Abstract
Purpose
Bacterial quantification by qPCR is considered the gold standard for microbial molecular diagnosis. However, a fundamental pre-requisite in this methodology is the designing of specific primers for the bacterium of interest. With the increase in bacterial genome sequencing data in the recent years, it has become possible to design specific primers that can be used to quantify different strains of the same bacterial species.
Methods
To develop a real-time PCR (qPCR) protocol for the specific quantification of Azospirillum baldaniorum Sp245 strain (old Azospirillum brasilense), the Sp245 genome sequence was fragmented into small contigs with 500 base pairs each, and analyzed for similarity against the NCBI non-redundant database. A. baldaniorum-specific contigs were used to design the primers. The best pair of primers was used to quantify these bacteria after inoculation in different cultivars of Brachiaria, grown under field conditions.
Results
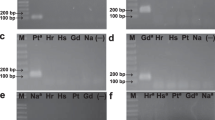
Our results showed that the primer pair Sp245p10 was highly specific for the Sp245 strain in the Brachiaria root and shoot field under different conditions. The qPCR assay using these primers showed differences among cultivars in the number of bacteria detected in plants after inoculation. Additionally, the number of bacteria observed in the roots was higher than that in the shoots.
Conclusion
The qPCR methodology using a Sp245 strain-specific primer may be used to monitor A. baldaniorum inoculated into other plants and may find potential application in field experiments.





Similar content being viewed by others
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. https://doi.org/10.1093/nar/25.17.3389
Antonio CS, Rouws LFM, Teixeira KRS, Reis VM (2016) Diazotrophic bacteria associated to sugarcane varieties cropped at northeast region of Brazil. Rev Bras Cienc Agra 11:272–280. https://doi.org/10.5039/agraria.v11i4a5393
Baldani JI, Baldani VLD (2005) History on the biological nitrogen fixation research in graminaceous plants: special emphasis on the Brazilian experience. An Acad Bras Ciênc 77:549–579. https://doi.org/10.1590/S0001-37652005000300014
Baldani VLD, Alvarez MAdB, Baldani JI et al (1986a) Establishment of inoculated Azospirillum spp. in the rhizosphere and in roots of field grown wheat and sorghum. Plant Soil 90:35–46. https://doi.org/10.1007/BF02277385
Baldani JI, Baldani VLD, Seldin L et al (1986b) Characterization of Herbaspirillurn seropedicae gen. nov., sp. nov., a Root- Associated Nitrogen-Fixing Bacterium. Int J Syst Evol Microbiol 36:86–93. https://doi.org/10.1099/00207713-36-1-86
Bishop CJ, Aanensen DM, Jordan GE et al (2009) Assigning strains to bacterial species via the internet BMC Biol 7:3. https://doi.org/10.1186/1741-7007-7-3
Boa Sorte PMF, Simoes-Araujo JL, de Melo LHV et al (2014) Development of a real-time PCR assay for the detection and quantification of Gluconacetobacter diazotrophicus in sugarcane grown under field conditions. Afric J Microbio Res 8:2937–2946. https://doi.org/10.5897/AJMR2014.6779
Braun-Kiewnick A, Lehmann A, Rezzonico F et al (2012) Development of species-, strain- and antibiotic biosynthesis-specific quantitative PCR assays for Pantoea agglomerans as tools for biocontrol monitoring. J Microbiol Meth 90:315–320. https://doi.org/10.1016/j.mimet.2012.06.004
Breda FAF, da Silva TFR, Dos Santos SG et al (2019) Modulation of nitrogen metabolism of maize plants inoculated with Azospirillum brasilense and Herbaspirillum seropedicae. Arch Microbiol 201:547–558. https://doi.org/10.1007/s00203-018-1594-z
Brunk CF, Li J, Avaniss-Aghajani E (2002) Analysis of specific bacteria from environmental samples using a quantitative polymerase chain reaction. Curr Issues in Mol Biol 4:13–18
