Abstract
Background
Considering the increase in the number of HCC patients, it is critical to predict the survival of patients. Although ferroptosis is closely related to HCC progression, predicting the survival of HCC patients through ferroptosis-related genes is challenging.
Methods
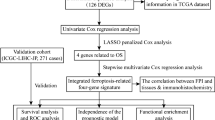
RNA-seq and clinical data of HCC in the TCGA database were analyzed to establish a prognostic model, and ICGC and GSE14520 data were used for validation. Risk score was constructed with 5 genes identified by univariate and LASSO Cox regression analysis. Risk score, TNM stage and cirrhosis were incorporated to construct a nomogram through univariate and multivariate Cox regression analysis.
Results
Five genes identified from 70 ferroptosis-related DEGs were used to construct a gene signature that predicts survival of HCC patients in the TCGA cohort. PCA and heatmap showed clear differences between patients in different score groups. Next, risk score, TNM stage and cirrhosis were combined in a nomogram for overall survival prediction. Survival analysis indicated that the overall survival of the low-risk group was significantly higher than that of the high-risk group. The data from the GSE14520 cohort confirmed satisfactory nomogram performance. Furthermore, KEGG and GO functional enrichment analyses indicated that the difference in overall survival between risk groups was closely related to immune-related pathways. Further analyses implied that an immune-suppressive tumor microenvironment might contribute to the difference in the prognosis between risk groups.
Conclusion
The nomogram based on ferroptosis-related genes showed good performance for predicting the prognosis of HCC patients. The model may provide a reference for the evaluation of HCC patients by targeting ferroptosis.






Similar content being viewed by others
References
Forner A, Reig M, Bruix J. Hepatocellular carcinoma. Lancet 2018;391(10127):1301–1314
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2021;71(3):209–249
Heimbach JK, Kulik LM, Finn RS, Sirlin CB, Abecassis MM, Roberts LR, et al. AASLD guidelines for the treatment of hepatocellular carcinoma. Hepatology 2018;67(1):358–380
Dixon SJ, Lemberg KM, Lamprecht MR, Skouta R, Zaitsev EM, Gleason CE, et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 2012;149(5):1060–1072
Stockwell BR, Friedmann Angeli JP, Bayir H, Bush AI, Conrad M, Dixon SJ, et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell 2017;171(2):273–285
Yang WS, SriRamaratnam R, Welsch ME, Shimada K, Skouta R, Viswanathan VS, et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 2014;156(1–2):317–331
Eling N, Reuter L, Hazin J, Hamacher-Brady A, Brady NR. Identification of artesunate as a specific activator of ferroptosis in pancreatic cancer cells. Oncoscience 2015;2(5):517–532
Louandre C, Marcq I, Bouhlal H, Lachaier E, Godin C, Saidak Z, et al. The retinoblastoma (Rb) protein regulates ferroptosis induced by sorafenib in human hepatocellular carcinoma cells. Cancer Lett 2015;356(2 Pt B):971–977
Sun X, Ou Z, Chen R, Niu X, Chen D, Kang R, et al. Activation of the p62-Keap1-NRF2 pathway protects against ferroptosis in hepatocellular carcinoma cells. Hepatology 2016;63(1):173–184
Yuan H, Li X, Zhang X, Kang R, Tang D. CISD1 inhibits ferroptosis by protection against mitochondrial lipid peroxidation. Biochem Biophys Res Commun 2016;478(2):838–844
Sun X, Niu X, Chen R, He W, Chen D, Kang R, et al. Metallothionein-1G facilitates sorafenib resistance through inhibition of ferroptosis. Hepatology 2016;64(2):488–500
