Abstract
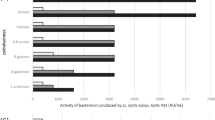
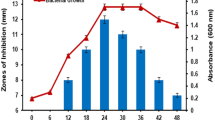
The focus of the present study was to characterize antimicrobial peptide produced by potential probiotic cultures of Enterococcus durans DB-1aa (MCC4243), Lactiplantibacillus plantarum Cu2-PM7 (MCC4246) and Limosilactobacillus fermentum Cu3-PM8 (MCC4233) against Staphylococus aureus MTCC 96 and Escherichia coli MTCC118. The growth kinetic assay revealed 24 h of incubation to be optimum for bacteriocin production. The partially purified compound of all the three selected cultures after ion-exchange chromatography was found to be thermoresistant and stable under a wide range of pH. The compound was sensitive to proteinase-K, but resistant to trypsin, α-amylase and lipase. Comparatively, bacteriocins from L. fermentum Cu3-PM8 and L. plantarum Cu2-PM7 showed higher stability under studied parameter, hence was taken up for further investigation. The apparent molecular weight of bacteriocin from L. fermentum MCC4233 and L. plantarum MCC4246 was found to be 3.5 kDa. Further, plantaricin gene from MCC4246 was characterized in silico. The translated partial amino acid sequence of the plnA gene in MCC4246 displayed 48 amino acids showing 100 % similarity with plantaricin A of Lactobacillus plantarum (WP_0036419). The sequence revealed 7 β sheets, 6 α sheets, 6 predicted coils and 9 predicted turns. The predicted properties of the peptide included an isoelectric point of 10.82 and a hydrophobicity of 48.6 %. The molecular approach of using Geneious Prime software and protein prediction data base for characterization of bacteriocin is novel and predicts “KSSAYSLQMGATAIKQVKKLFKKWGW” to be a peptide responsible for antimicrobial activity. The study provides information about a broad spectrum bacteriocin in native probiotic culture and paves a way towards its application in functional foods as a biopreservative agent.
Graphic abstract






Similar content being viewed by others
Data availability
Data sharing not applicable to this article as no datasets were generated or analyzed during the current study.
References
Amortegui J, Rodriguez-Lopez A, Rodriguez D, Carrascal AK, Almeciga-Diaz CJ, Melendez ADP, Sanchez OF (2014) Characterization of a new bacteriocin from Lactobacillus plantarum LE5 and LE27 isolated from ensiled corn. Appl Biochem Biotechnol 172(7):3374–3389
Arief II, Budiman C, Jenie BSL, Andreas E, Yuneni A (2015) Plantaricin IIA-1A5 from Lactobacillus plantarum IIA-1A5 displays bactericidal activity against Staphylococcus aureus. Benef Microbes 6(4):603–613
Armstrong DR, Berrisford JM, Conroy MJ, Gutmanas A, Anyango S, Choudhary P, Clark AR, Dana JM, Deshpande M, Dunlop R, Gane P, Gáborová R, Gupta D, Haslam P, Koča J, Mak L, Mir S, Mukhopadhyay A, Nadzirin N, Nair S, Paysan-Lafosse T, Pravda L, Sehnal D, Salih O, Smart O, Tolchard J, Varadi M, Svobodova-Vařeková R, Zaki H, Kleywegt GJ, Velankar S (2020) PDBe: improved findability of macromolecular structure data in the PDB. Nucleic Acids Res 48(D1):D335–D343
Aureli P, Capurso L, Castellazzi AM, Clerici M, Giovannini M, Morelli L, Poli A, Pregliasco F, Salvini F, Zuccotti GV (2011) Probiotics and health: an evidence-based review. Pharmacl Res 63(5):366–376
Barbosa MS, Todorov SD, Belguesmia Y, Choiset Y, Rabesona H, Ivanova IV, Chobert JM, Haertle T, Franco BDGM (2013) Purification and characterization of the bacteriocin produced by Lactobacillus sakei MBSa1 isolated from Brazilian salami. J Appl Microbiol 116:1195–1208
Barboza-Corona JE, Vazquez-Acosta H, Bideshi DK, Salcedo-Hernandez R (2007) Bacteriocin-like inhibitor substances produced by Mexican strains of Bacillus thuringiensis. Arch Microbiol 187:117–1126
Bello OO, Babalola OO, Adegboye M, Fashola MO, Bello TK (2018) Partial purification, characterization and application of bacteriocin from bacteria isolated Parkia biglobosa seed. NE Sci 3(2):72–94
Ben Omar N, Abriouel H, Keleke S, Sanchez Valenzuela A, Martinez-Canamero M, Lucas Lopez R, Ortega E, Galvez A (2008) Bacteriocin producing Lactobacillus strains isolated from poto poto, a Congolese fermented maize product, and genetic fingerprinting of their plantaricin operons. Int J Food Microbiol 30:18–25
Bindu A, Lakshmidevi N (2020) Identification and in vitro evaluation of probiotic attributes of lactic acid bacteria isolated from fermented food sources. Arch Microbiol 203(2):579–595
