(11):2271–2272
Bailey VL, Bond-Lamberty B, DeAngelis K, Grandy AS, Hawkes CV, Heckman K, Lajtha K, Phillips RP, Sulman BN, Todd‐Brown KE (2018) Soil carbon cycling proxies: understanding their critical role in predicting climate change feedbacks. Glob Change Biol 24(3):895–905
Ballantyne IVF, Billings SA (2018) Model formulation of microbial CO2 production and efficiency can significantly influence short and long term soil C projections. ISME J 12(6):1395–1403
Barnett SE, Buckley DH (2020) Simulating metagenomic stable isotope probing datasets with MetaSIPSim. BMC Bioinform 21(1):37
Batjes NH (2016) Harmonized soil property values for broad-scale modelling (WISE30sec) with estimates of global soil carbon stocks. Geoderma 269:61–68
Berardi D, Brzostek E, Blanc-Betes E, Davison B, DeLucia EH, Hartman MD, Kent J, Parton WJ, Saha D, Hudiburg TW (2020) 21st‐century biogeochemical modeling: challenges for century‐based models and where do we go from here? GCB Bioenergy 12(10):774–788
Bernard L, Mougel C, Maron PA, Nowak V, Lévêque J, Henault C, Haichar FeZ, Berge O, Marol C, Balesdent J (2007) Dynamics and identification of soil microbial populations actively assimilating carbon from 13 C-labelled wheat residue as estimated by DNA‐and RNA‐SIP techniques. Environ Microbiol 9(3):752–764
Billings SA, Ballantyne IVF (2013) How interactions between microbial resource demands, soil organic matter stoichiometry, and substrate reactivity determine the direction and magnitude of soil respiratory responses to warming. Glob Change Biol 19(1):90–102
Billings SA, Ziegler SE (2008) Altered patterns of soil carbon substrate usage and heterotrophic respiration in a pine forest with elevated CO2 and N fertilization. Glob Change Biol 14(5):1025–1036
Billings SA, Lajtha K, Malhotra A, Berhe AA, de Graaff MA, Earl S, Fraterrigo J, Georgiou K, Grandy S, Hobbie SE (2021) Soil organic carbon is not just for soil scientists: measurement recommendations for diverse practitioners. Ecol Appl 31(3):e02290
Blankinship JC, Berhe AA, Crow SE, Druhan JL, Heckman KA, Keiluweit M, Lawrence CR, Marín-Spiotta E, Plante AF, Rasmussen C (2018) Improving understanding of soil organic matter dynamics by triangulating theories, measurements, and models. Biogeochemistry 140(1):1–13
Bond-Lamberty B, Epron D, Harden J, Harmon ME, Hoffman F, Kumar J, David McGuire A, Vargas R (2016) Estimating heterotrophic respiration at large scales: challenges, approaches, and next steps. Ecosphere 7(6):e01380
Borer B, Or D (2021) Spatiotemporal metabolic modeling of bacterial life in complex habitats. Curr Opin Biotechnol 67:65–71
Borer B, Ataman M, Hatzimanikatis V, Or D (2019) Modeling metabolic networks of individual bacterial agents in heterogeneous and dynamic soil habitats (IndiMeSH). PLoS Comput Biol 15(6):e1007127
Bouskill N, Tang J, Riley WJ, Brodie EL (2012) Trait-based representation of biological nitrification: model development, testing, and predicted community composition. Front Microbiol 3:364
Buchkowski RW, Schmitz OJ, Bradford MA (2015) Microbial stoichiometry overrides biomass as a regulator of soil carbon and nitrogen cycling. Ecology 96(4):1139–1149
Buchkowski RW, Shaw AN, Sihi D, Smith GR, Keiser AD (2019) Constraining carbon and nutrient flows in soil with ecological stoichiometry. Front Ecol Evol 7:382
Burns RG (1982) Enzyme activity in soil: location and a possible role in microbial ecology. Soil Biol Biochem 14(5):423–427
Cai W, Lesnik KL, Wade MJ, Heidrich ES, Wang Y, Liu H (2019) Incorporating microbial community data with machine learning techniques to predict feed substrates in microbial fuel cells. Biosens Bioelectron 133:64–71
Cheng C-H, Lehmann J, Thies JE, Burton SD, Engelhard MH (2006) Oxidation of black carbon by biotic and abiotic processes. Org Geochem 37(11):1477–1488
Chowdhury TR, Lee J-Y, Bottos EM, Brislawn CJ, White RA, Bramer LM, Brown J, Zucker JD, Kim Y-M, Jumpponen A (2019) Metaphenomic responses of a native prairie soil microbiome to moisture perturbations. Msystems. https://doi.org/10.1128/mSystems.00061-19
Cleveland CC, Liptzin D (2007) C: N: P stoichiometry in soil: is there a “Redfield ratio” for the microbial biomass? Biogeochemistry 85(3):235–252
Cleveland CC, Reed SC, Townsend AR (2006) Nutrient regulation of organic matter decomposition in a tropical rain forest. Ecology 87(2):492–503