Bulgarelli D, Schlaeppi K, Spaepen S et al (2013) Structure and functions of the bacterial microbiota of plants. A Rev Plant Biol 64:807–838. https://doi.org/10.1146/annurev-arplant-050312-120106
Camelo A, Barreto CP, Vidal MS et al (2021) Field response of two seed propagated elephant grass genotypes to diazotrophic bacterial inoculation and in situ confocal microscopy colonization analyses. Symbiosis 83:41–53. https://doi.org/10.1007/s13199-020-00730-8
Cassán F, Diaz-Zorita M (2016) Azospirillum sp. in current agriculture: from the laboratory to the field. Soil Biol Bioch 103:117–130. https://doi.org/10.1016/j.soilbio.2016.08.020
Cassán FD, Okon Y, Creus CM (2015) Handbook for Azospirillum: Technical Issues and Protocols. Springer. https://doi.org/10.1007/978-3-319-06542-7
Cassán F, Coniglio A, López G et al (2020) Everything you must know about Azospirillum and its impact on agriculture and beyond. Biol Fertil Soils 56:461–479. https://doi.org/10.1007/s00374-020-01463-y
Chaparro JM, Badri DV, Bakker MG et al (2013) Root exudation of phytochemicals in Arabidopsis follows specific patterns that are developmentally programmed and correlate with soil microbial functions. PLoS One 8:1–10. https://doi.org/10.1371/journal.pone.0055731
Couillerot O, Poirier M-A, Prigent-Combaret C et al (2010a) Assessment of SCAR markers to design real-time PCR primers for rhizosphere quantification of Azospirillum brasilense phytostimulatory inoculants of maize. J Appl Microbiol 109:528–538. https://doi.org/10.1111/j.1365-2672.2010.04673.x
Couillerot O, Bouffaudab M-L, Baudoin E et al (2010b) Development of a real-time PCR method to quantify the PGPR strain Azospirillum lipoferum CRT1 on maize seedlings. Soil Biol Bioch 42:2298–2305. https://doi.org/10.1016/j.soilbio.2010.09.003
D’haene B, Vandesompele J, Hellemans J (2010) Accurate and objective copy number profiling using real-time quantitative PCR. Methods 50:4262–4270. https://doi.org/10.1016/j.ymeth.2009.12.007
Da Cunha ET, Pedrolo AM, Paludo F et al (2020) Azospirillum brasilense viable cells enumeration using propidium monoazide-quantitative PCR. Arch Microbiol 202:1653–1662. https://doi.org/10.1007/s00203-020-01877-0
Da Silva CG (2017) Uso da técnica de pcr em tempo real para quantificação de bactérias diazotróficas endofíticas em tecidos de cana-de-açúcar. https://tede.ufrrj.br/jspui/bitstream/jspui/4269/2/2017%20-%20Cleudison%20Gabriel%20Nascimento%20da%20Silva.pdf
da Silva MF, Reis VM (2009) Production, characterization and application of polyclonal antibodies against Azospirillum amazonense strain Am15. Bragantia 68:1–11. https://doi.org/10.1590/S0006-87052009000100001
Díaz-Zorita M, Canigia MVF, Bravo OÁ et al (2015) Field evaluation of extensive crops inoculated with Azospirillum sp. In: Cassán F, Okon Y, Creus C (eds) Handbook for Azospirillum. Springer, Cham. https://doi.org/10.1007/978-3-319-06542-7_24
Dionisi HM, Harms G, Layton AC et al (2003) Power analysis for real-time pcr quantification of genes in activated sludge and analysis of the variability introduced by dna extraction. Appl Envir Microbiol 69:6597–6604. https://doi.org/10.1128/AEM.69.11.6597-6604.2003
Dos Santos Ferreira NS, Sant’Anna FH, Reis VM et al (2020) Genome-based reclassification of Azospirillum brasilense Sp245 as the type strain of Azospirillum baldaniorum sp. nov. Int J Syst Evol Microbiol 70:6203–6212. https://doi.org/10.1099/ijsem.0.004517
Dos Santos CLR, Alves GC, Macedo AVM et al (2017) Contribution of a mixed inoculant containing strains of Burkholderia spp. and Herbaspirillum ssp. to the growth of three sorghum genotypes under increased nitrogen fertilization levels. Appl Soil Ecol 113:96–106. https://doi.org/10.1016/j.apsoil.2017.02.008
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin 19:11–15
Faleiro AC, Pereira TP, Espindula E et al (2013) Real time PCR detection targeting nifA gene of plant growth promoting bacteria Azospirillum brasilense strain FP2 in maize roots. Symbiosis 61:125–113. https://doi.org/10.1007/s13199-013-0262-y