Cintas L, Herranz C, Hernández P, Casaus M, Nes L (2001) Review: bacteriocins of lactic acid bacteria. Food Sci Technol Int 7:281–305
Cotter PD, Ross RP, Hill C (2013) Bacteriocins-a viable alternative to antibiotics? Nature Rev Microbiol 11:95–105
Croxen MA, Law RJ, Scholz R, Keeney KM, Wlodarska M, Finlay BB (2013) Recent Advances in understanding enteric pathogenic Escherichia coli. Clin Microbiol Rev 26(4):822–880
Devi SM, Halami PM (2019) Genetic variation of pln loci among probiotic Lactobacillus plantarum group strains with antioxidant and cholesterol lowering ability. Probiot Antimicrob Proteins 11:11–22
Diep DB, Havarstein LS, Nissen-Meyer, Nes IF (1994) The gene encoding plantaricin A, a bacteriocin from Lactobacillus plantarum C11, is located on the same transcription unit as an agr-like regulatory system. Appl Environ Microbiol 60:160–166
Diep DB, Havarstein LS, Nes IF (1996) Characterization of the locus responsible for the bacteriocin production in Lactobacillus plantarum C11. J Bacteriol 178:4472–4483
Diep DB, Straume D, Kjos M, Torres C, Nes IF (2009) An overview of the mosaic bacteriocin pln loci from Lactobacillus plantarum. Peptides 30:1562–1574
Drider D, Fimland G, Héchard Y, McMullen LM, Prevost H (2006) The continuing story of class IIa bacteriocins. Microbiol Mol Biol Rev 70:564–582
Ferchichi M, Frère J, Mabrouk K, Manai M (2001) Lactococcin MMFII, a novel class IIa bacteriocin produced by Lactococcus lactis MMFII, isolated from a Tunisian dairy product. FEMS Microbiol Lett 205:49–55
Hernández D, Cardell E, Zárate V (2005) Antimicrobial activity of lactic acid bacteria isolated from Tenerife cheese: initial characterization of plantaricin TF711, a bacteriocin-like substance produced by Lactobacillus plantarum TF711. J Appl Microbiol 99:77–84
Kadariya J, Smith TC, Thapaliya D (2014) Staphylococcus aureusemclose and Staphylococcal food-borne disease: an ongoing challenge in public health. Emerg Microb Concerns Food Saf New Control Meas. https://doi.org/10.1155/2014/827965
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Kristiansen PE, Fimland G, Mantzilas D, Nissen-Meyer (2005) Structure and mode of action of the membrane-permeabilizing antimicrobial peptide pheromone Plantaricin A. J Biol Chem 280(4):22945–22950
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lee NK, Paik HD (2001) Partial characterization of lactocin NK24, a newly identified bacteriocin of Lactococcus lactis NK24 isolated from Jeot-gal. Food Microbiol 18:17–24
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the folin phenol reagent. J Biol Chem 193:265–275
Maldonado A, Jimenez-Diaz R, Ruiz-Barba JL (2004) Induction of plantaricin production in Lactobacillus plantarum NC8 after co-culture with specific gram positive bacteria is mediated by an autoinduction mechanism. J Bacteriol 86:1556–1564
Martinez RC, Wachsman M, Torres NI, LeBlanc JG, Todorov SD, Franco BD (2013) Biochemical, antimicrobial and molecular characterization of a noncytotoxic bacteriocin produced by Lactobacillus plantarum ST71KS. Food Microbiol 34:376–381
Messi P, Bondi M, Sabia C, Battini R, Manicardi G (2001) Detection and preliminary characterization of a bacteriocin (plantaricin 35d) produced by a Lactobacillus plantarum strain. Int J Food Microbiol 64(1–2):93–198
Milioni C, Martínez B, Degl’Innocenti S, Turchi B, Fratini F, Cerri D, Fischetti R (2015) A novel bacteriocin produced by Lactobacillus plantarum LpU4 as a valuable candidate for biopreservation in artisanal raw milk cheese. Dairy Sci Technol 95:479–494
Mora D, Fortina MG, Parini C, Daffonchio D, Manachini PL (2000) Genomic subpopulations within the species Pediococcus acidilactici detected by multilocus typing analysis: relationship between pediocin AcHPA-1 producing and non-producing strains. Microbiology 146:2027–2038
Mourad K, Halima ZK, Nour-Eddine K (2005) Detection and activity of plantaricin OL15 a bacteriocin produced by Lactobacillus plantarum OL15 isolated from Algerian fermented olives. Grasas Aceites 56:192–197
Nes IF, Holo H (2000) Class II antimicrobial peptides from lactic acid bacteria. Biopolymers 55:50–61
Noda M, Miyauchi R, Danshiitsoodol N, Matoba Y, Kumagai T, Sugiyama M (2018) Expression of genes involved in bacteriocin production and self-resistance in Lactobacillus brevis 174A is mediated by two regulatory proteins. Appl Environ Microbiol 84:e02707–e02717
