Abstract
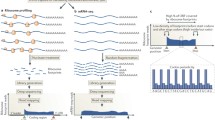
Ribosome profiling (riboseq) has opened the possibilities for the genome-wide studies of translation in all living organisms. This method is based on deep sequencing of mRNA fragments protected by the ribosomes from hydrolysis by ribonucleases, the so-called ribosomal footprints (RFPs). Ribosomal profiling together with RNA sequencing allows not only to identify with a reasonable accuracy translated reading frames in the transcriptome, but also to track changes in gene expression in response to various stimuli. Notably, ribosomal profiling in its classical version has certain limitations. The size of the selected mRNA fragments is 25-35 nts, while RFPs of other sizes are usually omitted from analysis. Also, ribosomal profiling “averages” the data from all ribosomes and does not allow to study specific ribosomal complexes associated with particular translation factors. However, recently developed modifications of ribosomal profiling provide answers to a number of questions. Thus, it has become possible to analyze not only elongating, but also scanning and reinitiating ribosomes, to study events associated with the collision of ribosomes during mRNA translation, to discover new ways of cotranslational assembly of multisubunit protein complexes during translation, and to selectively isolate ribosomal complexes associated with certain protein factors. New data obtained using these modified approaches provide a better understanding of the mechanisms of translation regulation and the functional roles of translational apparatus components.



Similar content being viewed by others
Abbreviations
- AMP-PNP:
-
non-hydrolysable analogue of ATP
- eIF:
-
eukaryotic initiation factor
- ORF:
-
open reading frame
- PIC:
-
preinitiation complex
- RFP:
-
ribosomal footprint
- RSC:
-
ribosomal scanning complex
- RQC:
-
quality control on translating ribosomes
- UTR:
-
untranslated region
References
Castles, J. J., and Singer, M. F. (1969) Degradation of polyuridylic acid by ribonuclease II: protection by ribosomes, J. Mol. Biol., 40, 1-17, https://doi.org/10.1016/0022-2836(69)90292-7.
Wolin, S. L., and Walter, P. (1988) Ribosome pausing and stacking during translation of a eukaryotic mRNA, EMBO J., 7, 3559-3569.
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R., and Weissman, J. S. (2009) Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling, Science, 324, 218-223, https://doi.org/10.1126/science.1168978.
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M., and Weissman, J. S. (2012) The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments, Nat. Prot., 7, 1534-1550, https://doi.org/10.1038/nprot.2012.086.
Lee, S., Liu, B., Lee, S., Huang, S. X., Shen, B., and Qian, S. B. (2012) Global mapping of translation initiation sites in mammalian cells at single-nucleotide resolution, Proc. Natl. Acad. Sci. USA, 109, E2424-2432, https://doi.org/10.1073/pnas.1207846109.
Ingolia, N. T., Lareau, L. F., and Weissman, J. S. (2011) Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes, Cell, 147, 789-802, https://doi.org/10.1016/j.cell.2011.10.002.
Wu, C. C., Zinshteyn, B., Wehner, K. A., and Green, R. (2019) High-resolution ribosome profiling defines discrete ribosome elongation states and translational regulation during cellular stress, Mol. Cell, 73, 959-970.e955, https://doi.org/10.1016/j.molcel.2018.12.009.
Lareau, L. F., Hite, D. H., Hogan, G. J., and Brown, P. O. (2014) Distinct stages of the translation elongation cycle revealed by sequencing ribosome-protected mRNA fragments, eLife, 3, e01257, https://doi.org/10.7554/eLife.01257.
Ingolia, N. T., Hussmann, J. A., and Weissman, J. S. (2019) Ribosome profiling: global views of translation, Cold Spring Harb. Perspect. Biol., 11, a032698, https://doi.org/10.1101/cshperspect.a032698.