Felten AV, Défago G, Maurhofer M (2010) Quantification of Pseudomonas fluorescens strains F113, CHA0 and Pf153 in the rhizosphere of maize by strain-specific real-time PCR unaffected by the variability of DNA extraction efficiency. J Microbiol Methods 81:108–115. https://doi.org/10.1016/j.mimet.2010.02.003
Ferreira DF (2014) Sisvar: a guide for its bootstrap procedures in multiple comparisons. Ciência e Agrotecnologia 38:109–112. https://doi.org/10.1590/S1413-70542014000200001
Franz P, Betat H, Mörl M (2016) Genotyping bacterial and fungal pathogens using sequence variation in the gene for the CCA-adding enzyme. BMC Microbiol 16:1–11. https://doi.org/10.1186/s12866-016-0670-2
Fukami J, Nogueira MA, Araujo RS et al (2016) Accessing inoculation methods of maize and wheat with Azospirillum brasilense. AMB Express 6:1–13. https://doi.org/10.1186/s13568-015-0171-y
Fukami J, Cerezini P, Hungria M (2018) Azospirillum: benefits that go far beyond biological nitrogen fixation. AMB Express 8:73. https://doi.org/10.1186/s13568-018-0608-1
Galindo FS, Buzetti S, Rodrigues WL et al (2020) Inoculation of Azospirillum brasilense associated with silicon as a liming source to improve nitrogen fertilization in wheat crops. Sci Rep 10:6160. https://doi.org/10.1038/s41598-020-63095-4
Gouda S, Kerry RG, Das G et al (2018) Revitalization of plant growth promoting rhizobacteria for sustainable development in agriculture. Microbiol Res 206:131–140. https://doi.org/10.1016/j.micres.2017.08.016
Hungria M, Campo RJ, Souza EM et al (2010) Inoculation with selected strains of Azospirillum brasilense and A. lipoferum improves yields of maize and wheat in Brazil. Plant Soil 331:413–425. https://doi.org/10.1007/s11104-009-0262-0
Hungria M, Nogueira MA, Araujo RS (2015) Soybean seed co-inoculation with Bradyrhizobium spp. and Azospirillum brasilense: a new biotechnological tool to improve yield and sustainability. American J Plant Sci 6:811–817. https://doi.org/10.4236/ajps.2015.66087
Hungria M, Nogueira MA, Araujo RS (2016) Inoculation of Brachiaria spp. with the plant growth-promoting bacterium Azospirillum brasilense: an environment-friendly component in the reclamation of degraded pastures in the tropics. Agriculture. Ecosystems and Environment 221:125–131. https://doi.org/10.1016/j.agee.2016.01.024
Kang MJ, Kima MH, Hwang DJ et al (2012) Quantitative in planta PCR assay for specific detection of Xanthomonas oryzae pv. Oryzicola using putative membrane protein based primer set. Crop Prot 40:22–27. https://doi.org/10.1016/j.cropro.2012.04.014
Kuang J, Yan X, Genders AJ et al (2018) An overview of technical considerations when using quantitative real-time PCR analysis of gene expression in human exercise research. PLoS One 13:e0196438. https://doi.org/10.1371/journal.pone.0196438
Lareen A, Burton F, Schäfer P (2016) Plant root-microbe communication in shaping root microbiomes. Plant Mol Biol 90:575–587. https://doi.org/10.1007/s11103-015-0417-8
Leite RDC, dos Santos JGD, Silva EL et al (2019) Productivity increase, reduction of nitrogen fertiliser use and drought-stress mitigation by inoculation of Marandu grass (Urochloa brizantha) with Azospirillum brasilense. Crop and Pasture Science 70:61–67. https://doi.org/10.1071/CP18105
Li W, Raoult D, Fournier P-E (2009) Bacterial strain typing in the genomic era. FEMS Microbiol Rev 33:892–916. https://doi.org/10.1111/j.1574-6976.2009.00182.x
Li X, Jensen BB, Højberg O, Noel SJ, Canibe N (2018) Development of a species-specific TaqMan-MGB real-time PCR assay to quantify Olsenella scatoligenes in pigs offered a chicory root-based diet. AMB Express 8:2018. https://doi.org/10.1186/s13568-018-0627-y