Cotrufo MF, Wallenstein MD, Boot CM, Denef K, Paul E (2013) The Microbial Efficiency-Matrix Stabilization (MEMS) framework integrates plant litter decomposition with soil organic matter stabilization: do labile plant inputs form stable soil organic matter? Global Change Biol 19(4):988–995
Craig ME, Mayes MA, Sulman BN, Walker AP (2021) Biological mechanisms may contribute to soil carbon saturation patterns. Glob Change Biol 27(12):2633–2644
Crowther TW, Van den Hoogen J, Wan J, Mayes MA, Keiser A, Mo L, Averill C, Maynard DS (2019) The global soil community and its influence on biogeochemistry. Science 365(6455):eaav0550
Cuevas DA, Edirisinghe J, Henry CS, Overbeek R, O’Connell TG, Edwards RA (2016) From DNA to FBA: how to build your own genome-scale metabolic model. Front Microbiol 7:907
Dangal SR, Sanderman J, Wills S, Ramirez-Lopez L (2019) Accurate and precise prediction of soil properties from a large mid-infrared spectral library. Soil Syst 3(1):11
De Clercq D, Jalota D, Shang R, Ni K, Zhang Z, Khan A, Wen Z, Caicedo L, Yuan K (2019) Machine learning powered software for accurate prediction of biogas production: A case study on industrial-scale Chinese production data. J Clean Prod 218:390–399
Del Giorgio PA, Cole JJ (1998) Bacterial growth efficiency in natural aquatic systems. Annu Rev Ecol Syst 29(1):503–541
Dong C, Chen J (2019) Optimization of process parameters for anaerobic fermentation of corn stalk based on least squares support vector machine. Bioresour Technol 271:174–181
Duarte NC, Herrgård MJ, Palsson B (2004) Reconstruction and validation of Saccharomyces cerevisiae iND750, a fully compartmentalized genome-scale metabolic model. Genome Res 14(7):1298–1309
Dumont MG, Murrell JC (2005) Stable isotope probing—linking microbial identity to function. Nat Rev Microbiol 3(6):499–504
Dungait JA, Hopkins DW, Gregory AS, Whitmore AP (2012) Soil organic matter turnover is governed by accessibility not recalcitrance. Glob Change Biol 18(6):1781–1796
Dwivedi D, Tang J, Bouskill N, Georgiou K, Chacon SS, Riley WJ (2019) Abiotic and biotic controls on soil organo–mineral interactions: developing model structures to analyze why soil organic matter persists. Rev Mineral Geochem 85(1):329–348
Elser JJ, Dobberfuhl DR, MacKay NA, Schampel JH (1996) Organism size, life history, and N:P stoichiometry: toward a unified view of cellular and ecosystem processes. Bioscience 46(9):674–684
Elser J, Sterner RW, Gorokhova Ea, Fagan W, Markow T, Cotner JB, Harrison J, Hobbie SE, Odell G, Weider L (2000) Biological stoichiometry from genes to ecosystems. Ecol Lett 3(6):540–550
Elser J, Acharya K, Kyle M, Cotner J, Makino W, Markow T, Watts T, Hobbie S, Fagan W, Schade J (2003) Growth rate–stoichiometry couplings in diverse biota. Ecol Lett 6(10):936–943
Fatichi S, Manzoni S, Or D, Paschalis A (2019) A Mechanistic model of microbially mediated soil biogeochemical processes: A reality check. Global Biogeochem Cycles 33(6):620–648
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88(6):1354–1364
Finn D, Yu J, Penton CR (2020) Soil quality shapes the composition of microbial community stress response and core cell metabolism functional genes. Appl Soil Ecol 148:103483
Follows MJ, Dutkiewicz S (2011) Modeling diverse communities of marine microbes. Annu Rev Mar Sci 3:427–451
Freney J (1986) Forms and reactions of organic sulfur compounds in soils. In: Tabatabai MA (ed) Sulfur in agriculture, vol 27. American Society of Agronomy Inc, Madison, pp 207–232
Frey SD, Lee J, Melillo JM, Six J (2013) The temperature response of soil microbial efficiency and its feedback to climate. Nat Clim Chang 3(4):395–398
Frost PC, Evans-White MA, Finkel ZV, Jensen TC, Matzek V (2005) Are you what you eat? Physiological constraints on organismal stoichiometry in an elementally imbalanced world. Oikos 109(1):18–28
Garayburu-Caruso VA, Stegen JC, Song H-S, Renteria L, Wells J, Garcia W, Resch CT, Goldman AE, Chu RK, Toyoda J (2020) Carbon limitation leads to thermodynamic regulation of aerobic metabolism. Environ Sci Technol Lett. https://doi.org/10.1021/acs.estlett.0c00258
Ge T, Luo Y, Singh BP (2020) Resource stoichiometric and fertility in soil. Biol Fertil Soils. https://doi.org/10.1007/s00374-020-01513-5
Geyer KM, Kyker-Snowman E, Grandy AS, Frey SD (2016) Microbial carbon use efficiency: accounting for population, community, and ecosystem-scale controls over the fate of metabolized organic matter. Biogeochemistry 127(2–3):173–188
