Abstract
Macrophages exhibit a spectrum of activation states ranging from classical to alternative activation1. Alternatively, activated macrophages are involved in diverse pathophysiological processes such as confining tissue parasites2, improving insulin sensitivity3 or promoting an immune-tolerant microenvironment that facilitates tumour growth and metastasis4. Recently, the metabolic regulation of macrophage function has come into focus as both the classical and alternative activation programmes require specific regulated metabolic reprogramming5. While most of the studies regarding immunometabolism have focussed on the catabolic pathways activated to provide energy, little is known about the anabolic pathways mediating macrophage alternative activation. In this study, we show that the anabolic transcription factor sterol regulatory element binding protein 1 (SREBP1) is activated in response to the canonical T helper 2 cell cytokine interleukin-4 to trigger the de novo lipogenesis (DNL) programme, as a necessary step for macrophage alternative activation. Mechanistically, DNL consumes NADPH, partitioning it away from cellular antioxidant defences and raising reactive oxygen species levels. Reactive oxygen species serves as a second messenger, signalling sufficient DNL, and promoting macrophage alternative activation. The pathophysiological relevance of this mechanism is validated by showing that SREBP1/DNL is essential for macrophage alternative activation in vivo in a helminth infection model.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All the data presented in this manuscript are available upon reasonable request to the corresponding authors. Transfer of materials requires materials transfer agreements. The RNA-seq dataset is deposited in the Gene Expression Omnibus under accession number GSE179066. Source data are provided with this paper.
References
Murray, P. J. et al. Macrophage activation and polarization: nomenclature and experimental guidelines. Immunity 41, 14–20 (2014).
Chen, F. et al. An essential role for TH2-type responses in limiting acute tissue damage during experimental helminth infection. Nat. Med. 18, 260–266 (2012).
Chawla, A., Nguyen, K. D. & Goh, Y. P. Macrophage-mediated inflammation in metabolic disease. Nat. Rev. Immunol. 11, 738–749 (2011).
Mantovani, A., Allavena, P., Sica, A. & Balkwill, F. Cancer-related inflammation. Nature 454, 436–444 (2008).
O’Neill, L. A. & Pearce, E. J. Immunometabolism governs dendritic cell and macrophage function. J. Exp. Med. 213, 15–23 (2016).
Gerrick, K. Y. et al. Transcriptional profiling identifies novel regulators of macrophage polarization. PLoS ONE 13, e0208602 (2018).
Czimmerer, Z. et al. The transcription factor STAT6 mediates direct repression of inflammatory enhancers and limits activation of alternatively polarized macrophages. Immunity 48, 75–90 (2018).
Byles, V. et al. The TSC-mTOR pathway regulates macrophage polarization. Nat. Commun. 4, 2834 (2013).
Li, Y. et al. AMPK phosphorylates and inhibits SREBP activity to attenuate hepatic steatosis and atherosclerosis in diet-induced insulin-resistant mice. Cell Metab. 13, 376–388 (2011).
Walker, A. K. et al. Conserved role of SIRT1 orthologs in fasting-dependent inhibition of the lipid/cholesterol regulator SREBP. Genes Dev. 24, 1403–1417 (2010).
Haschemi, A. et al. The sedoheptulose kinase CARKL directs macrophage polarization through control of glucose metabolism. Cell Metab. 15, 813–826 (2012).
Shay, A. E. et al. IL-4 upregulates cyclooxygenase-1 expression in macrophages. J. Biol. Chem. 292, 14544–14555 (2017).
York, A. G. et al. Limiting cholesterol biosynthetic flux spontaneously engages type I IFN signaling. Cell 163, 1716–1729 (2015).
Sun, L. P., Seemann, J., Goldstein, J. L. & Brown, M. S. Sterol-regulated transport of SREBPs from endoplasmic reticulum to Golgi: Insig renders sorting signal in Scap inaccessible to COPII proteins. Proc. Natl Acad. Sci. USA 104, 6519–6526 (2007).
Neill, D. R. et al. Nuocytes represent a new innate effector leukocyte that mediates type-2 immunity. Nature 464, 1367–1370 (2010).
Allen, J. E. & Sutherland, T. E. Host protective roles of type 2 immunity: parasite killing and tissue repair, flip sides of the same coin. Semin. Immunol. 26, 329–340 (2014).
