Abstract
Bacillus cereus is one of the most common foodborne pathogens found in various kinds of staple foods such as rice and wheat. A rapid and accurate detection method for this pathogen is highly desirable for the sustainable production of relevant food products. While several classical and molecular-based detection methods are available for the identification of B. cereus, they suffered one or more limitations such as the requirement for a tedious and time-consuming process, less than ideal specificity, and the lack of portability. Herein, we developed the first paper-based sensing device that exhibits high species specificity with sufficiently low limit of detection for the visual detection of specific DNA sequences of B. cereus. The success is attributed to the strategic planning of fabrication in various dimensions including thorough bioinformatics search for highly specific genes, the use of the pyrrolidinyl peptide nucleic acid (PNA) probe whose selectivity advantage is well documented, and an effective PNA immobilization and DNA-binding visualization method with an internal cross-checking system for validating the results. Testing in rice matrices indicates that the sensor is capable of detecting and distinguishing B. cereus from other bacterial species. Hence, this paper-based sensor has potential to be adopted as a practical means to detect B. cereus in food industries.
Graphical abstract






Similar content being viewed by others
References
Granum PE, Lund T. Bacillus cereus and its food poisoning toxins. FEMS Microbiol Lett. 1997;157(2):223–8.
Frenzel E, Kranzler M, Stark TD, Hofmann T, Ehling-Schulz M. The endospore-forming pathogen Bacillus cereus exploits a small colony variant-based diversification strategy in response to aminoglycoside exposure. mBio. 2015;6(6):e01172–15.
Setlow P. Mechanisms which contribute to the long-term survival of spores of Bacillus species. J Appl Bacteriol. 1994;76(S23):49S–60S.
Ramarao N, Tran S-L, Marin M, Vidic J. Advanced methods for detection of Bacillus cereus and its pathogenic factors. Sensors. 2020;20(9):2667.
Schulten SM, In’t Veld PH, Nagelkerke NJD, Scotter S, de Buyser ML, Rollier P, Lahellec C. Evaluation of the ISO 7932 standard for the enumeration of Bacillus cereus in foods. Int J. Food Microbiol. 2000;57(1):53–61.
Chon J-W, Kim Y-J, Kim D-H, Song K-Y, Kim H, Seo K-H. Supplementation of modified mannitol-yolk-polymyxin B agar with cefuroxime for quantitative detection of Bacillus cereus in food. J Food Sci. 2019;84(1):133–7.
Zhang Z, Feng L, Xu H, Liu C, Shah NP, Wei H. Detection of viable enterotoxin-producing Bacillus cereus and analysis of toxigenicity from ready-to-eat foods and infant formula milk powder by multiplex PCR. J Dairy Sci. 2016;99(2):1047–55.
Sadek ZI, Abdel-Rahman MA, Azab MS, Darwesh OM, Hassan MS. Microbiological evaluation of infant foods quality and molecular detection of Bacillus cereus toxins relating genes. Toxicol. Rep. 2018;5:871–7.
Sánchez-Chica J, Correa MM, Aceves-Diez AE, Castañeda-Sandoval LM. A novel method for direct detection of Bacillus cereus toxin genes in powdered dairy products. Int. Diary J. 2020;103:104625.
Yabutani M, Agata N, Ohta M. A new rapid and sensitive detection method for cereulide-producing Bacillus cereus using a cycleave real-time PCR. Lett Appl Microbiol. 2009;48(6):698–704.
Fischer C, Hünniger T, Jarck J-H, Frohnmeyer E, Kallinich C, Haase I, Hahn U, Fischer M. Food sensing: aptamer-based trapping of Bacillus cereus spores with specific detection via real time PCR in milk. J Agric Food Chem. 2015;63(36):8050–7.
Cattani F, Barth VC Jr, Nasário JSR, Ferreira CAS, Oliveira SD. Detection and quantification of viable Bacillus cereus group species in milk by propidium monoazide quantitative real-time PCR. J Dairy Sci. 2016;99(4):2617–24.
Kim K, Seo J, Wheeler K, Park C, Kim D, Park S, Kim W, Chung SI, Leighton T. Rapid genotypic detection of Bacillus anthracis and the Bacillus cereus group by multiplex real-time PCR melting curve analysis. FEMS Immunol Med Microbiol. 2005;43(2):301–10.
Oliwa-Stasiak K, Molnar CI, Arshak K, Bartoszcze M, Adley CC. Development of a PCR assay for identification of the Bacillus cereus group species. J Appl Microbiol. 2010;108(1):266–73.
Helgason E, Økstad OA, Caugant DA, Johansen HA, Fouet A, Mock M, Hegna I, Kolstø A-B. Bacillus anthracis, Bacillus cereus, and Bacillus thuringiensis—one species on the basis of genetic evidence. Appl Environ Microbiol. 2000;66(6):2627–30.
Muangkaew P, Vilaivan T. Modulation of DNA and RNA by PNA. Bioorg Med Chem Lett. 2020;30(9):127064.
Patel R, Sarma S, Shukla A, Parmar P, Goswami D, Saraf M. Walking through the wonder years of artificial DNA: peptide nucleic acid. Mol Biol Rep. 2020;47(10):8113–31.
Vilaivan T. Pyrrolidinyl PNA with α/β-dipeptide backbone: from development to applications. Acc Chem Res. 2015;48(6):1645–56.
Laopa PS, Vilaivan T, Hoven VP. Positively charged polymer brush-functionalized filter paper for DNA sequence determination following dot blot hybridization employing a pyrrolidinyl peptide nucleic acid probe. Analyst. 2013;138(1):269–77.