Brar, G. A., and Weissman, J. S. (2015) Ribosome profiling reveals the what, when, where and how of protein synthesis, Nat. Rev. Mol. Cell Biol., 16, 651-664, https://doi.org/10.1038/nrm4069.
Kiniry, S. J., Michel, A. M., and Baranov, P. V. (2020) Computational methods for ribosome profiling data analysis, Wiley Interdiscip. Rev. RNA, 11, e1577, https://doi.org/10.1002/wrna.1577.
Ingolia, N. T. (2014) Ribosome profiling: new views of translation, from single codons to genome scale, Nat. Rev. Genet., 15, 205-213, https://doi.org/10.1038/nrg3645.
Andreev, D. E., O’Connor, P. B., Loughran, G., Dmitriev, S. E., Baranov, P. V., and Shatsky, I. N. (2017) Insights into the mechanisms of eukaryotic translation gained with ribosome profiling, Nucleic Acids Res., 45, 513-526, https://doi.org/10.1093/nar/gkw1190.
Calviello, L., and Ohler, U. (2017) Beyond read-counts: Ribo-seq data analysis to understand the functions of the transcriptome, Trends Genet., 33, 728-744, https://doi.org/10.1016/j.tig.2017.08.003.
Michel, A. M., and Baranov, P. V. (2013) Ribosome profiling: a Hi-Def monitor for protein synthesis at the genome-wide scale, Wiley Interdiscip. Rev. RNA, 4, 473-490, https://doi.org/10.1002/wrna.1172.
Kozak, M. (1978) How do eucaryotic ribosomes select initiation regions in messenger RNA? Cell, 15, 1109-1123, https://doi.org/10.1016/0092-8674(78)90039-9.
Kozak, M. (1980) Evaluation of the “scanning model” for initiation of protein synthesis in eucaryotes, Cell, 22, 7-8, https://doi.org/10.1016/0092-8674(80)90148-8.
Hinnebusch, A. G. (2014) The scanning mechanism of eukaryotic translation initiation, Annu. Rev. Biochem., 83, 779-812, https://doi.org/10.1146/annurev-biochem-060713-035802.
Hinnebusch, A. G. (2017) Structural insights into the mechanism of scanning and start codon recognition in eukaryotic translation initiation, Trends Biochem. Sci., 42, 589-611, https://doi.org/10.1016/j.tibs.2017.03.004.
Tahmasebi, S., Sonenberg, N., Hershey, J. W. B., and Mathews, M. B. (2019) Protein synthesis and translational control: a historical perspective, Cold Spring Harb. Perspect. Biol., 11, a035584, https://doi.org/10.1101/cshperspect.a035584.
Hershey, J. W. B., Sonenberg, N., and Mathews, M. B. (2019) Principles of translational control, Cold Spring Harb. Perspect. Biol., 11, a032607, https://doi.org/10.1101/cshperspect.a032607.
Hinnebusch, A. G., Ivanov, I. P., and Sonenberg, N. (2016) Translational control by 5′-untranslated regions of eukaryotic mRNAs, Science, 352, 1413-1416, https://doi.org/10.1126/science.aad9868.
Valasek, L., Szamecz, B., Hinnebusch, A. G., and Nielsen, K. H. (2007) In vivo stabilization of preinitiation complexes by formaldehyde cross-linking, Methods Enzymol., 429, 163-183, https://doi.org/10.1016/S0076-6879(07)29008-1.
Archer, S. K., Shirokikh, N. E., Beilharz, T. H., and Preiss, T. (2016) Dynamics of ribosome scanning and recycling revealed by translation complex profiling, Nature, 535, 570-574, https://doi.org/10.1038/nature18647.
Pisarev, A. V., Kolupaeva, V. G., Yusupov, M. M., Hellen, C. U., and Pestova, T. V. (2008) Ribosomal position and contacts of mRNA in eukaryotic translation initiation complexes, EMBO J., 27, 1609-1621, https://doi.org/10.1038/emboj.2008.90.