Graham EB, Knelman JE, Schindlbacher A, Siciliano S, Breulmann M, Yannarell A, Beman J, Abell G, Philippot L, Prosser J (2016) Microbes as engines of ecosystem function: when does community structure enhance predictions of ecosystem processes? Front Microbiol 7:214
Grandy AS, Sinsabaugh RL, Neff JC, Stursova M, Zak DR (2008) Nitrogen deposition effects on soil organic matter chemistry are linked to variation in enzymes, ecosystems and size fractions. Biogeochemistry 91(1):37–49
van Groenigen K-J, Six J, Hungate BA, de Graaff M-A, Van Breemen N, Van Kessel C (2006) Element interactions limit soil carbon storage. Proc Natl Acad Sci 103(17):6571–6574
Guo X, Gao Q, Yuan M, Wang G, Zhou X, Feng J, Shi Z, Hale L, Wu L, Zhou A (2020) Gene-informed decomposition model predicts lower soil carbon loss due to persistent microbial adaptation to warming. Nat Commun 11(1):1–12
Hall EK, Bernhardt ES, Bier RL, Bradford MA, Boot CM, Cotner JB, del Giorgio PA, Evans SE, Graham EB, Jones SE (2018) Understanding how microbiomes influence the systems they inhabit. Nat Microbiol 3(9):977–982
Heinken A, Basile A, Thiele I (2021) Advances in constraint-based modelling of microbial communities. Curr Opin Syst Biol. https://doi.org/10.1016/j.coisb.2021.05.007
Henry CS, Bernstein HC, Weisenhorn P, Taylor RC, Lee JY, Zucker J, Song HS (2016) Microbial community metabolic modeling: a community data-driven network reconstruction. J Cell Physiol 231(11):2339–2345
Herron PM, Stark JM, Holt C, Hooker T, Cardon ZG (2009) Microbial growth efficiencies across a soil moisture gradient assessed using 13 C-acetic acid vapor and 15 N-ammonia gas. Soil Biol Biochem 41(6):1262–1269
Heuck C, Weig A, Spohn M (2015) Soil microbial biomass C: N: P stoichiometry and microbial use of organic phosphorus. Soil Biol Biochem 85:119–129
Hill BH, Elonen CM, Jicha TM, Kolka RK, Lehto LL, Sebestyen SD, Seifert-Monson LR (2014) Ecoenzymatic stoichiometry and microbial processing of organic matter in northern bogs and fens reveals a common P-limitation between peatland types. Biogeochemistry 120(1–3):203–224
Hunter KS, Wang Y, Van Cappellen P (1998) Kinetic modeling of microbially-driven redox chemistry of subsurface environments: coupling transport, microbial metabolism and geochemistry. J Hydrol 209(1–4):53–80
Jameson E, Taubert M, Coyotzi S, Chen Y, Eyice Ö, Schäfer H, Murrell JC, Neufeld JD, Dumont MG (2017) DNA-, RNA-, and protein-based stable-isotope probing for high-throughput biomarker analysis of active microorganisms. In: Jameson E, Taubert M, Coyotzi S, Chen Y, Eyice Ö, Schäfer H, Murrell JC, Neufeld JD, Dumont MG (eds) Metagenomics. Springer, New York, pp p57-74
Jansson JK, Hofmockel KS (2018) The soil microbiome—from metagenomics to metaphenomics. Curr Opin Microbiol 43:162–168
Jenkinson DS (1990) The turnover of organic carbon and nitrogen in soil. Philos Trans R Soc Lond B Biol Sci 329(1255):361–368
Jilling A, Keiluweit M, Contosta AR, Frey S, Schimel J, Schnecker J, Smith RG, Tiemann L, Grandy AS (2018) Minerals in the rhizosphere: overlooked mediators of soil nitrogen availability to plants and microbes. Biogeochemistry 139(2):103–122
Jing X, Chen X, Fang J, Ji C, Shen H, Zheng C, Zhu B (2020) Soil microbial carbon and nutrient constraints are driven more by climate and soil physicochemical properties than by nutrient addition in forest ecosystems. Soil Biol Biochem 141:107657
Kaiser M, Ellerbrock R (2005) Functional characterization of soil organic matter fractions different in solubility originating from a long-term field experiment. Geoderma 127(3–4):196–206
Kallenbach C, Grandy AS, Frey S, Diefendorf A (2015) Microbial physiology and necromass regulate agricultural soil carbon accumulation. Soil Biol Biochem 91:279–290
Kallenbach CM, Frey SD, Grandy AS (2016) Direct evidence for microbial-derived soil organic matter formation and its ecophysiological controls. Nat Commun 7(1):1–10
Keiluweit M, Nico PS, Johnson MG, Kleber M (2010) Dynamic molecular structure of plant biomass-derived black carbon (biochar). Environ Sci Technol 44(4):1247–1253
Kessell AK, McCullough HC, Auchtung JM, Bernstein HC, Song H-S (2020) Predictive interactome modeling for precision microbiome engineering. Curr Opin Chem Eng 30:77–85
Khan KS, Mack R, Castillo X, Kaiser M, Joergensen RG (2016) Microbial biomass, fungal and bacterial residues, and their relationships to the soil organic matter C/N/P/S ratios. Geoderma 271:115–123