Wang Q, Bin C, Xue Q, Gao Q, Huang A, Wang K, et al. GSTZ1 sensitizes hepatocellular carcinoma cells to sorafenib-induced ferroptosis via inhibition of NRF2/GPX4 axis. Cell Death Dis 2021;12(5):426
Sun J, Zhou C, Zhao Y, Zhang X, Chen W, Zhou Q, et al. Quiescin sulfhydryl oxidase 1 promotes sorafenib-induced ferroptosis in hepatocellular carcinoma by driving EGFR endosomal trafficking and inhibiting NRF2 activation. Redox Biol 2021;41:101942
Gui J, Li H. Penalized Cox regression analysis in the high-dimensional and low-sample size settings, with applications to microarray gene expression data. Bioinformatics 2005;21(13):3001–3008
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 2012;16(5):284–287
Becht E, Giraldo NA, Lacroix L, Buttard B, Elarouci N, Petitprez F, et al. Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol 2016;17(1):218
Newman AM, Liu CL, Green MR, Gentles AJ, Feng W, Xu Y, et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 2015;12(5):453–457
Sartorius K, Sartorius B, Aldous C, Govender PS, Madiba TE. Global and country underestimation of hepatocellular carcinoma (HCC) in 2012 and its implications. Cancer Epidemiol 2015;39(3):284–290
Nie J, Lin B, Zhou M, Wu L, Zheng T. Role of ferroptosis in hepatocellular carcinoma. J Cancer Res Clin Oncol 2018;144(12):2329–2337
Yu H, Guo P, Xie X, Wang Y, Chen G. Ferroptosis, a new form of cell death, and its relationships with tumourous diseases. J Cell Mol Med 2017;21(4):648–657
Jiraskova A, Novotny J, Novotny L, Vodicka P, Pardini B, Naccarati A, et al. Association of serum bilirubin and promoter variations in HMOX1 and UGT1A1 genes with sporadic colorectal cancer. Int J Cancer 2012;131(7):1549–1555
Barker HE, Cox TR, Erler JT. The rationale for targeting the LOX family in cancer. Nat Rev Cancer 2012;12(8):540–552
Erler JT, Bennewith KL, Nicolau M, Dornhofer N, Kong C, Le QT, et al. Lysyl oxidase is essential for hypoxia-induced metastasis. Nature 2006;440(7088):1222–1226
Bouez C, Reynaud C, Noblesse E, Thepot A, Gleyzal C, Kanitakis J, et al. The lysyl oxidase LOX is absent in basal and squamous cell carcinomas and its knockdown induces an invading phenotype in a skin equivalent model. Clin Cancer Res 2006;12(5):1463–1469
Bai T, Yokobori T, Altan B, Ide M, Mochiki E, Yanai M, et al. High STMN1 level is associated with chemo-resistance and poor prognosis in gastric cancer patients. Br J Cancer 2017;116(9):1177–1185
Leung TW, Tang AM, Zee B, Lau WY, Lai PB, Leung KL, et al. Construction of the Chinese University Prognostic Index for hepatocellular carcinoma and comparison with the TNM staging system, the Okuda staging system, and the Cancer of the Liver Italian Program staging system: a study based on 926 patients. Cancer 2002;94(6):1760–1769
Cillo U, Vitale A, Grigoletto F, Farinati F, Brolese A, Zanus G, et al. Prospective validation of the Barcelona Clinic Liver Cancer staging system. J Hepatol 2006;44(4):723–731
Gomez EV, Rodriguez YS, Bertot LC, Gonzalez AT, Perez YM, Soler EA, et al. The natural history of compensated HCV-related cirrhosis: a prospective long-term study. J Hepatol 2013;58(3):434–444
Polaris OC. Global prevalence, treatment, and prevention of hepatitis B virus infection in 2016: a modelling study. Lancet Gastroenterol Hepatol 2018;3(6):383–403
Wan S, Nie Y, Zhu X. Development of a prognostic scoring model for predicting the survival of elderly patients with hepatocellular carcinoma. PeerJ 2020;8:e8497
Zhang W, Tan Y, Shen S, Jiang L, Yan L, Yang J, et al. Prognostic nomogram for hepatocellular carcinoma in adolescent and young adult patients after hepatectomy. Oncotarget 2017;8(63):106393–106404