Kozak, M. (1977) Nucleotide sequences of 5′-terminal ribosome-protected initiation regions from two reovirus messages, Nature, 269, 391-394, https://doi.org/10.1038/269390a0.
Lazarowitz, S. G., and Robertson, H. D. (1977) Initiator regions from the small size class of reovirus messenger RNA protected by rabbit reticulocyte ribosomes, J. Biol. Chem., 252, 7842-7849, https://doi.org/10.1016/S0021-9258(17)41043-X.
Bohlen, J., Fenzl, K., Kramer, G., Bukau, B., and Teleman, A. A. (2020) Selective 40S footprinting reveals cap-tethered ribosome scanning in human cells, Mol. Cell, 79, 561-574.e565, https://doi.org/10.1016/j.molcel.2020.06.005.
Wagner, S., Herrmannova, A., Hronova, V., Gunisova, S., Sen, N. D., et al. (2020) Selective translation complex profiling reveals staged initiation and co-translational assembly of initiation factor complexes, Mol. Cell, 79, 546-560.e547, https://doi.org/10.1016/j.molcel.2020.06.004.
Giess, A., Torres Cleuren, Y. N., Tjeldnes, H., Krause, M., Bizuayehu, T. T., et al. (2020) Profiling of small ribosomal subunits reveals modes and regulation of translation initiation, Cell Rep., 31, 107534, https://doi.org/10.1016/j.celrep.2020.107534.
Elfakess, R., Sinvani, H., Haimov, O., Svitkin, Y., Sonenberg, N., and Dikstein, R. (2011) Unique translation initiation of mRNAs-containing TISU element, Nucleic Acids Res., 39, 7598-7609, https://doi.org/10.1093/nar/gkr484.
Kumar, P., Hellen, C. U., and Pestova, T. V. (2016) Toward the mechanism of eIF4F-mediated ribosomal attachment to mammalian capped mRNAs, Genes Dev., 30, 1573-1588, https://doi.org/10.1101/gad.282418.116.
Meyuhas, O., and Kahan, T. (2015) The race to decipher the top secrets of TOP mRNAs, Biochim. Biophys. Acta, 1849, 801-811, https://doi.org/10.1016/j.bbagrm.2014.08.015.
Tamarkin-Ben-Harush, A., Vasseur, J. J., Debart, F., Ulitsky, I., and Dikstein, R. (2017) Cap-proximal nucleotides via differential eIF4E binding and alternative promoter usage mediate translational response to energy stress, eLife, 6, e21907, https://doi.org/10.7554/eLife.21907.
Shatsky, I. N., Terenin, I. M., Smirnova, V. V., and Andreev, D. E. (2018) Cap-independent translation: what’s in a name? Trends Biochem. Sci., 43, 882-895, https://doi.org/10.1016/j.tibs.2018.04.011.
Oh, E., Becker, A. H., Sandikci, A., Huber, D., Chaba, R., et al. (2011) Selective ribosome profiling reveals the cotranslational chaperone action of trigger factor in vivo, Cell, 147, 1295-1308, https://doi.org/10.1016/j.cell.2011.10.044.
Lin, Y., Li, F., Huang, L., Polte, C., Duan, H., Fang, J., et al. (2020) eIF3 associates with 80S ribosomes to promote translation elongation, mitochondrial homeostasis, and muscle health, Mol. Cell, 79, 575-587.e577, https://doi.org/10.1016/j.molcel.2020.06.003.
Hashem, Y., des Georges, A., Dhote, V., Langlois, R., Liao, H. Y., et al. (2013) Structure of the mammalian ribosomal 43S preinitiation complex bound to the scanning factor DHX29, Cell, 153, 1108-1119, https://doi.org/10.1016/j.cell.2013.04.036.