Killinger BJ, Brandvold KR, Ramos-Hunter SJ, Wright AT (2019) Chemoproteomic analyses by activity-based protein profiling. In: Tao WA, Zhang Y (eds) Mass Spectrometry-Based Chemical Proteomics. Wiley, Hoboken, pp 67–99
Kirkby C, Kirkegaard J, Richardson A, Wade L, Blanchard C, Batten G (2011) Stable soil organic matter: a comparison of C: N: P: S ratios in Australian and other world soils. Geoderma 163(3–4):197–208
Kirkby CA, Richardson AE, Wade LJ, Batten GD, Blanchard C, Kirkegaard JA (2013) Carbon-nutrient stoichiometry to increase soil carbon sequestration. Soil Biol Biochem 60:77–86
Kirkby CA, Richardson AE, Wade LJ, Passioura JB, Batten GD, Blanchard C, Kirkegaard JA (2014) Nutrient availability limits carbon sequestration in arable soils. Soil Biol Biochem 68:402–409
Kuzyakov Y, Mason-Jones K (2018) Viruses in soil: Nano-scale undead drivers of microbial life, biogeochemical turnover and ecosystem functions. Soil Biol Biochem 127:305–317
Kyker-Snowman E, Wieder WR, Frey SD, Grandy AS (2020) Stoichiometrically coupled carbon and nitrogen cycling in the microbial-mineral carbon stabilization model version 1.0 (MIMICS-CN v1. 0). Geosci Model Dev 13(9):4413–4434
Lehmann J, Kleber M (2015) The contentious nature of soil organic matter. Nature 528(7580):60–68
Leue M, Ellerbrock RH, Gerke HH (2010) DRIFT mapping of organic matter composition at intact soil aggregate surfaces. Vadose Zone J 9(2):317–324
Li J, Wang G, Allison SD, Mayes MA, Luo Y (2014) Soil carbon sensitivity to temperature and carbon use efficiency compared across microbial-ecosystem models of varying complexity. Biogeochemistry 119(1–3):67–84
Liang C, Amelung W, Lehmann J, Kästner M (2019) Quantitative assessment of microbial necromass contribution to soil organic matter. Glob change Biol 25(11):3578–3590
Liaw A, Wiener M (2002) Classification and regression by randomForest. R News 2(3):18–22
Lieven C, Beber ME, Olivier BG, Bergmann FT, Ataman M, Babaei P, Bartell JA, Blank LM, Chauhan S, Correia K (2020) MEMOTE for standardized genome-scale metabolic model testing. Nat Biotechnol 38(3):272–276
Loira N, Dulermo T, Nicaud J-M, Sherman DJ (2012) A genome-scale metabolic model of the lipid-accumulating yeast Yarrowia lipolytica. BMC Syst Biol 6(1):1–9
Lu H, Li F, Sánchez BJ, Zhu Z, Li G, Domenzain I, Marcišauskas S, Anton PM, Lappa D, Lieven C (2019) A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism. Nat Commun 10(1):1–13
Malik AA, Puissant J, Buckeridge KM, Goodall T, Jehmlich N, Chowdhury S, Gweon HS, Peyton JM, Mason KE, van Agtmaal M (2018) Land use driven change in soil pH affects microbial carbon cycling processes. Nat Commun 9(1):1–10
Malik AA, Martiny JB, Brodie EL, Martiny AC, Treseder KK, Allison SD (2020) Defining trait-based microbial strategies with consequences for soil carbon cycling under climate change. ISME J 14(1):1–9
Manzoni S (2017) Flexible carbon-use efficiency across litter types and during decomposition partly compensates nutrient imbalances—results from analytical stoichiometric models. Front Microbiol 8:661
Manzoni S, Jackson RB, Trofymow JA, Porporato A (2008) The global stoichiometry of litter nitrogen mineralization. Science 321(5889):684–686
Manzoni S, Taylor P, Richter A, Porporato A, Ågren GI (2012) Environmental and stoichiometric controls on microbial carbon-use efficiency in soils. New Phytol 196(1):79–91
Manzoni S, Čapek P, Mooshammer M, Lindahl BD, Richter A, Šantrůčková H (2017) Optimal metabolic regulation along resource stoichiometry gradients. Ecol Lett 20(9):1182–1191
Margalef O, Sardans J, Fernández-Martínez M, Molowny-Horas R, Janssens I, Ciais P, Goll D, Richter A, Obersteiner M, Asensio D (2017) Global patterns of phosphatase activity in natural soils. Sci Rep 7(1):1–13
McClure RS, Lee J-Y, Chowdhury TR, Bottos EM, White RA, Kim Y-M, Nicora CD, Metz TO, Hofmockel KS, Jansson JK (2020) Integrated network modeling approach defines key metabolic responses of soil microbiomes to perturbations. Sci Rep 10(1):1–9
Middelboe M, Søndergaard M (1993) Bacterioplankton growth yield: seasonal variations and coupling to substrate lability and β-glucosidase activity. Appl Environ Microbiol 59(11):3916–3921
Milcu A, Heim A, Ellis RJ, Scheu S, Manning P (2011) Identification of general patterns of nutrient and labile carbon control on soil carbon dynamics across a successional gradient. Ecosystems 14(5):710–719
