Abstract
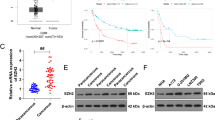
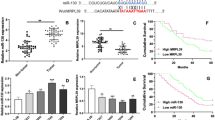
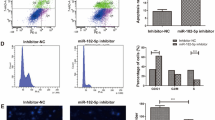
The malignant progression of oral squamous cell carcinoma (OSCC) has been confirmed to be mediated by a variety of factors, including circular RNA (circRNA). However, the role of circLPAR3 in OSCC development is still unclear. 70 paired OSCC tissues and normal control tissues were obtained from 70 OSCC patients. Quantitative real-time PCR was used to detect the expression of circLPAR3, microRNA (miR)-643, and high-mobility group box 2 (HMGB2). Cell proliferation, apoptosis, metastasis and stemness were assessed using cell counting kit 8 assay, colony-formation assay, flow cytometry, transwell assay and sphere formation assay. Marker protein expression and HMGB2 protein expression were determined by western blot analysis. The interaction between miR-643 and circLPAR3 or HMGB2 was confirmed by RNA pull-down assay, dual-luciferase reporter and RIP assay. The role of circLPAR3 in OSCC tumorigenesis was explored by constructing the xenograft models. Our data showed that circLPAR3 was highly expressed in OSCC tissues and cells. CircLPAR3 silencing suppressed OSCC cell proliferation, metastasis and stemness, while promoted apoptosis. On the mechanism, we discovered that circLPAR3 could sponge miR-643 to positive regulate HMGB2. MiR-643 overexpression had an inhibition effect on OSCC progression, and its inhibitor could reverse the negative regulation of circLPAR3 knockdown on OSCC progression. In addition, overexpressed HMGB2 also reversed the suppressive effect of circLPAR3 silencing on OSCC progression. Animal experiments results showed that downregulated circLPAR3 repressed OSCC tumorigenesis in vivo. Taken together, our data showed that circLPAR3 contributed to OSCC malignant progression through regulating the miR-643/HMGB2 axis.








Similar content being viewed by others
References
Bossi P, Alfieri S, Strojan P, Takes RP, Lopez F, Makitie A et al (2019) Prognostic and predictive factors in recurrent and/or metastatic head and neck squamous cell carcinoma: a review of the literature. Crit Rev Oncol Hematol 137:84–91
Chang J, Tan W, Ling Z, Xi R, Shao M, Chen M et al (2017) Genomic analysis of oesophageal squamous-cell carcinoma identifies alcohol drinking-related mutation signature and genomic alterations. Nat Commun 8:15290
Chen X, Yu J, Tian H, Shan Z, Liu W, Pan Z et al (2019) Circle RNA hsa_circRNA_100290 serves as a ceRNA for miR-378a to regulate oral squamous cell carcinoma cells growth via Glucose transporter-1 (GLUT1) and glycolysis. J Cell Physiol 234(11):19130–19140
Cheng H, Jiang W, Song Z, Li T, Li Y, Zhang L et al (2020) Circular RNA circLPAR3 facilitates esophageal squamous cell carcinoma progression through upregulating HMGB1 via sponging miR-375/miR-433. Onco Targets Ther 13:7759–7771
Cui G, Cai F, Ding Z, Gao L (2019) HMGB2 promotes the malignancy of human gastric cancer and indicates poor survival outcome. Hum Pathol 84:133–141
Fridman E, Na’ara S, Agarwal J, Amit M, Bachar G, Villaret AB et al (2018) The role of adjuvant treatment in early-stage oral cavity squamous cell carcinoma: an international collaborative study. Cancer 124(14):2948–2955
Gamez ME, Kraus R, Hinni ML, Moore EJ, Ma DJ, Ko SJ et al (2018) Treatment outcomes of squamous cell carcinoma of the oral cavity in young adults. Oral Oncol 87:43–48
Gao L, Zhao C, Li S, Dou Z, Wang Q, Liu J et al (2019) Circ-PKD2 inhibits carcinogenesis via the miR-204-3p/APC2 axis in oral squamous cell carcinoma. Mol Carcinog 58(10):1783–1794
Hamakawa H, Nakashiro K, Sumida T, Shintani S, Myers JN, Takes RP et al (2008) Basic evidence of molecular targeted therapy for oral cancer and salivary gland cancer. Head Neck 30(6):800–809
He ZH, Guo F, Hu XX, Luo ZY, Yi JW (2020) Knockdown of HMGB2 inhibits proliferation and invasion of renal tumor cells via the p-38MAPK pathway. Eur Rev Med Pharmacol Sci 24(9):4729–4737
Lei B, Tian Z, Fan W, Ni B (2019) Circular RNA: a novel biomarker and therapeutic target for human cancers. Int J Med Sci 16(2):292–301