Des Georges, A., Dhote, V., Kuhn, L., Hellen, C. U., Pestova, T. V., et al. (2015) Structure of mammalian eIF3 in the context of the 43S preinitiation complex, Nature, 525, 491-495, https://doi.org/10.1038/nature14891.
Eliseev, B., Yeramala, L., Leitner, A., Karuppasamy, M., Raimondeau, E., et al. (2018) Structure of a human cap-dependent 48S translation pre-initiation complex, Nucleic Acids Res., 46, 2678-2689, https://doi.org/10.1093/nar/gky054.
Brito Querido, J., Sokabe, M., Kraatz, S., Gordiyenko, Y., Skehel, J. M., et al. (2020) Structure of a human 48S translational initiation complex, Science, 369, 1220-1227, https://doi.org/10.1126/science.aba4904.
Valasek, L. S., Zeman, J., Wagner, S., Beznoskova, P., Pavlikova, Z., et al. (2017) Embraced by eIF3: structural and functional insights into the roles of eIF3 across the translation cycle, Nucleic Acids Res., 45, 10948-10968, https://doi.org/10.1093/nar/gkx805.
Mohammad, M. P., Munzarova Pondelickova, V., Zeman, J., Gunisova, S., and Valasek, L. S. (2017) In vivo evidence that eIF3 stays bound to ribosomes elongating and terminating on short upstream ORFs to promote reinitiation, Nucleic Acids Res., 45, 2658-2674, https://doi.org/10.1093/nar/gkx049.
Benitez-Cantos, M. S., Yordanova, M. M., O’Connor, P. B. F., Zhdanov, A. V., Kovalchuk, S. I., et al. (2020) Translation initiation downstream from annotated start codons in human mRNAs coevolves with the Kozak context, Genome Res., 30, 974-984, https://doi.org/10.1101/gr.257352.119.
Fresno, M., Jimenez, A., and Vazquez, D. (1977) Inhibition of translation in eukaryotic systems by harringtonine, Eur. J. Biochem., 72, 323-330, https://doi.org/10.1111/j.1432-1033.1977.tb11256.x.
Shirokikh, N. E., Dutikova, Y. S., Staroverova, M. A., Hannan, R. D., and Preiss, T. (2019) Migration of small ribosomal subunits on the 5′-untranslated regions of capped messenger RNA, Int. J. Mol. Sci., 20, 4464, https://doi.org/10.3390/ijms20184464.
Doring, K., Ahmed, N., Riemer, T., Suresh, H. G., Vainshtein, Y., et al. (2017) Profiling Ssb-nascent chain interactions reveals principles of Hsp70-assisted folding, Cell, 170, 298-311.e220, https://doi.org/10.1016/j.cell.2017.06.038.
Le Tallec, B., Barrault, M. B., Courbeyrette, R., Guerois, R., Marsolier-Kergoat, M. C., and Peyroche, A. (2007) 20S proteasome assembly is orchestrated by two distinct pairs of chaperones in yeast and in mammals, Mol. Cell, 27, 660-674, https://doi.org/10.1016/j.molcel.2007.06.025.
Rosenzweig, R., and Glickman, M. H. (2008) Chaperone-driven proteasome assembly, Biochem. Soc. Trans., 36, 807-812, https://doi.org/10.1042/BST0360807.
Shiber, A., Doring, K., Friedrich, U., Klann, K., Merker, D., et al. (2018) Cotranslational assembly of protein complexes in eukaryotes revealed by ribosome profiling, Nature, 561, 268-272, https://doi.org/10.1038/s41586-018-0462-y.
Kamenova, I., Mukherjee, P., Conic, S., Mueller, F., El-Saafin, F., et al. (2019) Co-translational assembly of mammalian nuclear multisubunit complexes, Nat. Commun., 10, 1740, https://doi.org/10.1038/s41467-019-09749-y.
Guydosh, N. R., and Green, R. (2014) Dom34 rescues ribosomes in 3′-untranslated regions, Cell, 156, 950-962, https://doi.org/10.1016/j.cell.2014.02.006.