Min K, Lehmeier CA, Billings SA (2016) Carbon availability modifies temperature responses of heterotrophic microbial respiration, carbon uptake affinity, and stable carbon isotope discrimination. Front Microbiol. https://doi.org/10.3389/fmicb.2016.02083
Moorhead DL, Lashermes G, Sinsabaugh RL (2012) A theoretical model of C-and N-acquiring exoenzyme activities, which balances microbial demands during decomposition. Soil Biol Biochem 53:133–141
Moorhead DL, Rinkes ZL, Sinsabaugh RL, Weintraub MN (2013) Dynamic relationships between microbial biomass, respiration, inorganic nutrients and enzyme activities: informing enzyme-based decomposition models. Front Microbiol 4:223
Mooshammer M, Wanek W, Hämmerle I, Fuchslueger L, Hofhansl F, Knoltsch A, Schnecker J, Takriti M, Watzka M, Wild B (2014) Adjustment of microbial nitrogen use efficiency to carbon: nitrogen imbalances regulates soil nitrogen cycling. Nat Commun 5:3694
Mooshammer M, Wanek W, Zechmeister-Boltenstern S, Richter AA (2014) Stoichiometric imbalances between terrestrial decomposer communities and their resources: mechanisms and implications of microbial adaptations to their resources. Front Microbiol 5:22
Müller T, Höper H (2004) Soil organic matter turnover as a function of the soil clay content: consequences for model applications. Soil Biol Biochem 36(6):877–888
Nannipieri P, Trasar-Cepeda C, Dick RP (2018) Soil enzyme activity: a brief history and biochemistry as a basis for appropriate interpretations and meta-analysis. Biol Fertil Soils 54(1):11–19
Nauck D, Klawonn F, Kruse R (1997) Foundations of neuro-fuzzy systems. Wiley, Hoboken
Naylor D, Sadler N, Bhattacharjee A, Graham EB, Anderton CR, McClure R, Lipton M, Hofmockel KS, Jansson JK (2020) Soil microbiomes under climate change and implications for carbon cycling. Annu Rev Environ Resour 45(1):29–59
Orth JD, Thiele I, Palsson B (2010) What is flux balance analysis? Nat Biotechnol 28(3):245–248
Parton W (1996) The CENTURY model. In: Parton W (ed) Evaluation of soil organic matter models. Springer, New York, pp 283–291
Patel KF, Fansler SJ, Campbell TP, Bond-Lamberty B, Smith AP, RoyChowdhury T, McCue LA, Varga T, Bailey VL (2021) Soil texture and environmental conditions influence the biogeochemical responses of soils to drought and flooding. Commun Earth Environ 2(1):1–9
Pepe-Ranney C, Campbell AN, Koechli CN, Berthrong S, Buckley DH (2016) Unearthing the ecology of soil microorganisms using a high resolution DNA-SIP approach to explore cellulose and xylose metabolism in soil. Front Microbiol 7:703
Peterson LE (2009) K-nearest neighbor. Scholarpedia 4(2):1883
Porta G, la Cecilia D, Guadagnini A, Maggi F (2018) Implications of uncertain bioreactive parameters on a complex reaction network of atrazine biodegradation in soil. Adv Water Resour 121:263–276
Qdais HA, Hani KB, Shatnawi N (2010) Modeling and optimization of biogas production from a waste digester using artificial neural network and genetic algorithm. Resour Conserv Recycl 54(6):359–363
Qiao Y, Wang J, Liang G, Du Z, Zhou J, Zhu C, Huang K, Zhou X, Luo Y, Yan L (2019) Global variation of soil microbial carbon-use efficiency in relation to growth temperature and substrate supply. Sci Rep 9(1):1–8
Rahimian Boogar A, Salehi H, Pourghasemi HR, Blaschke T (2019) Predicting habitat suitability and conserving juniperus spp. water, habitat using SVM and maximum entropy machine learning techniques. Water. https://doi.org/10.3390/w11102049
Rana P, Berry C, Ghosh P, Fong SS (2020) Recent advances on constraint-based models by integrating machine learning. Curr Opin Biotechnol 64:85–91
Redfield AC (1958) The biological control of chemical factors in the environment. Am Sci 46(3):230A–221
Reed DC, Algar CK, Huber JA, Dick GJ (2014) Gene-centric approach to integrating environmental genomics and biogeochemical models. Proc Natl Acad Sci 111(5):1879–1884
Riley W, Maggi F, Kleber M, Torn M, Tang J, Dwivedi D, Guerry N (2014) Long residence times of rapidly decomposable soil organic matter: application of a multi-phase, multi-component, and vertically resolved model (BAMS1) to soil carbon dynamics. Geosci Model Dev 7(4):1335–1355
Robertson AD, Paustian K, Ogle S, Wallenstein MD, Lugato E, Cotrufo MF (2019) Unifying soil organic matter formation and persistence frameworks: the MEMS model. Biogeosciences 16(6):1225–1248
