Abstract
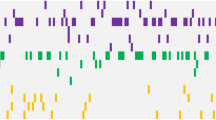
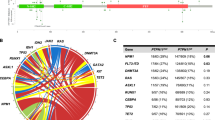
Germline DDX41 variants in myeloid neoplasms (MNs) are not uncommon, and we explored the prevalence and characterized the clinical and pathologic features in a cohort of 3132 unrelated adult MN patients. By targeted next-generation sequencing, we identified 28 patients (20 men and 8 women) with pathogenic germline DDX41 variants who developed acute myeloid leukemia (AML), in which only 3 (11%) had a family history (FH) of MNs. A subacute clinical course of cytopenia (mean duration of 11.2 months, range 0–72 months) prior to the initial AML diagnosis was accompanied by a low blast count (median at 30%, range 20–70%) in hypocellular marrows (93% of all patients), in vast contrast to the typical proliferative subtypes of AML in the elderly. Most patients had a normal karyotype (75%) and acquired a second DDX41 variant (69%). A favorable overall survival (OS) was observed in comparison to that of common subtypes of AML with wild-type DDX41 in age-matched patients. Our study demonstrated that the frequent germline pathogenic DDX41 variants characterized a clinically distinct AML entity. Features characteristic of DDX41-mutated AML include male predominance, often lack of FH, indolent course, low proliferative potential, frequent somatic DDX41 variants, and a favorable OS.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127:2391–405. https://doi.org/10.1182/blood-2016-03-643544.
Kennedy AL, Shimamura A. Genetic predisposition to MDS: clinical features and clonal evolution. Blood. 2019;133:1071–85. https://doi.org/10.1182/blood-2018-10-844662.
Rio-Machin A, Vulliamy T, Hug N, Walne A, Tawana K, Cardoso S, et al. The complex genetic landscape of familial MDS and AML reveals pathogenic germline variants. Nat Commun. 2020;11:1044. https://doi.org/10.1038/s41467-020-14829-5.
Sebert M, Passet M, Raimbault A, Rahme R, Raffoux E, Sicre de Fontbrune F, et al. Germline DDX41 mutations define a significant entity within adult MDS/AML patients. Blood. 2019;134:1441–4. https://doi.org/10.1182/blood.2019000909.
Tawana K, Drazer MW, Churpek JE. Universal genetic testing for inherited susceptibility in children and adults with myelodysplastic syndrome and acute myeloid leukemia: are we there yet? Leukemia. 2018;32:1482–92. https://doi.org/10.1038/s41375-018-0051-y.
Polprasert C, Schulze I, Sekeres MA, Makishima H, Przychodzen B, Hosono N, et al. Inherited and somatic defects in DDX41 in myeloid neoplasms. Cancer Cell. 2015;27:658–70. https://doi.org/10.1016/j.ccell.2015.03.017.
Cardoso SR, Ryan G, Walne AJ, Ellison A, Lowe R, Tummala H, et al. Germline heterozygous DDX41 variants in a subset of familial myelodysplasia and acute myeloid leukemia. Leukemia. 2016;30:2083–6. https://doi.org/10.1038/leu.2016.124.
Cheah JJC, Hahn CN, Hiwase DK, Scott HS, Brown AL. Myeloid neoplasms with germline DDX41 mutation. Int J Hematol. 2017;106:163–74. https://doi.org/10.1007/s12185-017-2260-y.
Quesada AE, Routbort MJ, DiNardo CD, Bueso-Ramos CE, Kanagal-Shamanna R, Khoury JD, et al. DDX41 mutations in myeloid neoplasms are associated with male gender, TP53 mutations and high-risk disease. Am J Hematol. 2019;94:757–66. https://doi.org/10.1002/ajh.25486.
Qu S, Li B, Qin T, Xu Z, Pan L, Hu N, et al. Molecular and clinical features of myeloid neoplasms with somatic DDX41 mutations. Br J Haematol. 2021;192:1006–10. https://doi.org/10.1111/bjh.16668.
Choi EJ, Cho YU, Hur EH, Jang S, Kim N, Park HS, et al. Unique ethnic features of DDX41 mutations in patients with idiopathic cytopenia of undetermined significance, myelodysplastic syndrome, or acute myeloid leukemia. Haematologica. 2021. https://doi.org/10.3324/haematol.2020.270553.
