Abstract
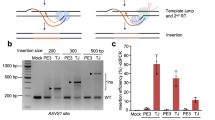
Genomic insertions, duplications and insertion/deletions (indels), which account for ~14% of human pathogenic mutations, cannot be accurately or efficiently corrected by current gene-editing methods, especially those that involve larger alterations (>100 base pairs (bp)). Here, we optimize prime editing (PE) tools for creating precise genomic deletions and direct the replacement of a genomic fragment ranging from ~1 kilobases (kb) to ~10 kb with a desired sequence (up to 60 bp) in the absence of an exogenous DNA template. By conjugating Cas9 nuclease to reverse transcriptase (PE-Cas9) and combining it with two PE guide RNAs (pegRNAs) targeting complementary DNA strands, we achieve precise and specific deletion and repair of target sequences via using this PE-Cas9-based deletion and repair (PEDAR) method. PEDAR outperformed other genome-editing methods in a reporter system and at endogenous loci, efficiently creating large and precise genomic alterations. In a mouse model of tyrosinemia, PEDAR removed a 1.38-kb pathogenic insertion within the Fah gene and precisely repaired the deletion junction to restore FAH expression in liver.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
A Reporting Summary for this article is available as a Supplementary Information file. The raw gel images underlying Figs. 1c,d, 2b,c,f,g, 3d and 4f and Supplementary Figs. 2d,e, 3a,b,e,f and 4a,b are provided as a Source Data file and an additional Supplementary Data file, respectively. The NCBI ClinVar database is accessible at https://www.ncbi.nlm.nih.gov/clinvar/. The raw DNA sequencing data are available at the NCBI Sequence Read Archive database under accession numbers PRJNA746292 and PRJNA746489. Source data are provided with this paper.
References
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014).
Cordaux, R. & Batzer, M. A. The impact of retrotransposons on human genome evolution. Nat. Rev. Genet. 10, 691–703 (2009).
Chen, J. M., Stenson, P. D., Cooper, D. N. & Ferec, C. A systematic analysis of LINE-1 endonuclease-dependent retrotranspositional events causing human genetic disease. Hum. Genet. 117, 411–427 (2005).
Hancks, D. C. & Kazazian, H. H. Roles for retrotransposon insertions in human disease. Mob. DNA 7, 9 (2016).
Wang, L., Norris, E. T. & Jordan, I. K. Human retrotransposon insertion polymorphisms are associated with health and disease via gene regulatory phenotypes. Front. Microbiol. 8, 1418 (2017).
Hancks, D. C. & Kazazian, H. H. Jr. Active human retrotransposons: variation and disease. Curr. Opin. Genet. Dev. 22, 191–203 (2012).
Qian, Y. et al. Identification of pathogenic retrotransposon insertions in cancer predisposition genes. Cancer Genet. 216-217, 159–169 (2017).
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Kato, T. et al. Creation of mutant mice with megabase-sized deletions containing custom-designed breakpoints by means of the CRISPR/Cas9 system. Sci. Rep. 7, 59 (2017).
Hara, S. et al. Microinjection-based generation of mutant mice with a double mutation and a 0.5 Mb deletion in their genome by the CRISPR/Cas9 system. J. Reprod. Dev. 62, 531–536 (2016).
Wang, L. et al. Large genomic fragment deletion and functional gene cassette knock-in via Cas9 protein mediated genome editing in one-cell rodent embryos. Sci. Rep. 5, 17517 (2015).
Yeh, C. D., Richardson, C. D. & Corn, J. E. Advances in genome editing through control of DNA repair pathways. Nat. Cell Biol. 21, 1468–1478 (2019).
Zheng, Q. et al. Precise gene deletion and replacement using the CRISPR/Cas9 system in human cells. Biotechniques 57, 115–124 (2014).
Cox, D. B., Platt, R. J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Liu, M. et al. Methodologies for improving HDR efficiency. Front. Genet. 9, 691 (2018).
Anzalone, A. V. et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576, 149–157 (2019).
Matsoukas, I. G. Prime editing: genome editing for rare genetic diseases without double-strand breaks or donor DNA. Front. Genet. 11, 528 (2020).
Liu, P. et al. Improved prime editors enable pathogenic allele correction and cancer modelling in adult mice. Nat. Commun. 12, 2121 (2021).
Jang, H. et al. Prime editing enables precise genome editing in mouse liver and retina. Preprint at bioRxiv https://doi.org/10.1101/2021.01.08.425835 (2021).
Schene, I. F. et al. Prime editing for functional repair in patient-derived disease models. Nat. Commun. 11, 5352 (2020).
Jiang, Y. Y. et al. Prime editing efficiently generates W542L and S621I double mutations in two ALS genes in maize. Genome Biol. 21, 257 (2020).
Song, X., Huang, H., Xiong, Z., Ai, L. & Yang, S. CRISPR–Cas9D10A nickase-assisted genome editing in Lactobacillus casei. Appl. Environ. Microbiol. 83, 1259–1275 (2017).
Cho, S. W. et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 24, 132–141 (2014).
Sfeir, A. & Symington, L. S. Microhomology-mediated end joining: a back-up survival mechanism or dedicated pathway? Trends Biochem. Sci 40, 701–714 (2015).
Bhargava, R., Onyango, D. O. & Stark, J. M. Regulation of single-strand annealing and its role in genome maintenance. Trends Genet. 32, 566–575 (2016).
Kim, H. K. et al. Predicting the efficiency of prime editing guide RNAs in human cells. Nat. Biotechnol. 39, 198–206 (2021).