Hastings, R. H., Folkesson, H. G. & Matthay, M. A. Mechanisms of alveolar protein clearance in the intact lung. Am. J. Physiol. Lung Cell. Mol. Physiol. 286, L679–L689 (2004).
Berthiaume, Y., Albertine, K. H., Grady, M., Fick, G. & Matthay, M. A. Protein clearance from the air spaces and lungs of unanesthetized sheep over 144 h. J. Appl Physiol. 67, 1887–1897 (1989).
Chen, F. et al. Monocyte-derived alveolar macrophages mediate resistance to migrating helminths through depletion of arginine availability. Preprint at bioRxiv https://doi.org/10.1101/2020.10.02.322388 (2020).
Shimano, H. & Sato, R. SREBP-regulated lipid metabolism: convergent physiology—divergent pathophysiology. Nat. Rev. Endocrinol. 13, 710–730 (2017).
Wei, X. et al. Fatty acid synthesis configures the plasma membrane for inflammation in diabetes. Nature 539, 294–298 (2016).
Carroll, R. G. et al. An unexpected link between fatty acid synthase and cholesterol synthesis in pro-inflammatory macrophage activation. J. Biol. Chem. 293, 5509–5521 (2018).
Griess, B., Mir, S., Datta, K. & Teoh-Fitzgerald, M. Scavenging reactive oxygen species selectively inhibits M2 macrophage polarization and their pro-tumorigenic function in part, via Stat3 suppression. Free Radic. Biol. Med. 147, 48–60 (2020).
Kapoor, N. et al. Transcription factors STAT6 and KLF4 implement macrophage polarization via the dual catalytic powers of MCPIP. J. Immunol. 194, 6011–6023 (2015).
He, C. et al. Mitochondrial Cu,Zn-superoxide dismutase mediates pulmonary fibrosis by augmenting H2O2 generation. J. Biol. Chem. 286, 15597–15607 (2011).
He, C., Ryan, A. J., Murthy, S. & Carter, A. B. Accelerated development of pulmonary fibrosis via Cu,Zn-superoxide dismutase-induced alternative activation of macrophages. J. Biol. Chem. 288, 20745–20757 (2013).
Zhang, Y. et al. ROS play a critical role in the differentiation of alternatively activated macrophages and the occurrence of tumor-associated macrophages. Cell Res. 23, 898–914 (2013).
Sanin, D. E. et al. Mitochondrial membrane potential regulates nuclear gene expression in macrophages exposed to prostaglandin E2. Immunity 49, 1021–1033 e1026 (2018).
Huang, S. C. et al. Cell-intrinsic lysosomal lipolysis is essential for alternative activation of macrophages. Nat. Immunol. 15, 846–855 (2014).
Vats, D. et al. Oxidative metabolism and PGC-1β attenuate macrophage-mediated inflammation. Cell Metab. 4, 13–24 (2006).
Van den Bossche, J. & van der Windt, G. J. W. Fatty acid oxidation in macrophages and T cells: time for reassessment? Cell Metab. 28, 538–540 (2018).
Divakaruni, A. S. et al. Etomoxir inhibits macrophage polarization by disrupting CoA homeostasis. Cell Metab. 28, 490–503 (2018).
Nomura, M. et al. Fatty acid oxidation in macrophage polarization. Nat. Immunol. 17, 216–217 (2016).
Namgaladze, D. & Brune, B. Fatty acid oxidation is dispensable for human macrophage IL-4-induced polarization. Biochim. Biophys. Acta 1841, 1329–1335 (2014).
Wang, F. et al. Glycolytic stimulation is not a requirement for M2 macrophage differentiation. Cell Metab. 28, 463–475 (2018).
Lewis, C. A. et al. Tracing compartmentalized NADPH metabolism in the cytosol and mitochondria of mammalian cells. Mol. Cell 55, 253–263 (2014).
Rouzer, C. A., Scott, W. A., Griffith, O. W., Hamill, A. L. & Cohn, Z. A. Glutathione metabolism in resting and phagocytizing peritoneal macrophages. J. Biol. Chem. 257, 2002–2008 (1982).
Mestas, J. & Hughes, C. C. Of mice and not men: differences between mouse and human immunology. J. Immunol. 172, 2731–2738 (2004).