Baliakas P, Tesi B, Wartiovaara-Kautto U, Stray-Pedersen A, Friis LS, Dybedal I, et al. Nordic guidelines for germline predisposition to myeloid neoplasms in adults: recommendations for genetic diagnosis, clinical management and follow-up. Hemasphere. 2019;3:e321. https://doi.org/10.1097/HS9.0000000000000321.
Berger G, van den Berg E, Sikkema-Raddatz B, Abbott KM, Sinke RJ, Bungener LB, et al. Re-emergence of acute myeloid leukemia in donor cells following allogeneic transplantation in a family with a germline DDX41 mutation. Leukemia. 2017;31:520–2. https://doi.org/10.1038/leu.2016.310.
Li PB, Williams M, Zeng G, Lei L, White T, Xie W, et al. Genetic landscape and disease spectrum of hematologic neoplasms with germline DDX41 variants. 2021.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24. https://doi.org/10.1038/gim.2015.30.
D’Agostino M, Zaccaria GM, Ziccheddu B, Rustad EH, Genuardi E, Capra A, et al. Early relapse risk in patients with newly diagnosed multiple myeloma characterized by next-generation sequencing. Clin Cancer Res. 2020;26:4832–41. https://doi.org/10.1158/1078-0432.CCR-20-0951.
Shah V, Johnson DC, Sherborne AL, Ellis S, Aldridge FM, Howard-Reeves J, et al. Subclonal TP53 copy number is associated with prognosis in multiple myeloma. Blood. 2018;132:2465–9. https://doi.org/10.1182/blood-2018-06-857250.
Corre J, Cleynen A, Robiou du Pont S, Buisson L, Bolli N, Attal M, et al. Multiple myeloma clonal evolution in homogeneously treated patients. Leukemia. 2018;32:2636–47. https://doi.org/10.1038/s41375-018-0153-6.
Kortum KM, Mai EK, Hanafiah NH, Shi CX, Zhu YX, Bruins L, et al. Targeted sequencing of refractory myeloma reveals a high incidence of mutations in CRBN and Ras pathway genes. Blood. 2016;128:1226–33. https://doi.org/10.1182/blood-2016-02-698092.
Juliusson G, Jadersten M, Deneberg S, Lehmann S, Mollgard L, Wennstrom L, et al. The prognostic impact of FLT3-ITD and NPM1 mutation in adult AML is age-dependent in the population-based setting. Blood Adv. 2020;4:1094–101. https://doi.org/10.1182/bloodadvances.2019001335.
Sakaguchi M, Yamaguchi H, Najima Y, Usuki K, Ueki T, Oh I, et al. Prognostic impact of low allelic ratio FLT3-ITD and NPM1 mutation in acute myeloid leukemia. Blood Adv. 2018;2:2744–54. https://doi.org/10.1182/bloodadvances.2018020305.
Schnittger S, Bacher U, Kern W, Alpermann T, Haferlach C, Haferlach T. Prognostic impact of FLT3-ITD load in NPM1 mutated acute myeloid leukemia. Leukemia. 2011;25:1297–304. https://doi.org/10.1038/leu.2011.97.
Lewinsohn M, Brown AL, Weinel LM, Phung C, Rafidi G, Lee MK, et al. Novel germ line DDX41 mutations define families with a lower age of MDS/AML onset and lymphoid malignancies. Blood. 2016;127:1017–23. https://doi.org/10.1182/blood-2015-10-676098.
Vairo FPE, Ferrer A, Cathcart-Rake E, King RL, Howard MT, Viswanatha DS, et al. Novel germline missense DDX41 variant in a patient with an adult-onset myeloid neoplasm with excess blasts without dysplasia. Leuk Lymphoma. 2019;60:1337–9. https://doi.org/10.1080/10428194.2018.1522443.
Li MM, Datto M, Duncavage EJ, Kulkarni S, Lindeman NI, Roy S, et al. Standards and guidelines for the interpretation and reporting of sequence variants in cancer: a joint consensus recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J Mol Diagn. 2017;19:4–23. https://doi.org/10.1016/j.jmoldx.2016.10.002.