Nowroozi J, Mirzaii M, Norouzi M (2004) Study of Lactobacillus as probiotic bacteria. Iranian J Public Health 33:1–7
Remiger A, Ehrmann MA, Vogel RF (1996) Identification of bacteriocin-encoding genes in Lactobacilli by polymerase chain reaction (PCR). System Appl Microbiol 19:28–34
Riley MA, Wertz JE (2002) Bacteriocins: evolution, ecology and application. Ann Rev Microbiol 56:117–137
Rodrigues E, Calzada J, Arques JK, Rodriguez JM, Nunez M, Medina M (2005) Antimicrobial activity of pediocin-producing Lactococcus lactis on Listeria monocytogenes, Staphylococcus aureus and Escherichia coli O157:H7 in cheese. Int Dairy J 5:51–57
Rojo-Bezares B, Sáenz Y, Navarro L, Jiménez-Díaz R, Zarazaga M, Ruiz-Larrea F, Torres C (2008) Characterization of a new organization of the plantaricin locus in the inducible bacteriocin-producing Lactobacillus plantarum J23 of grape must origin. Arch Microbiol 189:491–499
Song DF, Zhu MY, Gu Q (2014) Purification and characterization of plantaricin ZJ5, a new bacteriocin produced by Lactobacillus plantarum ZJ5. PLoS ONE 9(8):e105549
Sowani HM, Thorat P (2012) Antimicrobial activity studies of bactoriocin produced by Lactobacilli isolates from carrot kanji. Online J Biol Sci 12:6–10
Todorov SD (2009) Bacteriocins from Lactobacillus plantarum-production, genetic organization and mode of action. Braz J Microbiol 40:209–221
Todorov SD, Dicks LMT (2005) Lactobacillus plantarum isolated from molasses produces bacteriocins active against Gram-negative bacteria. Enz Microbial Technol 36:318–326
Todorov SD, Dicks LMT (2006) Screening for bacteriocin-producing lactic acid bacteria from boza, a traditional cereal beverage from Bulgaria: comparison of the bacteriocins. Process Biochem 46:11–19
Todorov SD, van Reenen CA, Dicks LM (2004) Optimization of bacteriocin production by Lactobacillus plantarum ST13BR, a strain isolated from barley beer. J Gen Appl Microbiol 50:49–157
Todorov SD, Ho P, Vaz-Velho M, Dicks LM (2010) Characterization of bacteriocins produced by two strains of Lactobacillus plantarum isolated from Beloura and Chouriço, traditional pork products from Portugal. Meat Sci 84:334–343
Tomé E, Todorov SD, Gibbs PA, Teixeira PC (2009) Partial characterization of nine bacteriocins produced by lactic acid bacteria isolated from cold-smoked salmon with activity against Listeria monocytogenes. Food Biotechnol 23:50–73
Xie J, Zhang R, Shang C, Guo Y (2009) Isolation and characterization of a bacteriocin produced by an isolated Bacillus subtilis LFB112 that exhibits antimicrobial activity against domestic animal pathogens. Afr J Biotechnol 8(20):5611–5619
Zapata S, Muñoz J, Ruiz O, Montoya O, Gutierrez P (2009) Isolation of Lactobacillus plantarum LPBM10 and partial characterization of its bacteriocin. Vitae 16:75–82
Zheng J, Wittouck S, Salvetti E, Franz CMAP, Harris HMB, Mattarelli P, O’Toole PW, Pot B, Vandamme P, Walter J, Watanabe K, Wuyts S, Felis GE, Gänzle MG, Lebeer S (2020) A taxonomic note on the genus Lactobacillus: description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. Int J Syst Evol Microbiol 70:2782–2858
Acknowledgements
This work was support by the University Grants Commission for the award of a NFSC research fellowship.
Author information
Authors and Affiliations
Contributions
NL and AB conceived and designed the experiments. AB conducted the experiments and wrote the manuscript. Both the authors read and approved the manuscript for submission.
Corresponding author
Ethics declarations
Conflict of interest
Dr. Lakshmidevi and Mrs. Amrutha Bindu declare that they have no conflict of interest.
Consent for publication
This manuscript does not contain any individual person ‘s data.
Ethical approval
This article does not contain any studies with humans or animals.
Informed consent
Informed consent was obtained from both the authors of the study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bindu, A., Lakshmidevi, N. In vitro and in silico approach for characterization of antimicrobial peptides from potential probiotic cultures against Staphylococcus aureus and Escherichia coli. World J Microbiol Biotechnol 37, 172 (2021). https://doi.org/10.1007/s11274-021-03135-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-021-03135-x