Maneelun N, Vilaivan T. Dual pyrene-labeled pyrrolidinyl peptide nucleic acid as an excimer-to-monomer switching probe for DNA sequence detection. Tetrahedron. 2013;69(51):10805–10.
Boonlua C, Ditmangklo B, Reenabthue N, Suparpprom C, Poomsuk N, Siriwong K, Vilaivan T. Pyrene-labeled pyrrolidinyl peptide nucleic acid as a hybridization-responsive DNA probe: comparison between internal and terminal labeling. RSC Adv. 2014;4(17):8817–27.
Jirakittiwut N, Panyain N, Nuanyai T, Vilaivan T, Praneenararat T. Pyrrolidinyl peptide nucleic acids immobilised on cellulose paper as a DNA sensor. RSC Adv. 2015;5(31):24110–4.
Yotapan N, Nim-anussornkul D, Vilaivan T. Pyrrolidinyl peptide nucleic acid terminally labeled with fluorophore and end-stacking quencher as a probe for highly specific DNA sequence discrimination. Tetrahedron. 2016;72(49):7992–9.
Kangkamano T, Numnuam A, Limbut W, Kanatharana P, Vilaivan T, Thavarungkul P. Pyrrolidinyl PNA polypyrrole/silver nanofoam electrode as a novel label-free electrochemical miRNA-21 biosensor. Biosens. Bioelectron. 2018;102:217–25.
Cecchini F, Iacumin L, Fontanot M, Comuzzo P, Comi G, Manzano M. Dot blot and PCR for Brettanomyces bruxellensis detection in red wine. Food Control. 2013;34(1):40–6.
Trinh TND, Lee NY. A foldable isothermal amplification microdevice for fuchsin-based colorimetric detection of multiple foodborne pathogens. Lab Chip. 2019;19(8):1397–405.
Nguyen QH, Kim MI. Nanomaterial-mediated paper-based biosensors for colorimetric pathogen detection. TrAC, Trends Anal. Chem. 2020;132:116038.
Na M, Zhang S, Liu J, Ma S, Han Y, Wang Y, He Y, Chen H, Chen X. Determination of pathogenic bacteria―Bacillus anthrax spores in environmental samples by ratiometric fluorescence and test paper based on dual-emission fluorescent silicon nanoparticles. J. Hazard. Mater. 2020;386:121956.
Jokerst JC, Adkins JA, Bisha B, Mentele MM, Goodridge LD, Henry CS. Development of a paper-based analytical device for colorimetric detection of select foodborne pathogens. Anal Chem. 2012;84(6):2900–7.
Saengsawang N, Ruang-areerate T, Kesakomol P, Thita T, Mungthin M, Dungchai W. Development of a fluorescent distance-based paper device using loop-mediated isothermal amplification to detect Escherichia coli in urine. Analyst. 2020;145(24):8077–86.
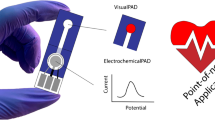
Sharafeldin M, Davis JJ. Point of care sensors for infectious pathogens. Anal Chem. 2021;93(1):184–97.
Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016;33(7):1870–4.
Vilaivan T, Srisuwannaket C. Hybridization of pyrrolidinyl peptide nucleic acids and DNA: selectivity, base-pairing specificity, and direction of binding. Org Lett. 2006;8(9):1897–900.
Jirakittiwut N, Munkongdee T, Wongravee K, Sripichai O, Fucharoen S, Praneenararat T, Vilaivan T. Visual genotyping of thalassemia by using pyrrolidinyl peptide nucleic acid probes immobilized on carboxymethylcellulose-modified paper and enzyme-induced pigmentation. Microchim Acta. 2020;187(4):238.
Winichagoon P, Saechan V, Sripanich R, Nopparatana C, Kanokpongsakdi S, Maggio A, Fucharoen S. Prenatal diagnosis of beta-thalassaemia by reverse dot-blot hybridization. Prenat Diagn. 1999;19(5):428–35.
Orelma H, Teerinen T, Johansson L-S, Holappa S, Laine J. CMC-modified cellulose biointerface for antibody conjugation. Biomacromolecules. 2012;13(4):1051–8.
Public Health England. Ready-to-eat foods: microbiological safety assessment guidelines. (2009). https://www.gov.uk/government/publications/ready-to-eat-foods-microbiological-safety-assessment-guidelines. Accessed 16 Aug 2021.
Food Standards Australia New Zealand. Compendium of microbiological criteria for food (2018). https://www.foodstandards.gov.au/publications/pages/compendium-of-microbiological-criteria-for-food.aspx. Accessed 16 Aug 2021.
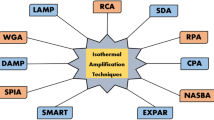
Zhao Y, Chen F, Li Q, Wang L, Fan C. Isothermal amplification of nucleic acids. Chem Rev. 2015;115(22):12491–545.
Funding
This work was supported by Thailand Science Research and Innovation (TSRI) (formerly the Thailand Research Fund, TRF) grant no. RGU6180001 (to TV) and MRG6280238 (to TP). NJ thanks the Science Achievement Scholarship of Thailand for his Ph.D. scholarship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(PDF 773 kb)
Rights and permissions
About this article
Cite this article
Jirakittiwut, N., Patipong, T., Cheiwchanchamnangij, T. et al. Paper-based sensor from pyrrolidinyl peptide nucleic acid for the efficient detection of Bacillus cereus. Anal Bioanal Chem 413, 6661–6669 (2021). https://doi.org/10.1007/s00216-021-03633-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-021-03633-9