Liang C, Wang J, Liu A, Wu Y (2021) Tumor promoting long non-coding RNA CASC15 affects HMGB2 expression by sponging miR-582–5p in colorectal cancer. J Gene Med 4:e3308
Lin X, Chen Y (2018) Identification of potentially functional CircRNA-miRNA-mRNA regulatory network in hepatocellular carcinoma by integrated microarray analysis. Med Sci Monit Basic Res 24:70–78
Liu L, Chen J, Cai X, Yao Z, Huang J (2019) Progress in targeted therapeutic drugs for oral squamous cell carcinoma. Surg Oncol 31:90–97
Lopez-Rosas I, Lopez-Camarillo C, Salinas-Vera YM, Hernandez-de la Cruz ON, Palma-Flores C, Chavez-Munguia B et al (2018) Entamoeba histolytica up-regulates MicroRNA-643 to promote apoptosis by targeting XIAP in human epithelial colon cells. Front Cell Infect Microbiol 8:437
McCauley MJ, Zimmerman J, Maher LJ 3rd, Williams MC (2007) HMGB binding to DNA: single and double box motifs. J Mol Biol 374(4):993–1004
Meng Y, Zhao EY, Zhou Y, Qiang DX, Wang S, Shi L et al (2020) Circular RNA hsa_circ_0011946 promotes cell growth, migration, and invasion of oral squamous cell carcinoma by upregulating PCNA. Eur Rev Med Pharmacol Sci 24(3):1226–1232
Panarese I, Aquino G, Ronchi A, Longo F, Montella M, Cozzolino I et al (2019) Oral and oropharyngeal squamous cell carcinoma: prognostic and predictive parameters in the etiopathogenetic route. Expert Rev Anticancer Ther 19(2):105–119
Shao Y, Song Y, Xu S, Li S, Zhou H (2020) Expression profile of circular RNAs in oral squamous cell carcinoma. Front Oncol 10:533616
Sheng R, Li X, Wang Z, Wang X (2020) Circular RNAs and their emerging roles as diagnostic and prognostic biomarkers in ovarian cancer. Cancer Lett 473:139–147
Shi Y, Fang N, Li Y, Guo Z, Jiang W, He Y et al (2020) Circular RNA LPAR3 sponges microRNA-198 to facilitate esophageal cancer migration, invasion, and metastasis. Cancer Sci 111(8):2824–2836
Su W, Shen Y, Wang Y, Wang F, Hong X, Chen Y et al (2021) CircPHIP promotes oral squamous cell carcinoma progression by sponging miR-142-5p and regulating PHIP and ACTN4 expression. Mol Ther Nucleic Acids 23:185–199
Syed N, Chavan S, Sahasrabuddhe NA, Renuse S, Sathe G, Nanjappa V et al (2015) Silencing of high-mobility group box 2 (HMGB2) modulates cisplatin and 5-fluorouracil sensitivity in head and neck squamous cell carcinoma. Proteomics 15(2–3):383–393
Wang H, Xing D, Ren D, Feng W, Chen Y, Zhao Z et al (2017) MicroRNA643 regulates the expression of ZEB1 and inhibits tumorigenesis in osteosarcoma. Mol Med Rep 16(4):5157–5164
Wu ZB, Cai L, Lin SJ, Xiong ZK, Lu JL, Mao Y et al (2013) High-mobility group box 2 is associated with prognosis of glioblastoma by promoting cell viability, invasion, and chemotherapeutic resistance. Neuro Oncol 15(9):1264–1275
Xiao M, Song H, You Y, Liu M, Yang X, Wang Y (2021) Metastasis of oral squamous cell carcinoma to the parotid lymph nodes. Int J Oral Maxillofac Surg 50(4):437–443
Xu F, Ni M, Li J, Cheng J, Zhao H, Zhao J et al (2020) Circ0004390 promotes cell proliferation through sponging miR-198 in ovarian cancer. Biochem Biophys Res Commun 526(1):14–20
Yang G, Zhang Y, Yang J (2019) Identification of potentially functional CircRNA-miRNA-mRNA regulatory network in gastric carcinoma using bioinformatics analysis. Med Sci Monit 25:8777–8796
Yang Y, Ci HS, Mao YL, Li JW, Zuo JH (2020) CircRNA_002178 promotes the proliferation and migration of oral squamous cell carcinoma cells by activating the Akt/mTOR pathway. Eur Rev Med Pharmacol Sci 24(11):6122–6130
Yue P, Rong X, Zhuang X, Sha HJ, Li JM, Xin L et al (2014) Cloning and expression analysis of a novel high-mobility group box 2 homologue from Lampetra japonica. Fish Physiol Biochem 40(2):625–634
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Fu, Y., Qiu, C., Yang, Y. et al. CircLPAR3 Acts as an Oncogene in Oral Squamous Cell Carcinoma Through Regulating the miR-643/HMGB2 Network. Biochem Genet 60, 882–898 (2022). https://doi.org/10.1007/s10528-021-10134-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-021-10134-y