Robert, U. et al. Inhibition of acetyl-CoA carboxylase suppresses fatty acid synthesis and tumor growth of non-small-cell lung cancer in preclinical models. Nat. Med. 22, 1108–1119 (2016).
Liu, C. et al. Treg cells promote the SREBP1-dependent metabolic fitness of tumor-promoting macrophages via repression of CD8+ T cell-derived Interferon-gamma. Immunity 51, 381–397 e386 (2019).
Agius, L., Meredith, E. J. & Sherratt, H. S. Stereospecificity of the inhibition by etomoxir of fatty acid and cholesterol synthesis in isolated rat hepatocytes. Biochem. Pharmacol. 42, 1717–1720 (1991).
Oishi, Y. et al. SREBP1 contributes to resolution of pro-inflammatory TLR4 signaling by reprogramming fatty acid metabolism. Cell Metab. 25, 412–427 (2017).
Im, S. S. et al. Linking lipid metabolism to the innate immune response in macrophages through sterol regulatory element binding protein-1a. Cell Metab. 13, 540–549 (2011).
Lee, J. H. et al. SREBP-1a-stimulated lipid synthesis is required for macrophage phagocytosis downstream of TLR4-directed mTORC1. Proc. Natl Acad. Sci. USA 115, E12228–E12234 (2018).
Reboldi, A. et al. Inflammation. 25-hydroxycholesterol suppresses interleukin-1-driven inflammation downstream of type I interferon. Science 345, 679–684 (2014).
Guo, C. et al. Cholesterol homeostatic regulator SCAP-SREBP2 integrates NLRP3 inflammasome activation and cholesterol biosynthetic signaling in macrophages. Immunity 49, 842–856 (2018).
Feingold, K. R. et al. Mechanisms of triglyceride accumulation in activated macrophages. J. Leukoc. Biol. 92, 829–839 (2012).
Rosas-Ballina, M., Guan, X. L., Schmidt, A. & Bumann, D. Classical activation of macrophages leads to lipid droplet formation without de novo fatty acid synthesis. Front Immunol. 11, 131 (2020).
Jenkins, S. J. et al. Local macrophage proliferation, rather than recruitment from the blood, is a signature of TH2 inflammation. Science 332, 1284–1288 (2011).
Pello, O. M. et al. Role of c-MYC in alternative activation of human macrophages and tumor-associated macrophage biology. Blood 119, 411–421 (2012).
Clausen, B. E., Burkhardt, C., Reith, W., Renkawitz, R. & Forster, I. Conditional gene targeting in macrophages and granulocytes using LysMcre mice. Transgenic Res. 8, 265–277 (1999).
Tan, C. Y. et al. Brown adipose tissue thermogenic capacity is regulated by Elovl6. Cell Rep. 13, 2039–2047 (2015).
Kim, S. K., Oh, E., Yun, M., Lee, S. B. & Chae, G. T. Palmitate induces cisternal ER expansion via the activation of XBP-1/CCTα-mediated phospholipid accumulation in RAW 264.7 cells. Lipids Health Dis. 14, 73 (2015).
Nakano, A., Harada, T., Morikawa, S. & Kato, Y. Expression of leukocyte common antigen (CD45) on various human leukemia/lymphoma cell lines. Acta Pathol. Jpn 40, 107–115 (1990).
Kopf, M., Schneider, C. & Nobs, S. P. The development and function of lung-resident macrophages and dendritic cells. Nat. Immunol. 16, 36–44 (2015).
Misharin, A. V., Morales-Nebreda, L., Mutlu, G. M., Budinger, G. R. & Perlman, H. Flow cytometric analysis of macrophages and dendritic cell subsets in the mouse lung. Am. J. Respir. Cell Mol. Biol. 49, 503–510 (2013).
Guth, A. M. et al. Lung environment determines unique phenotype of alveolar macrophages. Am. J. Physiol. Lung Cell. Mol. Physiol. 296, L936–L946 (2009).
Hussell, T. & Bell, T. J. Alveolar macrophages: plasticity in a tissue-specific context. Nat. Rev. Immunol. 14, 81–93 (2014).
Austyn, J. M. & Gordon, S. F4/80, a monoclonal antibody directed specifically against the mouse macrophage. Eur. J. Immunol. 11, 805–815 (1981).