Lin SY, Hameed A, Shen FT et al (2014) Description of Niveispirillum fermenti gen. nov., sp. nov., isolated from a fermentor in Taiwan, transfer of Azospirillum irakense (1989) as Niveispirillum irakense comb. nov., and reclassification of Azospirillum amazonense (1983) as Nitrospirillum amazonense gen. nov. Antonie van Leeuwenhoek 105:1149–1162. https://doi.org/10.1007/s10482-014-0176-6
Marks BB, Megías M, Ollero FJ et al (2015) Maize growth promotion by inoculation with Azospirillum brasilense and metabolites of rhizobium tropici enriched on lipo-chitooligosaccharides (LCOs). AMB Express 5:71. https://doi.org/10.1186/s13568-015-0154-z
Martini M, Moruzzi S, Ermacora P et al (2015) Quantitative real-time PCR and high-resolution melting (HRM) analysis for strain-specific monitoring of fluorescent pseudomonads used as biocontrol agents against soil-borne pathogens of food crops. Trends Food Sci Technol 46:277–285. https://doi.org/10.1016/j.tifs.2015.10.017
Martins MR, Jantalia CP, Reis VM et al (2018) Impact of plant growth-promoting bacteria on grain yield, protein content, and urea-15N recovery by maize in a Cerrado Oxisol. Plant Soil 422:239–250. https://doi.org/10.1007/s11104-017-3193-1
Mehnaz S (2015) Azospirillum: a biofertilizer for every crop. In: N., A. (Ed.). Plant microbes Symbiosis: applied facets. New Delhi: Springer, New Delhi 2015:297–314
Monteiro RA, Schmidt MA, Valter A et al (2008) Early colonization pattern of maize (Zea mays L. Poales, Poaceae) roots by Herbaspirillum seropedicae (Burkholderiales, Oxalobacteraceae). Gen Mol Biol 31:932–937. https://doi.org/10.1590/S1415-47572008005000007
Nabti E, Sahnoune M, Adjrad S et al (2007) A halophilic and osmotolerant Azospirillum brasilense strain from algerian soil restores wheat growth under saline conditions. Engineering in Life Sciences 7:354–360. https://doi.org/10.1002/elsc.200720201
Oliveira ALM, Canuto EdL, Reis VM et al (2003) Response of micropropagated sugarcane varieties to inoculation with endophytic diazotrophic bacteria. Braz J Microbiol 34:59–61. https://doi.org/10.1590/S1517-83822003000500020
Pedrosa FO, Yates MG (1984) Regulation of nitrogen fixation (nif) of Azospirillum brasilense by Nifa and Ntr (Gln) type gene-products. FEMS Microbiol Let 23:95–101. https://doi.org/10.1111/j.1574-6968.1984.tb01042.x
Pereg L, De-Bashan L,E, Bashan Y (2015) Assessment of affinity and specificity of Azospirillum for plants. Plant Soil 399:389–414. https://doi.org/10.1007/s11104-015-2778-9
Pereira TP, do Amaral FP, Dall’Asta P et al (2014) Real-time PCR quantification of the plant growth promoting bacteria Herbaspirillum seropedicae strain SmR1 in maize roots. Mol Biotechnol 56:660–670. https://doi.org/10.1007/s12033-014-9742-4
Pereira W, Oliveira RP, Pereira A et al (2021) Nitrogen acquisition and 15N-fertiliser recovery efficiency of sugarcane cultivar RB92579 inoculated with five diazotrophs. Nutr Cycl Agroecosyst 119:37–50. https://doi.org/10.1007/s10705-020-10100-x
Pollard AT, Okubara PA (2019) Real-time PCR quantification of Fusarium avenaceum in soil and seeds. J Microbiol Meth 157:21–30. https://doi.org/10.1016/j.mimet.2018.12.009
Raynaud X, Jaillard B, Leadley PW (2008) Plants may alter competition by modifying nutrient bioavailability in rhizosphere: a modeling approach. Am Nat 171:44–58. https://doi.org/10.1086/523951
Reis VM, Baldani VLD, Baldani JI (2015) Isolation, identification and biochemical characterization of Azospirillum spp. and other nitrogen-fixing Bacteria. In: Cassán F, Okon Y, Creus C (eds) Handbook for Azospirillum. Springer, Cham. https://doi.org/10.1007/978-3-319-06542-7_1
Rihs JD, Brenner DJ, Weaver RE et al (1993) Roseomonas, a new genus associated with bacteremia and other human infections. J Clin Microbiol 31:3275–3283. https://doi.org/10.1128/jcm.31.12.3275-3283.1993
Ruiz-Villalba A, van Pelt-Verkuil E, Gunst QD et al (2017) Amplification of nonspecific products in quantitative polymerase chain reactions (qPCR). Biomol Detect Quantif 14:7–18. https://doi.org/10.1016/j.bdq.2017.10.001