Mir, A. et al. Heavily and fully modified RNAs guide efficient SpyCas9-mediated genome editing. Nat. Commun. 9, 2641 (2018).
Certo, M. T. et al. Tracking genome engineering outcome at individual DNA breakpoints. Nat. Methods 8, 671–676 (2011).
Zhan, H., Li, A., Cai, Z., Huang, W. & Liu, Y. Improving transgene expression and CRISPR–Cas9 efficiency with molecular engineering-based molecules. Clin. Transl Med. 10, e194 (2020).
Chen, R. et al. Enrichment of transiently transfected mesangial cells by cell sorting after cotransfection with GFP. Am. J. Physiol. 276, F777–F785 (1999).
Homann, S. et al. A novel rapid and reproducible flow cytometric method for optimization of transfection efficiency in cells. PLoS ONE 12, e0182941 (2017).
Pham, C. T., MacIvor, D. M., Hug, B. A., Heusel, J. W. & Ley, T. J. Long-range disruption of gene expression by a selectable marker cassette. Proc. Natl Acad. Sci. USA 93, 13090–13095 (1996).
Grompe, M. et al. Loss of fumarylacetoacetate hydrolase is responsible for the neonatal hepatic dysfunction phenotype of lethal albino mice. Genes Dev. 7, 2298–2307 (1993).
Paulk, N. K. et al. Adeno-associated virus gene repair corrects a mouse model of hereditary tyrosinemia in vivo. Hepatology 51, 1200–1208 (2010).
Choi, J. et al. Precise genomic deletions using paired prime editing. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01025-z (in the press).
VanLith, C. J. et al. Ex vivo hepatocyte reprograming promotes homology-directed DNA repair to correct metabolic disease in mice after transplantation. Hepatol. Commun. 3, 558–573 (2019).
Dutta, A. et al. Microhomology-mediated end joining is activated in irradiated human cells due to phosphorylation-dependent formation of the XRCC1 repair complex. Nucleic Acids Res. 45, 2585–2599 (2016).
Aida, T. et al. Gene cassette knock-in in mammalian cells and zygotes by enhanced MMEJ. BMC Genomics 17, 979 (2016).
Warby, S. C. et al. CAG expansion in the Huntington disease gene is associated with a specific and targetable predisposing haplogroup. Am. J. Hum. Genet. 84, 351–366 (2009).
Wang, Y. et al. Identification of a Xist silencing domain by Tiling CRISPR. Sci. Rep. 9, 2408 (2019).
He, W. et al. De novo identification of essential protein domains from CRISPR–Cas9 tiling-sgRNA knockout screens. Nat. Commun. 10, 4541 (2019).
Xue, W. et al. Response and resistance to NF-κB inhibitors in mouse models of lung adenocarcinoma. Cancer Discov. 1, 236–247 (2011).
Magoc, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Acknowledgements
We thank C. Mello, P. Zamore, S. Wolfe, T. Flotte and E. Sontheimer for discussions and E. Haberlin for editing the manuscript. We thank E. Sontheimer (UMass Medical School) for providing the HEK293T-TLR cell line and M. Grompe (Oregon Health & Science University) for providing the FahΔExon5 mice. We thank Y. Liu, Y. Gu and E. Kittler in the UMass Morphology, Flow Cytometry and Deep Sequencing Cores for support. W.X. was supported by grants from the National Institutes of Health (DP2HL137167, P01HL131471 and UG3HL147367), American Cancer Society (129056-RSG-16-093), the Lung Cancer Research Foundation and the Cystic Fibrosis Foundation. T.J. was supported by grants from National Institutes of Health (K99HL153940).
Author information
Authors and Affiliations
Contributions
T.J. and W.X. designed the study. T.J. performed experiments. T.J., X.-O.Z. and Z.W. analyzed the data. T.J. and W.X. wrote the manuscript with comments from all authors.
Corresponding author
Ethics declarations
Competing interests
UMass has filed a patent application on this work. W.X. is a consultant for the Cystic Fibrosis Foundation Therapeutics Lab. The other authors declare no competing interests.
Additional information
Peer review information Nature Biotechnology thanks Daesik Kim and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8 and Tables 1–3.
Supplementary Data
Unprocessed gel and blot images for Supplementary Figs. 2d,e, 3a,b,e,f and 4a,b.
Source data
Source Data Fig. 1
Unprocessed gels and blots for Fig. 1c,d.
Source Data Fig. 2
Unprocessed gels and blots for Fig. 2b,c,f,g.
Source Data Fig. 3
Unprocessed gels and blots for Fig. 3d.
Source Data Fig. 4
Unprocessed gels and blots for Fig. 4f.
Rights and permissions
About this article
Cite this article
Jiang, T., Zhang, XO., Weng, Z. et al. Deletion and replacement of long genomic sequences using prime editing. Nat Biotechnol 40, 227–234 (2022). https://doi.org/10.1038/s41587-021-01026-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-01026-y
This article is cited by
-
Targeted genome-modification tools and their advanced applications in crop breeding
Nature Reviews Genetics (2024)
-
CRISPR technologies for genome, epigenome and transcriptome editing
Nature Reviews Molecular Cell Biology (2024)
-
BacPE: a versatile prime-editing platform in bacteria by inhibiting DNA exonucleases
Nature Communications (2024)
-
Precise genome-editing in human diseases: mechanisms, strategies and applications
Signal Transduction and Targeted Therapy (2024)
-
A prime editor mouse to model a broad spectrum of somatic mutations in vivo
Nature Biotechnology (2024)