Feng, Y. H. & Mao, H. Expression and preliminary functional analysis of Siglec-F on mouse macrophages. J. Zhejiang Univ. Sci. B 13, 386–394 (2012).
Azzu, V. et al. Suppression of insulin-induced gene 1 (INSIG1) function promotes hepatic lipid remodelling and restrains NASH progression. Mol. Metab. https://doi.org/10.1016/j.molmet.2021.101210 (2021).
Matute-Bello, G. et al. An official American Thoracic Society workshop report: features and measurements of experimental acute lung injury in animals. Am. J. Respir. Cell Mol. Biol. 44, 725–738 (2011).
Jenkins, B., Ronis, M. & Koulman, A. LC–MS lipidomics: exploiting a simple high-throughput method for the comprehensive extraction of lipids in a ruminant fat dose–response study. Metabolites https://doi.org/10.3390/metabo10070296 (2020).
Chen, L. et al. NADPH production by the oxidative pentose-phosphate pathway supports folate metabolism. Nat. Metab. 1, 404–415 (2019).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Huber, W., von Heydebreck, A., Sultmann, H., Poustka, A. & Vingron, M. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 18, S96–S104 (2002).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Varemo, L., Nielsen, J. & Nookaew, I. Enriching the gene-set analysis of genome-wide data by incorporating directionality of gene expression and combining statistical hypotheses and methods. Nucleic Acids Res. 41, 4378–4391 (2013).
Horton, J. D. et al. Combined analysis of oligonucleotide microarray data from transgenic and knockout mice identifies direct SREBP target genes. Proc. Natl. Acad. Sci. USA 100, 12027–12032 (2003).
Acknowledgements
Lyz2-Cre mice were a kind gift from S. Jackowski. We thank D. Hart, S. Grocott, C. Beresford, J. Bacon, L. McKinven, E. Rasijeff and A. Lukasik and J. Warner from the Histology core for their excellent technical assistance. This research was supported by the Cambridge NIHR BRC Cell Phenotyping Hub. In particular, we thank E. Perez for advice and support in flow cytometry. We thank M. Murphy and H. Prag from the MRC Mitochondrial Biology Unit who kindly helped with the Seahorse experiments. We thank C. Frezza, L. Tronci and E. Nikitopoulou from the MRC cancer unit for their help in stable isotope tracer experimental design. We thank the IMS Genomics and Transcriptomics core for the RNA-seq analysis. This work was supported by the British Heart Foundation (RG/18/7/33636), the MRC (MC_UU_00014/2) and the FP7 MITIN (223450). K.P. was a recipient of a fellowship from the Wellcome Trust. A.N.J.M. and E.J. are supported by the Wellcome Trust (100963/Z/13/Z) and the MRC (U105178805). J.L. is a recipient fellowship of the British Heart Foundation. A.D. was a Marie-Curie Early-Stage Researcher supported by the European Union’s Horizon 2020 research and innovation programme (675585 Marie-Curie ITN ‘SymBioSys’) to J.S.-R. A.K. is supported by the Wellcome Trust (106260/Z/14/Z) and an ERC award (648889). P.F. is supported by the Science Foundation Ireland (10/IN.1/B3004). The IMS Genomics and Transcriptomics and Histology cores (B.M.-A., B.Y.H.L. and M.K.M.) are funded by the UK MRC Metabolic Disease Unit (MRC_MC_UU_12012/5) and a Wellcome Trust Strategic Award (100574/Z/12/Z). The Disease Model Core is part of the MRC Metabolic Diseases Unit (MRC_MC_UU_12012/5) and Wellcome Trust Strategic Award (100574/Z/12/Z).
Author information
Authors and Affiliations
Contributions
G.B. conceived the study, designed, performed and analysed the results of experiments; S.V. provided advice on design and discussion and analysed the lipidomic experiments; K.P. significantly helped with experiments; A.D. performed the bioinformatics pathway analysis; H.E.J. performed the helminths infection and collected the tissues; A.-C.G. helped with the bioinformatics analysis; B.M.-A. scored the lungs histology; B.L. analysed the RNA-seq; M.K.M. performed the library preparation; J.L. and M.D. provided technical assistance; S.C. provide expertise in the lipid synthesis assay; A.K. provided expertise on the manuscript; P.G.F. and A.N.J.M. provided expertise and designed the helminth infection study; A.V.-P. supervised the entire study; and G.B., S.V. and A.V.-P. prepared the manuscript. All authors contributed to and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
K.P. is currently employed by AstraZeneca. J.S.-R. has received funding from GSK and Sanofi and consultant fees from Travere Therapeutics. The other authors declare no competing interests.