Sá GCR, Carvalho CLM, Moreira A et al (2019) Biomass yield, nitrogen accumulation and nutritive value of mavuno grass inoculated with plant growth-promoting bacteria. Commun Soil Sci Plant Anal 50:151931–151942. https://doi.org/10.1080/00103624.2019.1648498
Schloter M, Hartmann A (1998) Endophytic and surface colonization of wheat roots (Triticum aestivum) by different Azospirillum brasilense strains studied with strain-specific monoclonal antibodies. Symbiosis 25:159–179
Shameer S, Prasad TNVKV (2018) Plant growth promoting rhizobacteria for sustainable agricultural practices with special reference to biotic and abiotic stresses. Plant Growth Regul 84:603–615. https://doi.org/10.1007/s10725-017-0365-1
Spaepen S, Dobbelaere S, Croonenborghs A, Vanderleyden J (2008) Effects of Azospirillum brasilense indole-3-acetic acid production on inoculated wheat plants. Plant Soil 312:15–23. https://doi.org/10.1007/s11104-008-9560-1
Spaepen S, Vanderleyden J, Okon Y (2009) Plant growth-promoting actions of Rhizobacteria. In: LOON, L. C. VAN (Ed.). Advances in botanical research. 1. Ed. Elsevier Ltd 51:283–320
Stets MI, Alqueres SMC, de Souza EM et al (2015) Quantification of Azospirillum brasilense FP2 bacteria in wheat roots by strain-specific quantitative PCR. Appl Environ Microbiol 81:196700–196709. https://doi.org/10.1128/AEM.01351-15
Tarrand JJ, Krieg NR, Dobereiner J (1978) A taxonomic study of the Spirillum lipoferum group, with descriptions of a new genus, Azospirillum gen. nov. and two species, Azospirillum lipoferum (Beijerinck) comb. nov. and Azospirillum brasilense sp. nov.. Can J Microbiol 24:967–980. https://doi.org/10.1139/m78-160
Thermofisher (2016) Real-time PCR handbook. https://www.ffclrp.usp.br/divulgacao/emu/real_time/manuais/Apostila%20qPCR-Handbook.pdf. Accessed 2 Dec 2019
Timmusk S, Behers L, Muthoni J et al (2017) Perspectives and challenges of microbial application for crop improvement. Front Plant Sci 8:1–10. https://doi.org/10.3389/fpls.2017.00049
Untergasser A, Cutcutache I, Koressaar T et al (2012) Primer3- new capabilities and interfaces. Nucleic Acids Res 40:e115. https://doi.org/10.1093/nar/gks596
Urrea-Valencia S, Etto RM, Takahashi WY et al (2021) Detection of Azospirillum brasilense by qPCR throughout a maize field trial. Appl Soil Ecol 160:103849. https://doi.org/10.1016/j.apsoil.2020.103849
Zhang T, Fang HHP (2006) Applications of real-time polymerase chain reaction for quantification of microorganisms in environmental samples. Appl Microbiol Biotech 70:281–289. https://doi.org/10.1007/s00253-006-0333-6
Acknowledgements
We would like to express our gratitude to Johanna Döbereiner Biological Resource Center (https://www.embrapa.br/agrobiologia/crb-jd) at Embrapa Agrobiology for providing bacterial strains for this study.
Funding
This research was supported by a research grant from Embrapa (n° 12.13.08.004.00.02.006). Isis C. Soares and Rafael Sanches Pacheco were supported by fellowship from the Coordination of Personnel Improvement of Higher Education (CAPES—finance Code 001), Ministry of Education, Brazil, while Jean Luiz Simões-Araujo is recipient Researcher fellowship from the National Council of Scientific and Technological Development (CNPq, process n° 310642/2019–1). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Katharina Pawlowski.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 1498 kb)
Rights and permissions
About this article
Cite this article
Soares, I.C., Pacheco, R.S., da Silva, C.G.N. et al. Real-time PCR method to quantify Sp245 strain of Azospirillum baldaniorum on Brachiaria grasses under field conditions. Plant Soil 468, 525–538 (2021). https://doi.org/10.1007/s11104-021-05137-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-021-05137-y