Rumpel C, Kögel-Knabner I (2011) Deep soil organic matter—a key but poorly understood component of terrestrial C cycle. Plant soil 338(1):143–158
Sadler NC, Wright AT (2015) Activity-based protein profiling of microbes. Curr Opin Chem Biol 24:139–144
Safavian SR, Landgrebe D (1991) A survey of decision tree classifier methodology. IEEE Trans Syst Man Cybern 21(3):660–674
Saifuddin M, Bhatnagar JM, Segrè D, Finzi AC (2019) Microbial carbon use efficiency predicted from genome-scale metabolic models. Nat Commun 10(1):1–10
Sarle WS (1994) Neural networks and statistical models
Schimel J (2021) The Democracy of dirt: relating micro-scale dynamics to macro-scale ecosystem function. In: Hurst CJ (ed) The Foundation Stone of the Biosphere Microbes. Springer, Cham, pp 89–102
Schimel J, Becerra CA, Blankinship J (2017) Estimating decay dynamics for enzyme activities in soils from different ecosystems. Soil Biol Biochem 114:5–11
Seaver SM, Liu F, Zhang Q, Jeffryes J, Faria JP, Edirisinghe J, Mundy M, Chia N, Noor E, Beber ME (2020) The ModelSEED Database for the integration of metabolic annotations and the reconstruction, comparison, and analysis of metabolic models for plants, fungi, and microbes. Nucleic Acids Res. https://doi.org/10.1093/nar/gkaa746
Sierra CA, Müller M (2015) A general mathematical framework for representing soil organic matter dynamics. Ecol Monogr 85(4):505–524
Sihi D, Gerber S, Inglett PW, Inglett KS (2016) Comparing models of microbial–substrate interactions and their response to warming. Biogeosciences 13(6):1733–1752
Sinsabaugh RL, Shah JJF (2011) Ecoenzymatic stoichiometry of recalcitrant organic matter decomposition: the growth rate hypothesis in reverse. Biogeochemistry 102(1–3):31–43
Sinsabaugh RL, Lauber CL, Weintraub MN, Ahmed B, Allison SD, Crenshaw C, Contosta AR, Cusack D, Frey S, Gallo ME (2008) Stoichiometry of soil enzyme activity at global scale. Ecol Lett 11(11):1252–1264
Sinsabaugh RL, Manzoni S, Moorhead DL, Richter A (2013) Carbon use efficiency of microbial communities: stoichiometry, methodology and modelling. Ecol Lett 16(7):930–939
Sinsabaugh RL, Turner BL, Talbot JM, Waring BG, Powers JS, Kuske CR, Moorhead DL, Follstad Shah JJ (2016) Stoichiometry of microbial carbon use efficiency in soils. Ecol Monogr 86(2):172–189
Six J, Conant RT, Paul EA, Paustian K (2002) Stabilization mechanisms of soil organic matter: implications for C-saturation of soils. Plant Soil 241(2):155–176
Six J, Frey S, Thiet R, Batten K (2006) Bacterial and fungal contributions to carbon sequestration in agroecosystems. Soil Sci Soc Am J 70(2):555–569
Smith EM, Prairie YT (2004) Bacterial metabolism and growth efficiency in lakes: the importance of phosphorus availability. Limnol Oceanogr 49(1):137–147
Song H-S, Stegen JC, Graham EB, Lee J-Y, Garayburu-Caruso V, Nelson WC, Chen X, Moulton JD, Scheibe TD (2020) Representing organic matter thermodynamics in biogeochemical reactions via substrate-explicit modeling. Front Microbiol. https://doi.org/10.3389/fmicb.2020.531756
Spohn M (2016) Element cycling as driven by stoichiometric homeostasis of soil microorganisms. Basic Appl Ecol 17(6):471–478
Sterner RW (1995) Elemental stoichiometry of species in ecosystems. In: Sterner RW (ed) Linking species & ecosystems. Springer, New York, pp p240-252
Sterner RW, Elser JJ (2002) Ecological stoichiometry: the biology of elements from molecules to the biosphere. Princeton University Press, Princeton
Stewart CE, Paustian K, Conant RT, Plante AF, Six J (2007) Soil carbon saturation: concept, evidence and evaluation. Biogeochemistry 86(1):19–31
Stone BW, Li J, Koch BJ, Blazewicz SJ, Dijkstra P, Hayer M, Hofmockel KS, Liu X-JA, Mau RL, Morrissey EM (2021) Nutrients cause consolidation of soil carbon flux to small proportion of bacterial community. Nat Commun 12(1):1–9
Strickland MS, Rousk J (2010) Considering fungal: bacterial dominance in soils–methods, controls, and ecosystem implications. Soil Biol Biochem 42(9):1385–1395
Sullivan BW, Alvarez-Clare S, Castle SC, Porder S, Reed SC, Schreeg L, Townsend AR, Cleveland CC (2014) Assessing nutrient limitation in complex forested ecosystems: alternatives to large-scale fertilization experiments. Ecology 95(3):668–681
Sulman BN, Phillips RP, Oishi AC, Shevliakova E, Pacala SW (2014) Microbe-driven turnover offsets mineral-mediated storage of soil carbon under elevated CO 2. Nat Clim Chang 4(12):1099–1102