Additional information
Peer review information Primary handling editors: George Caputa & Elena Bellafante. Nature Metabolism thanks Yu Li, Di Wang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 IL-4 induced SREBP1 activation is downstream of AKT and STAT6 signalling.
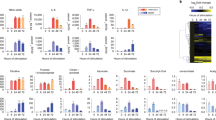
a, Protein expression of phosphorylated (Ser473) and total Akt in Vehicle (DMSO) or MK-2206-treated BMM in response to IL-4. Representative picture of n = 4 biological replicates. b, mRNA expression over BK of the SREBP1 target genes Fasn and Scd2 AKT in Vehicle (DMSO) or MK-2206-treated BMMФ in response to IL-4. Data are presented as the mean ± SEM of n = 8 biological replicates from 2 independent experiments. c, Gene expression analysis of the SREBP target genes in WT and Stat6-/- BMMФ. Data analysed from the publicly available dataset (GSE106706)8 with FDR<0.05 and Fc>2. Data has been analysed using a 2-way ANOVA followed by Sidak post-hoc test.
Extended Data Fig. 2 SIRT1 and AMPK are not responsible for SREBP1 activation in response to IL-4.
a-d, mRNA expression of the SREBP1 target genes Fasn and Scd2 and of the macrophages alternative activation markers Retnla and Mgl2 in BMMФ treated with SIRT1 activator II (a) or SIRT1 inhibitor (EX-527, b) or the AMPK inhibitor Compound C (c) or the AMPK activator AICAR (d) in response to IL-4. Data are presented as the mean ± SEM of n = 4 biological replicates per group. e and f, mRNA expression over BK of the SREBP1 target genes in 25-hydroxycholesterol-treated (25HC, n = 8 biological replicates from 2 independent experiments) (e) or SREBP-1c KO (n = 3 biological replicates) (f) BMMФ in response to IL-4. Data has been analysed using a 2-way ANOVA followed by a Dunnett (a-d) or Sidak (e and f) post-hoc test.
Extended Data Fig. 3 SREBP1 but not SREBP2 activation is required for macrophage alternative activation.
a and b, mRNA expression of Retnla and Mgl1 (a) and Tnf and Il1b (b) in WT and SCAP-KO BMMФ, 4h and 24h post IL-4 stimulation. mRNA expression over BK as mean ± SEM of n = 8 biological replicates from 2 independent experiments. c, Alternative activation of Vehicle (EtOH) or 25-hydroxycholesterol (25HC)-treated BMMФ in response to IL-4. Alternative activation was assessed by the expression of RELMα and CD206 by flow cytometry. The quantification of the number of alternatively activated macrophages is presented as mean ± SEM in d. Data of n = 4 biological replicates per group. e, Expression of the macrophage alternative activation markers Mrc1, Mgl1 Retlna and Arg1 in Vehicle (EtOH) or 25-hydroxycholesterol (25HC)-treated BMMФ in response to IL-4. mRNA expression over BK as mean ± SEM of n = 4 biological replicates per group. f, Expression of the macrophage alternative activation markers of Mrc1, Arg1 and Mgl1 in WT and SREBP1c-KO BMMФ in response to IL-4. mRNA expression over BK of n = 3 biological replicates. g, mRNA expression over BK of RAW264.7 macrophages transfected with siRNA against SREBP1, SREBP2 or both in response to IL-4. Data is expressed as mean ± SEM of n = 6 different experiments. Data has been analysed using a 2-way ANOVA followed by Sidak (a, b, d-f) or Tukey (g) post-hoc test.