Wan G, Gao F, Chen J, Li Y, Geng M, Sun L, et al. Nomogram prediction of individual prognosis of patients with hepatocellular carcinoma. BMC Cancer 2017;17(1):91
Liang JY, Wang DS, Lin HC, Chen XX, Yang H, Zheng Y, et al. A novel ferroptosis-related gene signature for overall survival prediction in patients with hepatocellular carcinoma. Int J Biol Sci 2020;16(13):2430–2441
Sangro B, Gomez-Martin C, de la Mata M, Inarrairaegui M, Garralda E, Barrera P, et al. A clinical trial of CTLA-4 blockade with tremelimumab in patients with hepatocellular carcinoma and chronic hepatitis C. J Hepatol 2013;59(1):81–88
Mazzaferro V, Romito R, Schiavo M, Mariani L, Camerini T, Bhoori S, et al. Prevention of hepatocellular carcinoma recurrence with alpha-interferon after liver resection in HCV cirrhosis. Hepatology 2006;44(6):1543–1554
Kawada N, Imanaka K, Kawaguchi T, Tamai C, Ishihara R, Matsunaga T, et al. Hepatocellular carcinoma arising from non-cirrhotic nonalcoholic steatohepatitis. J Gastroenterol 2009;44(12):1190–1194
Lee BP, Vittinghoff E, Dodge JL, Cullaro G, Terrault NA. National trends and long-term outcomes of liver transplant for alcohol-associated liver disease in the United States. JAMA Intern Med 2019;179(3):340–348
Liu Z, Jiang Y, Yuan H, Fang Q, Cai N, Suo C, et al. The trends in incidence of primary liver cancer caused by specific etiologies: results from the Global Burden of Disease Study 2016 and implications for liver cancer prevention. J Hepatol 2019;70(4):674–683
Li J, Zou B, Yeo YH, Feng Y, Xie X, Lee DH, et al. Prevalence, incidence, and outcome of non-alcoholic fatty liver disease in Asia, 1999–2019: a systematic review and meta-analysis. Lancet Gastroenterol Hepatol 2019;4(5):389–398
Yamashita T, Kitao A, Matsui O, Hayashi T, Nio K, Kondo M, et al. Gd-EOB-DTPA-enhanced magnetic resonance imaging and alpha-fetoprotein predict prognosis of early-stage hepatocellular carcinoma. Hepatology 2014;60(5):1674–1685
Hsu CY, Shen YC, Yu CW, Hsu C, Hu FC, Hsu CH, et al. Dynamic contrast-enhanced magnetic resonance imaging biomarkers predict survival and response in hepatocellular carcinoma patients treated with sorafenib and metronomic tegafur/uracil. J Hepatol 2011;55(4):858–865
Waidmann O, Koberle V, Bettinger D, Trojan J, Zeuzem S, Schultheiss M, et al. Diagnostic and prognostic significance of cell death and macrophage activation markers in patients with hepatocellular carcinoma. J Hepatol 2013;59(4):769–779
Acknowledgements
We would like to thank the National Natural Science Foundation of China for financial support.
Funding
This study was supported by the National Natural Science Foundation of China (82070574), the Natural Science Foundation Team Project of Guangdong Province (2018B030312009), and the Fundamental Research Funds for the Central Universities (19ykpy29).
Author information
Authors and Affiliations
Contributions
SW and YL contributed equally to this study. SW designed and wrote the manuscript. YL screened the literature and collected data. ML analyzed the data. BW critically revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors Sizhe Wan, Yiming Lei, Mingkai Li, Bin Wu declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wan, S., Lei, Y., Li, M. et al. A prognostic model for hepatocellular carcinoma patients based on signature ferroptosis-related genes. Hepatol Int 16, 112–124 (2022). https://doi.org/10.1007/s12072-021-10248-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12072-021-10248-w