Han, P., Shichino, Y., Schneider-Poetsch, T., Mito, M., Hashimoto, S., et al. (2020) Genome-wide survey of ribosome collision, Cell Rep., 31, 107610, https://doi.org/10.1016/j.celrep.2020.107610.
Brown, A., Shao, S., Murray, J., Hegde, R. S., and Ramakrishnan, V. (2015) Structural basis for stop codon recognition in eukaryotes, Nature, 524, 493-496, https://doi.org/10.1038/nature14896.
McCaughan, K. K., Brown, C. M., Dalphin, M. E., Berry, M. J., and Tate, W. P. (1995) Translational termination efficiency in mammals is influenced by the base following the stop codon, Proc. Natl. Acad. Sci. USA, 92, 5431-5435, https://doi.org/10.1073/pnas.92.12.5431.
Zhao, T., Chen, Y. M., Li, Y., Wang, J., Chen, S., et al. (2021) Disome-seq reveals widespread ribosome collisions that promote cotranslational protein folding, Genome Biol., 22, 16, https://doi.org/10.1186/s13059-020-02256-0.
Arpat, A. B., Liechti, A., De Matos, M., Dreos, R., Janich, P., and Gatfield, D. (2020) Transcriptome-wide sites of collided ribosomes reveal principles of translational pausing, Genome Res., 30, 985-999, https://doi.org/10.1101/gr.257741.119.
Joazeiro, C. A. P. (2019) Mechanisms and functions of ribosome-associated protein quality control, Nat. Rev. Mol. Cell Biol., 20, 368-383, https://doi.org/10.1038/s41580-019-0118-2.
Ikeuchi, K., Izawa, T., and Inada, T. (2018) Recent progress on the molecular mechanism of quality controls induced by ribosome stalling, Front. Genet., 9, 743, https://doi.org/10.3389/fgene.2018.00743.
Joazeiro, C. A. P. (2017) Ribosomal stalling during translation: providing substrates for ribosome-associated protein quality control, Annu. Rev. Cell Dev. Biol., 33, 343-368, https://doi.org/10.1146/annurev-cellbio-111315-125249.
Defenouillere, Q., and Fromont-Racine, M. (2017) The ribosome-bound quality control complex: from aberrant peptide clearance to proteostasis maintenance, Curr. Genet., 63, 997-1005, https://doi.org/10.1007/s00294-017-0708-5.
Wu, C. C., Peterson, A., Zinshteyn, B., Regot, S., and Green, R. (2020) Ribosome collisions trigger general stress responses to regulate cell fate, Cell, 182, 404-416.e414, https://doi.org/10.1016/j.cell.2020.06.006.
Pochopien, A. A., Beckert, B., Kasvandik, S., Berninghausen, O., Beckmann, R., et al. (2021) Structure of Gcn1 bound to stalled and colliding 80S ribosomes, Proc. Natl. Acad. Sci. USA, 118, e2022756118, https://doi.org/10.1073/pnas.2022756118.
Meydan, S., and Guydosh, N. R. (2020) Disome and Trisome profiling reveal genome-wide targets of ribosome quality control, Mol. Cell, 79, 588-602.e586, https://doi.org/10.1016/j.molcel.2020.06.010.
Tuck, A. C., Rankova, A., Arpat, A. B., Liechti, L. A., Hess, D., et al. (2020) Mammalian RNA decay pathways are highly specialized and widely linked to translation, Mol. Cell, 77, 1222-1236.e1213, https://doi.org/10.1016/j.molcel.2020.01.007.
Young, D. J., Makeeva, D. S., Zhang, F., Anisimova, A. S., Stolboushkina, E. A., et al. (2018) Tma64/eIF2D, Tma20/MCT-1, and Tma22/DENR recycle post-termination 40S subunits in vivo, Mol. Cell, 71, 761-774.e765, https://doi.org/10.1016/j.molcel.2018.07.028.