Extended Data Fig. 4 Macrophage SREBP1 activation is required for immune response to helminth infection.
a, Neutrophil number presented as mean ± SEM in the lungs of naïve or 5- and 7-days post N. brasiliensis inoculation in WT and SCAP-KO mice. b, Eosinophil number presented as mean ± SEM in the lungs of naïve or 5- and 7-days post N. brasiliensis inoculation in WT and SCAP-KO mice. c, Percentage of alternatively activated alveolar macrophages in the lungs of naïve 5- and 7-days post N. brasiliensis inoculation in WT and SCAP-KO mice within the alveolar macrophage population. Alternative polarization was quantified by the expression of RELMα and CD206 by flow cytometry. d-f, Interstitial macrophages number (d), percentage of alternative activation (e) and number of alternatively activated interstitial macrophages (f) presented as mean ± SEM in the lungs of naïve or 5- and 7-days post N. brasiliensis inoculation in WT and SCAP-KO mice. g and h, Alveolar (g) and interstitial (h) neutrophil histological score in the lungs of naïve or 5- and 7-days post N. brasiliensis inoculation in WT and SCAP-KO mice. i, Correlation between the alveolar proteinaceous debris score and the number of alveolar macrophages 5 days post N. brasiliensis inoculation in WT and SCAP-KO mice. Pooled data as mean ± SEM n = 6-12 mice per group from 2 independent experiments. Data was analyzed using a two-way ANOVA followed by Sidak post-hoc test for comparison between genotypes at different days of post inoculation (a-h) or linear regression modelling (i).
Extended Data Fig. 5 Fatty acid synthesis in response to IL-4 requires SREBP1 activation.
a and b, Gene enrichment analysis of the pathways associated with de novo lipogenesis and SREBP activation from RNA sequencing comparing CTR and IL-4-treated BMMФ or of the interaction effect of IL-4 in WT and SCAP-KO BMMФ. Data from n = 6 biological replicates per group. c, Lipid synthesis rate in Lipopolysaccharide (LPS) or IL-4-treated BMMФ. The data represents the incorporation of radiolabelled 14C-acetate in the lipid fraction as mean ± SEM of n = 4 biological replicates per group. d, Proportion of newly synthesized palmitate per hour in control or IL-4 stimulated BMMФ. The data are presented as mean ± SEM of n = 4 biological replicates. e, Exogenous palmitate uptake rate of control or IL-4 stimulated BMMФ. The data are presented as mean ± SEM of n = 4 biological replicates. f, FAME composition in order of increasing chain length and desaturation of WT and SCAP-KO BMMФ in response to IL-4. Data is presented as mean ± SEM of n = 4 biological replicates. g, Total, essential and non-essential fatty acid content of WT and SCAP-KO BMMФ in response to IL-4. Data is presented as mean ± SEM of n = 4 biological replicates. h, Exogenous palmitate uptake rate of WT and SCAP-KO BMMФ in response to IL-4. The data are presented as mean ± SEM of n = 4 biological replicates. Statistical analysis of the RNAseq data is detailed in the methods section. Data has been analysed using a 2-way ANOVA followed by a Dunnett (c) or Sidak post-hoc test (f-h) or a two-tailed Student’s t-test (d and e).
Extended Data Fig. 6 Fatty acid but not cholesterol synthesis is required for macrophage alternative activation.
a, Lipid synthesis in BMMФ pre-treated with increasing doses of the FASN inhibitors C75 and cerulenin (Cer) 30 minutes prior 24h IL-4 stimulation. The data represents the incorporation of radiolabelled 14C-acetate in the lipid fraction as mean ± SEM of n = 7 biological replicates from 2 independent experiments. b, Expression of the macrophage alternative activation markers Mrc1, Mgl2, Arg1, Retlna and Il4i1 in Vehicle (DMSO) or C75 (10µM)-treated BMMФ in response to IL-4. mRNA expression over BK as mean ± SEM of n = 4 biological replicates per group. c, Alternative activation of BMMФ in response to IL-4 and pre-treated with increasing doses of Cerulenin (Cer). Alternative activation was assessed by flow cytometry using the co-expression of RELMα and CD301. The quantification of the number of M(IL-4) macrophages as mean ± SEM of n = 4 biological replicates is presented in d. e, Expression of the macrophage alternative activation markers Retlna and Mgl1 in Vehicle (DMSO) or cerulenin (2.5µg/mL)-treated BMMФ in response to IL-4. mRNA expression over BK as mean ± SEM of n = 4 biological replicates per group. f, Expression of the inflammatory cytokine Tnf and Il1b in Vehicle (DMSO) or C75 (10µM)-treated BMMФ in response to IL-4. mRNA expression over BK as mean ± SEM of n = 4 biological replicates per group. g and h, Expression of the SREBP2-target genes (g) and macrophage activation markers Tnf, Retlna and Mgl2 (h) in Vehicle (DMSO) or Simvastatin (10µM)-treated BMMФ in response to IL-4. mRNA expression over BK as mean ± SEM of n = 4 biological replicates per group. i, mRNA expression of the macrophage activation markers Retlna and Mgl2 in Vehicle (DMSO) or C75 (10µM)-treated BMMФ in response to IL-4 supplemented or not with HMG-CoA (1mM). mRNA expression over BK as mean ± SEM of n = 4 biological replicates per group. Data has been analysed using a 2-way ANOVA followed by a Dunnett (a and d) or Sidak post-hoc test (e-h) or one-way ANOVA followed by Tukey post-hoc test (i).
