Abstract
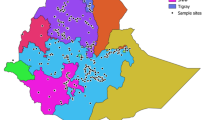
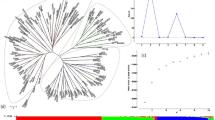
The cultivated barley ranks fourth cereal crop in the world. The demand for higher-yielding, nutritious, and better-adapted crop varieties has increased the need to exploit genebanks diversity. Thus, assessing the genetic diversity of barley is essential to determine the genetic relationship between lines and deploy novel alleles in breeding programs. Here, we used 14 polymorphic Simple Sequence Repeat (SSR) markers to assess genetic diversity, genetic relationship, and population structure of 113 barley lines originated from 14 countries, and the majority from Africa. The average alleles per locus of 5.36, Polymorphism Information Content (PIC) of 0.57, and genetic diversity of 0.61 indicate a high degree of genetic variation in this collection. The Analysis of Molecular Variance AMOVA showed genetic variation within countries to be higher (74%) than among countries (26%). The STRUCTURE analysis was consistent with neighbor-joining clustering and principal components analysis, which identified three main subpopulations clustered primarily according to their geographic origin. Growing environments, migration between and within countries might have caused a considerable genetic diversity observed in the North African barley germplasm. The results of this study contribute to the conservation and utilization of these barley germplasm.




Similar content being viewed by others
References
Bekele E (1983) Some measures of gene diversity analysis on landrace populations of Ethiopian barley. Hereditas 98:127–143
Blake VC et al (2019) GrainGenes: centralized small grain resources and digital platform for geneticists and breeders. Database. https://doi.org/10.1093/database/baz065
Cross R (1994) Geographical trends within a diverse spring barley collection as identified by agro/morphological and electrophoretic data. Theor Appl Genet 88:597–603. https://doi.org/10.1007/bf01240924
Dakir E-H, Ruiz M-L, García P, de la Vega MP (2002) Genetic variability evaluation in a Moroccan collection of barley, Hordeum vulgare L., by means of storage proteins and RAPDs. Genet Resour Crop Evol 49:619–631
Davila J, Loarce Y, Ramsay L, Waugh R, Ferrer E (1999) Comparison of RAMP and SSR markers for the study of wild barley genetic diversity. Hereditas. https://doi.org/10.1111/j.1601-5223.1999.00005.x
Earl DA (2012) Structure harvester: a website and program for visualizing structure output and implementing the Evanno method. Conser Gene Resour 4:359–361
Elakhdar A, Kumamaru T, Qualset CO, Brueggeman RS, Amer K, Capo-chichi L (2018) Assessment of genetic diversity in Egyptian barley (Hordeum vulgare L.) genotypes using SSR and SNP markers. Genet Resour Crop Evol 65:1937–1951. https://doi.org/10.1007/s10722-018-0666-x
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol. https://doi.org/10.1111/j.1365-294x.2005.02553.x
Ferreira JR, Pereira JF, Turchetto C, Minella E, Consoli L, Delatorre CA (2016) Assessment of genetic diversity in Brazilian barley using SSR markers. Genet Mol Biol 39:86–96
Food and Agriculture Organization of the Unites Nations (2020,December 22) FAOSTAT. http://www.fao.org/faostat/en/#data/QC/visualize. Accessed January 2 2021
Govindaraj M, Vetriventhan M, Srinivasan M (2015) Importance of genetic diversity assessment in crop plants and its recent advances: an overview of its analytical perspectives. Gene Res Int. https://doi.org/10.1155/2015/431487
Gupta P, Balyan H, Sharma P, Ramesh B (1996) Microsatellites in plants: a new class of molecular markers. Curr Sci 70:45–54
Hamza S, Hamida WB, Rebaï A, Harrabi M (2004) SSR-based genetic diversity assessment among Tunisian winter barley and relationship with morphological traits. Euphytica 135:107–118
Jilal A et al (2008) Genetic diversity of ICARDA’s worldwide barley landrace collection. Genet Resour Crop Evol 55:1221–1230. https://doi.org/10.1007/s10722-008-9322-1
Jin L, Chakraborty R (1994) Estimation of genetic distance and coefficient of gene diversity from single-probe multilocus DNA fingerprinting data. Mol Biol Evol 11:120–127. https://doi.org/10.1093/oxfordjournals.molbev.a040086
Kantartzi SK (2013) Microsatellites: methods and protocols. Humana Press
Khodayari H, Saeidi H, Roofigar AA, Rahiminejad MR, Pourkheirandish M, Komatsuda T (2012) Genetic diversity of cultivated barley landraces in Iran measured using microsatellites. Int J Biosci, Biochem Bioinfo 2:287. https://doi.org/10.7763/ijbbb.2012.v2.118
Kilian B et al (2006) Haplotype structure at seven barley genes: relevance to gene pool bottlenecks, phylogeny of ear type and site of barley domestication. Mol Genet Genomics 276:230–241
Kumlehn J, Stein N (2014) Biotechnological approaches to barley improvement. Springer, Berlin