Sulman BN, Shevliakova E, Brzostek ER, Kivlin SN, Malyshev S, Menge DN, Zhang X (2019) Diverse mycorrhizal associations enhance terrestrial C storage in a global model. Global Biogeochem Cycles 33(4):501–523
Takriti M, Wild B, Schnecker J, Mooshammer M, Knoltsch A, Lashchinskiy N, Alves RJE, Gentsch N, Gittel A, Mikutta R (2018) Soil organic matter quality exerts a stronger control than stoichiometry on microbial substrate use efficiency along a latitudinal transect. Soil Biol Biochem 121:212–220
Tang J, Riley WJ (2015) Weaker soil carbon–climate feedbacks resulting from microbial and abiotic interactions. Nat Clim Chang 5(1):56–60
Tang J, Riley WJ (2020) Linear two-pool models are insufficient to infer soil organic matter decomposition temperature sensitivity from incubations. Biogeochemistry 149(3):251–261
Taylor PG, Townsend AR (2010) Stoichiometric control of organic carbon–nitrate relationships from soils to the sea. Nature 464(7292):1178–1181
Tfaily MM, Chu RK, Tolić N, Roscioli KM, Anderton CR, Paša-Tolić L, Robinson EW, Hess NJ (2015) Advanced solvent based methods for molecular characterization of soil organic matter by high-resolution mass spectrometry. Anal Chem 87(10):5206–5215
Tfaily MM, Chu RK, Toyoda J, Tolić N, Robinson EW, Paša-Tolić L, Hess NJ (2017) Sequential extraction protocol for organic matter from soils and sediments using high resolution mass spectrometry. Anal Chim Acta 972:54–61
Thompson J, Johansen R, Dunbar J, Munsky B (2019) Machine learning to predict microbial community functions: an analysis of dissolved organic carbon from litter decomposition. PloS One. https://doi.org/10.1371/journal.pone.0215502
Todd-Brown K, Randerson J, Post W, Hoffman F, Tarnocai C, Schuur E, Allison S (2013) Causes of variation in soil carbon simulations from CMIP5 Earth system models and comparison with observations. Biogeosciences 10(3):1717–1736
Todd-Brown K, Randerson J, Hopkins F, Arora V, Hajima T, Jones C, Shevliakova E, Tjiputra J, Volodin E, Wu T (2014) Changes in soil organic carbon storage predicted by Earth system models during the 21st century. Biogeosciences 11(8):2341–2356
Van de Waal DB, Elser JJ, Martiny AC, Sterner RW, Cotner JB (2018) Progress in ecological stoichiometry. Front Microbiol. https://doi.org/10.3389/fmicb.2018.01957
Wang L (2005) Support vector machines: theory and applications. Springer, New York
Wang B, Allison SD (2021) Drought legacies mediated by trait trade-offs in soil microbiomes. Ecosphere 12(6):e03562
Wang YP, Houlton BZ (2009) Nitrogen constraints on terrestrial carbon uptake: Implications for the global carbon-climate feedback. Geophys Res Lett. https://doi.org/10.1029/2009GL041009
Wang G, Jagadamma S, Mayes MA, Schadt CW, Steinweg JM, Gu L, Post WM (2015) Microbial dormancy improves development and experimental validation of ecosystem model. ISME J 9(1):226–237
Wang G, Huang W, Zhou G, Mayes MA, Zhou J (2020a) Modeling the processes of soil moisture in regulating microbial and carbon-nitrogen cycling. J Hydrol 585:124777
Wang L, Long F, Liao W, Liu H (2020b) Prediction of anaerobic digestion performance and identification of critical operational parameters using machine learning algorithms. Biores Technol 298:122495
Waring BG, Sulman BN, Reed S, Smith AP, Averill C, Creamer CA, Cusack DF, Hall SJ, Jastrow JD, Jilling A (2020) From pools to flow: The PROMISE framework for new insights on soil carbon cycling in a changing world. Glob Change Biol 26(12):6631–6643
Wei X, Zhu Z, Liu Y, Luo Y, Deng Y, Xu X, Liu S, Richter A, Shibistova O, Guggenberger G (2020) C: N: P stoichiometry regulates soil organic carbon mineralization and concomitant shifts in microbial community composition in paddy soil. Biol Fertil Soils. https://doi.org/10.1007/s00374-020-01468-7
Weinbauer MG (2004) Ecology of prokaryotic viruses. FEMS MicroBiol Rev 28(2):127–181
Were K, Bui DT, Dick ØB, Singh BR (2015) A comparative assessment of support vector regression, artificial neural networks, and random forests for predicting and mapping soil organic carbon stocks across an Afromontane landscape. Ecol Ind 52:394–403
Whidbey C, Wright AT (2018) Activity-based protein profiling—enabling multimodal functional studies of microbial communities. In: Whidbey C, Wright AT (eds) Activity-Based Protein Profiling. Springer, New York, pp p1-21