Extended Data Fig. 7 Fatty acid or cholesterol supplementation does not rescue the alternative activation of SCAP-KO or C75-treated macrophages.
mRNA expression of the SREBP1 target genes Fasn and Scd2 in C75-treated (a) or SCAP-KO (c) macrophages and of the macrophages alternative activation markers in C75-treated (b) or SCAP-KO (d) macrophages in response to IL-4 and/or palmitic acid (PA, 10 or 50 µM), oleic acid (OA, 10 or 50 µM) or water-soluble cholesterol (50 µM). mRNA expression of BK of n = 4 (C75) or n = 3 (SCAP-KO) biological replicates. Data has been analysed using a 2-way ANOVA followed by a Tukey post-hoc test.
Extended Data Fig. 8 Sources of the ROS in M(IL-4) cells.
a, Gene enrichment analysis of the pathways associated with redox homeostasis and response to oxidative stress in IL-4 treated BMMФ. Data from n = 6 biological replicates per group. Data of n = 6 biological replicates from 2 independent experiments per group. b, ROS accumulation in Vehicle (EtOH) or 25-hydroxycholesterol (25HC)-treated BMMФ in response to IL-4. ROS were quantified by the fluorescence ratio of CM-H2DCFDA over DNA (Hoechst). Data as mean ± SEM of n = 7 biological replicates from 2 independent experiments. c, ROS accumulation in Vehicle (DMSO) or cerulenin (1µg/mL)-treated BMMФ in response to IL-4. ROS were quantified by the median fluorescence intesnity of CM-H2DCFDA by flow cytometry. Data as mean ± SEM of n = 4 biological replicates. d, Reactive oxygen species (ROS) levels in BMMФ pre-treated with DPI (NADPH oxidase inhibitor), Allopurinol (xanthine oxidase inhibitor) or L-NAME (nitric oxide synthase inhibitor) prior 24h stimulation with IL-4 or LPS. ROS were quantified by the fluorescence ratio of CM-H2DCFDA over DNA (Hoechst). Data are presented as mean ± SEM of n = 4 biological replicates. e, Mitochondrial ROS production in response to IL-4. Mitochondrial ROS levels were determined by the fluorescence ratio of MitoSox over DNA (Hoechst). Data are presented as mean ± SEM of n = 8 biological replicates from 2 independent experiments. f, Time-course of fatty acid oxidation in response to IL-4 (10ng/mL). Data of n = 4 biological replicates. g and h, Oxygen consumption rate (OCR) in WT and SCAP-KO BMMФ (g) or in Vehicle (DMSO) or C75 (10 μM)-treated BMMФ (h) in control or IL-4 stimulated macrophages. OCR was monitored using an XF-96 Extracellular Flux Analyzer following the sequential treatments with oligomycin (oligo), FCCP and rotenone/antimycin (R/A). Data are presented as mean ± SEM of n = 4 (WT vs KO) and n = 8 from 2 independent experiments (DMSO vs C75) biological replicates per group. i, Fatty acid oxidation in IL-4 treated BMMФ in response to the AMPK activator AICAR (100 or 500 µM). Data of n = 4 biological replicates. j, Lipid synthesis in BMMФ treated with increasing doses of the AMPK activator AICAR in response to IL-4 (10ng/mL, 24h). Data of n = 4 biological replicates. k, Fatty acid oxidation assay of SCAP-KO macrophages in response to IL-4 (10ng/mL, 24h). Etomoxir (40 µM) was used as a negative control for FAO. Data of n = 4 biological replicates. l, Fatty acid oxidation assay in control or IL-4 stimulated macrophages treated or not with C75 (10 μM) or Cerulenin (1µg/mL) for 24h. Data of n = 4 biological replicates. m, ROS levels in C75 (10 μM) and/or Etomoxir (ETO, 40 μM)-treated BMMФ in response to IL-4. ROS were quantified