Lamara M, Zhang LY, Marchand S, Tinker NA, Belzile F (2013) Comparative analysis of genetic diversity in Canadian barley assessed by SSR, DarT, and pedigree data. Genome. https://doi.org/10.1139/gen-2013-0048
Li H, Zhou M, Liu C (2009) A major QTL conferring crown rot resistance in barley and its association with plant height. Theor Appl Genet 118:903–910
Li H, Chen G, Yan W (2015) Molecular characterization of barley 3H semi-dwarf genes. PLoS ONE. https://doi.org/10.1371/journal.pone.0120558
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Mahmoud MOM, Hamza S (2009) Genetic diversity in local barley accessions collected from different geographical regions of Tunisia. Plant Gene Resources. https://doi.org/10.1017/s1479262108162086
Meints B, Cuesta-Marcos A, Fisk S, Ross A, Hayes P (2016) Food barley quality improvement and germplasm utilization. In: Exploration, Identification and Utilization of Barley Germplasm, Elsevier. https://doi.org/10.1016/b978-0-12-802922-0.00003-0
Munoz-Amatriain M et al (2014) The USDA barley core collection: genetic diversity, population structure, and potential for genome-wide association studies. PLoS ONE. https://doi.org/10.1371/journal.pone.0094688
Naceur AB, Chaabane R, El-Faleh M, Abdelly C, Ramla D, Nada A, Sakr M (2012) Genetic diversity analysis of North Africa’s barley using SSR markers. J Gene Eng Biotech 10:13–21
Orabi J, Jahoor A, Backes G (2009) Genetic diversity and population structure of wild and cultivated barley from West Asia and North Africa. Plant Breed 128:606–614
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel Population genetic software for teaching and research. Mol Ecol Notes. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Perry DJ, Fernando U, Lee S-J (2014) Simple sequence repeat-based identification of Canadian malting barley varieties. Can J Plant Sci 94:485–496
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ren X, Nevo E, Sun D, Sun G (2013) Tibet as a potential domestication center of cultivated barley of China. PLoS ONE. https://doi.org/10.1371/journal.pone.0062700
Řepková J, Dreiseitl A (2010) Candidate markers for powdery mildew resistance genes from wild barley PI284752. Euphytica 175:283–292
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard R (1984) Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. Proceed National Acad Sci 81:8014–8018
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Singh B, Singh AK (2015) Marker-assisted plant breeding: principles and practices. Springer, Berlin
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Udupa SM, Robertson LD, Weigand F, Baum M, Kahl G (1999) Allelic variation at (TAA)n microsatellite loci in a world collection of chickpea (Cicer arietinum L) germplasm. Mol Gen Genet 261:354–363
Ullrich SE (2010) Barley: production, improvement, and uses. Wiley, USA
Varshney RK, Pandey MK, Chitikineni A (2018) Plant genetics and molecular biology: an introduction. In: Plant Genetics and Molecular Biology. Springer, pp 1–9. https://doi.org/10.1007/10_2017_45
von Bothmer R, Sato K, Komatsuda T, Yasuda S, Fischbeck G (2003) The domestication of cultivated barley Diversity in barley (Hordeum vulgare). In: von Bothmer R, van Hintum T, Knüpffer H, Sato K (eds) Diversity in barley. Elsevier, Nertherland
Wang IJ, Bradburd GS (2014) Isolation by environment. Mol Ecol. https://doi.org/10.1111/mec.12938
Wang J-m, Yang J-m, Zhu J-h, Jia Q-j, Tao Y-z (2010) Assessment of genetic diversity by simple sequence repeat markers among forty elite varieties in the germplasm for malting barley. Breed J Zhejiang University Sci B. https://doi.org/10.1631/jzus.b0900414
Zong-Yun F, Xian-Jun L, Zhang YZ, Hong-Qing L (2006) Genetic diversity analysis of Tibetan wild barley using SSR markers. Acta Genet Sin 33:917–928
Funding
This research was funded by FAO/ITPGRFA/EU, grant number W3B-PR-21-Morocco.
Author information
Authors and Affiliations
Contributions
SN contributed to investigation, formal analysis, data curation and writing original, ZG contributed to formal analysis, FH contributed to formal analysis and editing manuscript, HO contributed to supervision and conceptualization, MI contributed to supervision, SMU contributed to conceptualization, supervision, project administration and funding acquisition. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declared that they have no conflict of interest.
Human and animal rights
This study did not involve humans or animals.
Additional information
Communicated by A. Börner.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nyiraguhirwa, S., Grana, Z., Henkrar, F. et al. Genetic diversity and structure of a barley collection predominatly from North African region. CEREAL RESEARCH COMMUNICATIONS 50, 647–654 (2022). https://doi.org/10.1007/s42976-021-00209-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42976-021-00209-2