Wieder WR, Bonan GB, Allison SD (2013) Global soil carbon projections are improved by modelling microbial processes. Nat Clim Chang 3(10):909–912
Wieder W, Grandy A, Kallenbach C, Bonan G (2014) Integrating microbial physiology and physio-chemical principles in soils with the MIcrobial-MIneral Carbon Stabilization (MIMICS) model. Biogeosciences 11(14):3899–3917
Wieder WR, Cleveland CC, Smith WK, Todd-Brown K (2015) Future productivity and carbon storage limited by terrestrial nutrient availability. Nat Geosci 8(6):441–444
Wilhelm SW, Suttle CA (1999) Viruses and nutrient cycles in the sea: viruses play critical roles in the structure and function of aquatic food webs. Bioscience 49(10):781–788
Wilhelm RC, Singh R, Eltis LD, Mohn WW (2019) Bacterial contributions to delignification and lignocellulose degradation in forest soils with metagenomic and quantitative stable isotope probing. ISME J 13(2):413–429
Wilson RM, Neumann RB, Crossen KB, Raab NM, Hodgkins SB, Saleska SR, Bolduc B, Woodcroft BJ, Tyson GW, Chanton JP (2019) Microbial community analyses inform geochemical reaction network models for predicting pathways of greenhouse gas production. Front Earth Sci 7:59
Woolf D, Lehmann J (2019) Microbial models with minimal mineral protection can explain long-term soil organic carbon persistence. Sci Rep 9(1):1–8
Youngblut ND, Barnett SE, Buckley DH (2018) HTSSIP: an R package for analysis of high throughput sequencing data from nucleic acid stable isotope probing (SIP) experiments. PLoS One. https://doi.org/10.1371/journal.pone.0189616
Youngblut ND, Barnett SE, Buckley DH (2018) SIPSim: a modeling toolkit to predict accuracy and aid design of DNA-SIP experiments. Front Microbiol 9:570
Zakem EJ, Polz MF, Follows MJ (2020) Redox-informed models of global biogeochemical cycles. Nat Commun 11(1):1–10
Zampieri G, Vijayakumar S, Yaneske E, Angione C (2019) Machine and deep learning meet genome-scale metabolic modeling. PLoS Comput Biol 15(7):e1007084
Zegeye EK, Sadler NC, Lomas GX, Attah K, Jansson JK, Hofmockel KS, Anderton CR, Wright A (2020) Activity-based protein profiling of chitin catabolism. ChemBioChem. https://doi.org/10.1002/cbic.202000616
Zhang Q, Wang Y-P, Matear R, Pitman A, Dai Y (2014) Nitrogen and phosphorous limitations significantly reduce future allowable CO2 emissions. Geophys Res Lett 41(2):632–637
Zhang Y, Lavallee JM, Robertson AD, Even R, Ogle SM, Paustian K, Cotrufo MF (2021) Simulating measurable ecosystem carbon and nitrogen dynamics with the mechanistically-defined MEMS 2.0 model. Biogeosciences Discussions: 1–37
Zhao Q, Callister SJ, Thompson AM, Kukkadapu RK, Tfaily MM, Bramer LM, Qafoku NP, Bell SL, Hobbie SE, Seabloom EW (2020) Strong mineralogic control of soil organic matter composition in response to nutrient addition across diverse grassland sites. Sci Total Environ 736:137839
Acknowledgements
We thank Hyun-Seob Song, Josh Schimel, and one anonymous reviewer for helpful comments on this manuscript. This research was supported by the US Department of Energy Office of Biological and Environmental Research (BER) and is a contribution to the Scientific Focus Area ‘Phenotypic response of the soil microbiome to environmental perturbations’. Pacific Northwest National Laboratory is operated for the Department of Energy by Battelle Memorial Institute under contract DE-AC05-76RLO1830.
Funding
This research was supported by the US Department of Energy Office of Biological and Environmental Research (BER) and is a contribution to the Scientific Focus Area ‘Phenotypic response of the soil microbiome to environmental perturbations’. Pacific Northwest National Laboratory is operated for the Department of Energy by Battelle Memorial Institute under contract DE-AC05-76RLO1830.
Author information
Authors and Affiliations
Contributions
EBG conceived and wrote the manuscript with significant contributions from KSH.
Corresponding author
Ethics declarations
Conflict of interest
We declare no conflicts of interest/competing interests.
Additional information
Responsible Editor: Sharon A. Billings
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Graham, E.B., Hofmockel, K.S. Ecological stoichiometry as a foundation for omics-enabled biogeochemical models of soil organic matter decomposition. Biogeochemistry 157, 31–50 (2022). https://doi.org/10.1007/s10533-021-00851-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10533-021-00851-2