by the fluorescence ratio of CM-H2DCFDA over DNA (Hoechst). Data of n = 4 biological replicates. n and o, Mitochondrial ROS production in WT and SCAP-KO BMMФ (n = 12 biological replicates from 3 independent experiments) (n) or in Vehicle (DMSO) or C75 (10 μM)-treated (n = 12 biological replicates from 3 independent experiments) BMMФ (o) in M(IL-4) macrophages. Mitochondrial ROS levels were determined by the fluorescence ratio of MitoSox over DNA (Hoechst). p, Reduced glutathione (GSH) levels in C75 (10 μM)-treated BMMФ in response to IL-4. Data presented as mean ± SEM of n = 4 biological replicates. Data has been analysed using a 2-way ANOVA followed by a Sidak (b, c, k, l and n-p) or Dunnett (d and j) or Tukey (g and h) post-hoc test or a two-tailed Student’s t-test (e) or a one-way ANOVA followed by Dunnett post-hoc test (f and i).
Extended Data Fig. 9 ROS scavenging impairs macrophage alternative activation.
a, ROS levels in N-acetyl cysteine (NAC, 10 mM)-treated BMMФ in response to IL-4. ROS were quantified by the fluorescence ratio of CM-H2DCFDA over DNA (Hoechst). Data as mean ± SEM of n = 8 biological replicates from 2 independent experiments. b and c, Alternative activation of NAC-treated BMMФ in response to IL-4. Alternative activation was assessed by the expression of RELMα and CD206 by flow cytometry. The quantification of the number of M(IL-4) macrophages is presented as mean ± SEM in c. Data of n = 8 biological replicates from 2 independent experiments. d, mRNA expression over BK of Arg1, Mgl1 and Retnla in NAC-treated BMMФ in response to IL-4. Data as mean ± SEM of n = 4 biological replicates. Data was analysed using a two-way ANOVA followed by Sidak post-hoc test.
Extended Data Fig. 10
Schematic representation of the mechanism by which DNL is activated and sensed in alternatively activated macrophages.
Supplementary information
Supplementary Information
Gating strategy.
Supplementary Table 1
Pathway analysis of WT BMDMs in response to IL-4 and pathway analysis of the interaction effect of IL-4 in WT and SCAP-KO BMDMs.
Supplementary Tables 2–4
Mouse models details and antibody and primers lists.
Source data
Source Data Fig. 1
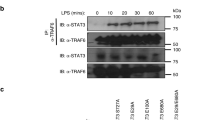
Uncropped western blots Fig. 1c.
Source Data Fig. 3
Uncropped western blots Fig. 3.
Source Data Extended Data Fig. 1
Uncropped western blots Extended Data Fig. 1a.
Rights and permissions
About this article
Cite this article
Bidault, G., Virtue, S., Petkevicius, K. et al. SREBP1-induced fatty acid synthesis depletes macrophages antioxidant defences to promote their alternative activation. Nat Metab 3, 1150–1162 (2021). https://doi.org/10.1038/s42255-021-00440-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-021-00440-5
This article is cited by
-
Regulation of genes involved in the metabolic adaptation of murine microglial cells in response to elevated HIF-1α mediated activation
Immunogenetics (2024)
-
Limited oxygen in standard cell culture alters metabolism and function of differentiated cells
The EMBO Journal (2024)
-
NAD+ metabolism-based immunoregulation and therapeutic potential
Cell & Bioscience (2023)
-
Adipose tissue macrophages: implications for obesity-associated cancer
Military Medical Research (2023)
-
Spatially resolved multi-omics highlights cell-specific metabolic remodeling and interactions in gastric cancer
Nature Communications (2023)