Gaikwad, S., Ghobakhlou, F., Young, D. J., Visweswaraiah, J., Zhang, H., and Hinnebusch, A. G. (2021) Reprogramming of translation in yeast cells impaired for ribosome recycling favors short, efficiently translated mRNAs, eLife, 10, e64283, https://doi.org/10.7554/eLife.64283.
Castelo-Szekely, V., De Matos, M., Tusup, M., Pascolo, S., Ule, J., and Gatfield, D. (2019) Charting DENR-dependent translation reinitiation uncovers predictive uORF features and links to circadian timekeeping via clock, Nucleic Acids Res., 47, 5193-5209, https://doi.org/10.1093/nar/gkz261.
Schuller, A. P., Wu, C. C., Dever, T. E., Buskirk, A. R., and Green, R. (2017) eIF5A functions globally in translation elongation and termination, Mol. Cell, 66, 194-205.e195, https://doi.org/10.1016/j.molcel.2017.03.003.
Kasari, V., Margus, T., Atkinson, G. C., Johansson, M. J. O., and Hauryliuk, V. (2019) Ribosome profiling analysis of eEF3-depleted Saccharomyces cerevisiae, Sci. Rep., 9, 3037, https://doi.org/10.1038/s41598-019-39403-y.
Zhou, F., Zhang, H., Kulkarni, S. D., Lorsch, J. R., and Hinnebusch, A. G. (2020) eIF1 discriminates against suboptimal initiation sites to prevent excessive uORF translation genome-wide, RNA, 26, 419-438, https://doi.org/10.1261/rna.073536.119.
Fijalkowska, D., Verbruggen, S., Ndah, E., Jonckheere, V., Menschaert, G., and Van Damme, P. (2017) eIF1 modulates the recognition of suboptimal translation initiation sites and steers gene expression via uORFs, Nucleic Acids Res., 45, 7997-8013, https://doi.org/10.1093/nar/gkx469.
Sen, N. D., Zhou, F., Harris, M. S., Ingolia, N. T., and Hinnebusch, A. G. (2016) eIF4B stimulates translation of long mRNAs with structured 5′-UTRs and low closed-loop potential but weak dependence on eIF4G, Proc. Natl. Acad. Sci. USA, 113, 10464-10472, https://doi.org/10.1073/pnas.1612398113.
Young, D. J., Guydosh, N. R., Zhang, F., Hinnebusch, A. G., and Green, R. (2015) Rli1/ABCE1 recycles terminating ribosomes and controls translation reinitiation in 3′-UTRs in vivo, Cell, 162, 872-884, https://doi.org/10.1016/j.cell.2015.07.041.
Tang, L., Morris, J., Wan, J., Moore, C., Fujita, Y., et al. (2017) Competition between translation initiation factor eIF5 and its mimic protein 5MP determines non-AUG initiation rate genome-wide, Nucleic Acids Res., 45, 11941-11953, https://doi.org/10.1093/nar/gkx808.
Sugiyama, H., Takahashi, K., Yamamoto, T., Iwasaki, M., Narita, M., et al. (2017) Nat1 promotes translation of specific proteins that induce differentiation of mouse embryonic stem cells, Proc. Natl. Acad. Sci. USA, 114, 340-345, https://doi.org/10.1073/pnas.1617234114.
Funding
This work was supported by the Russian Science Foundation (project no. 19-14-00152).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare no conflict of interest in financial or any other sphere. This article does not contain description of studies with the involvement of humans or animal subjects performed by any of the authors.
Rights and permissions
About this article
Cite this article
Andreev, D.E., Smirnova, V.V. & Shatsky, I.N. Modifications of Ribosome Profiling that Provide New Data on the Translation Regulation. Biochemistry Moscow 86, 1095–1106 (2021). https://doi.org/10.1134/S0006297921090054
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297921090054