Mendez-Ferrer S, Garcia-Fernandez M, de Castillejo CL. Convert and conquer: the strategy of chronic myelogenous leukemic cells. Cancer Cell. 2015;27:611–3. https://doi.org/10.1016/j.ccell.2015.04.012.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Disco. 2012;2:401–4. https://doi.org/10.1158/2159-8290.CD-12-0095.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1. https://doi.org/10.1126/scisignal.2004088.
Patkar N, Kakirde C, Shaikh AF, Salve R, Bhanshe P, Chatterjee G, et al. Clinical impact of panel-based error-corrected next generation sequencing versus flow cytometry to detect measurable residual disease (MRD) in acute myeloid leukemia (AML). Leukemia. 2021;35:1392–404. https://doi.org/10.1038/s41375-021-01131-6.
Maciejewski JP, Padgett RA, Brown AL, Muller-Tidow C. DDX41-related myeloid neoplasia. Semin Hematol. 2017;54:94–97. https://doi.org/10.1053/j.seminhematol.2017.04.007.
Polprasert C, Takeda J, Niparuck P, Rattanathammethee T, Pirunsarn A, Suksusut A, et al. Novel DDX41 variants in Thai patients with myeloid neoplasms. Int J Hematol. 2020;111:241–6. https://doi.org/10.1007/s12185-019-02770-3.
Sanders MA. Lifting the veil on germline DDX41 mutations. Blood. 2019;134:1368–70. https://doi.org/10.1182/blood.2019002982.
Varga RE, Schule R, Fadel H, Valenzuela I, Speziani F, Gonzalez M, et al. Do not trust the pedigree: reduced and sex-dependent penetrance at a novel mutation hotspot in ATL1 blurs autosomal dominant inheritance of spastic paraplegia. Hum Mutat. 2013;34:860–3. https://doi.org/10.1002/humu.22309.
Cooper DN, Krawczak M, Polychronakos C, Tyler-Smith C, Kehrer-Sawatzki H. Where genotype is not predictive of phenotype: towards an understanding of the molecular basis of reduced penetrance in human inherited disease. Hum Genet. 2013;132:1077–130. https://doi.org/10.1007/s00439-013-1331-2.
Al-Mulla F, Bland JM, Serratt D, Miller J, Chu C, Taylor GT. Age-dependent penetrance of different germline mutations in the BRCA1 gene. J Clin Pathol. 2009;62:350–6. https://doi.org/10.1136/jcp.2008.062646.
van Rijsingen IA, Nannenberg EA, Arbustini E, Elliott PM, Mogensen J, Hermans-van Ast JF, et al. Gender-specific differences in major cardiac events and mortality in lamin A/C mutation carriers. Eur J Heart Fail. 2013;15:376–84. https://doi.org/10.1093/eurjhf/hfs191.
Author information
Authors and Affiliations
Contributions
PL designed the study and drafted the manuscript. TW, WX, WC, DP, GZ, H-YW, MW, and SB collected patients’ clinical and family history, morphologic, cytogenetic, and molecular data. JV and TK examined patients and performed DDX41 germline testing. PL, TW, WX, WC, DP, GZ, H-YW, and SB interpreted and classified all variants by NGS testing. PL, MW, and JLP reviewed the bone marrow examination. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, P., White, T., Xie, W. et al. AML with germline DDX41 variants is a clinicopathologically distinct entity with an indolent clinical course and favorable outcome. Leukemia 36, 664–674 (2022). https://doi.org/10.1038/s41375-021-01404-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-021-01404-0
This article is cited by
-
Impaired binding affinity of YTHDC1 with METTL3/METTL14 results in R-loop accumulation in myelodysplastic neoplasms with DDX41 mutation
Leukemia (2024)
-
Ribosome profiling analysis reveals the roles of DDX41 in translational regulation
International Journal of Hematology (2023)
-
DDX41 coordinates RNA splicing and transcriptional elongation to prevent DNA replication stress in hematopoietic cells
Leukemia (2022)
-
Approach Toward Germline Predisposition Syndromes in Patients with Hematologic Malignancies
Current Hematologic Malignancy Reports (2022)
-
Germline and Somatic Defects in DDX41 and its Impact on Myeloid Neoplasms
Current Hematologic Malignancy Reports (2022)