Abstract
Objectives
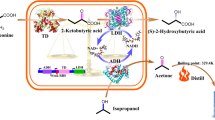
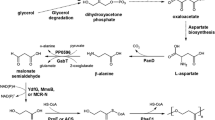
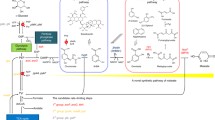
3,4-Dihydroxybutyric acid (3,4-DHBA) is a multifunctional C4 platform compound widely used for the synthesis of various materials, including pharmaceuticals. Although, a biosynthetic pathway for 3,4-DHBA production has been developed, its low yield still precludes large-scale use. Here, a heterologous four-step biosynthetic pathway was established in recombinant Escherichia coli (E. coli) using a combinatorial strategy.
Results
Several aldehyde dehydrogenases (ALDHs) were screened, using in vitro enzyme assays, to identify suitable catalysts for the dehydrogenation of 3,4-dihydroxybutanal (3,4-DHB) to 3,4-DHBA. A pathway containing glucose dehydrogenase (BsGDH) from Bacillus subtilis, d-xylonate dehydratase (YagF) from E. coli, benzoylformate decarboxylase (PpMdlC) from Pseudomonas putida and ALDH was introduced into E. coli, generating 3.04 g/L 3,4-DHBA from d-xylose (0.190 g 3,4-DHBA/g d-xylose). Disruption of competing pathways by deleting xylA, ghrA, ghrB and adhP contributed to an 87% increase in 3,4-DHBA accumulation. Expression of a fusion construct containing PpMdlC and YagF enhanced the 3,4-DHBA titer, producing the highest titer and yield reported thus far (7.71 g/L; 0.482 g 3,4-DHBA/g d-xylose).
Conclusions
These results showed that deleting genes from competing pathways and constructing fusion proteins significantly improved the titer and yield of 3,4-DHBA in engineered E. coli.




Similar content being viewed by others
References
Albertsen L, Chen Y, Bach LS, Rattleff S, Maury J, Brix S, Nielsen J, Mortensen UH (2011) Diversion of flux toward sesquiterpene production in Saccharomyces cerevisiae by fusion of host and heterologous enzymes. Appl Environ Microbiol 77:1033–1040
Brower PL, Butler DE, Deering CF, Le TV, Millar A, Nanninga TN, Roth D (1992) The synthesis of (4R-cis)-1,1-dimethylethyl 6-cyanomethyl-2,2-dimethyl-1,3-dioxane-4-acetate, a key intermediate for the preparation of CI-981, a highly potent, tissue selective inhibitor of HMG-CoA reductase. Tetrahedron Lett 33:2279–2282
Chen K, Li K, Deng J, Zhang B, Lin J, Wei D (2016) Carbonyl reductase identification and development of whole-cell biotransformation for highly efficient synthesis of (R)-[3,5-bis(trifluoromethyl)phenyl] ethanol. Microb Cell Fact 15:191
Choi KY, Wernick DG, Tat CA, Liao JC (2014) Consolidated conversion of protein waste into biofuels and ammonia using Bacillus subtilis. Metab Eng 23:53–61
Choi S, Song CW, Shin JH, Lee SY (2015) Biorefineries for the production of top building block chemicals and their derivatives. Metab Eng 28:223–239
Corey EJ, Niwa H, Knolle J (1978) Total synthesis of (S)-12-hydroxy-5,8,14-cis-10-trans-eicosatetraenoic acid (Samuelesson’s HETE). J Am Chem Soc 100(6):1942–1944
Dhamankar H, Tarasova Y, Martin CH, Prather KLJ (2014) Engineering E. coli for the biosynthesis of 3-hydroxy-γ-butyrolactone (3HBL) and 3,4-dihydroxybutyric acid (3,4-DHBA) as value-added chemicals from glucose as a sole carbon source. Metab Eng 25:72–81
Hollingsworth RI (1994) Process for the preparation of 3,4-dihydroxybutanoic acid and salts thereof. European Patent Office Publ. of Application with search report EP19910114350
Inoue K, Matsumoto M, Takahashi S (1991) Method of preparing optically active 3,4-dihydroxy butyric acid derivatives. European Patent Office Publ. of Application with search report EP0339618 B1
Kawaguchi H, Hasunuma T, Ogino C, Kondo A (2016) Bioprocessing of bio-based chemicals produced from lignocellulosic feedstocks. Curr Opin Biotechnol 42:30–39
Kim EE, Baker CT, Dwyer MD, Murcko MA, Rao BG, Tung RD, Navia MA (1995) Crystal structure of HIV-1 protease in complex with VX-478, a potent and orally bioavailable inhibitor of the enzyme. J Am Chem Soc 117(3):1181–1182
Li J, Zhang R, Xu Y, Xiao R, Li K, Liu H, Jiang J, Zhou X, Li L, Zhou L (2017) Ala258Phe substitution in Bacillus sp. YX-1 glucose dehydrogenase improves its substrate preference for xylose. Process Biochem 56:124–131
Liu XW, Wang HH, Chen JY, Li XT, Chen GQ (2009) Biosynthesis of poly(3-hydroxybutyrate-co-3-hydroxyvalerate) by recombinant Escherichia coli harboring propionyl-CoA synthase gene (prpE) or propionate permease gene (prpP). Biochem Eng J 43:72–77
Lu P, Feng MG, Li WF, Hu CX (2006) Construction and characterization of a bifunctional fusion enzyme of Bacillus-sourced beta-glucanase and xylanase expressed in Escherichia coli. FEMS Microbiol Lett 261:224–230
Ryosuke F, Shuhei N, Tsutomu T, Akihiko K (2018) Muconic acid production using gene-level fusion proteins in Escherichia coli. ACS Synth Biol 7(11):2698–2705
Sang HP, Lee SH, Sang YL (2010) Preparation of optically active β-amino acids from microbial polyester polyhydroxyalkanoates. J Chem Res 2001:498–499
Schweiger G, Buckel W (1984) On the dehydration of (R)-lactate in the fermentation of alanine to propionate by Clostridium propionicum. FEBS Lett 171:79–84
Shieh HM, Prestwich GD (1982) Chiral, biomimetic total synthesis of(-)-aplysistatin. Tetrahedron Lett 23(45):4643–4646
Sivaramakrishnan S, Spudich JA (2011) Systematic control of protein interaction using a modular ER/K α-helix linker. Proc Natl Acad Sci USA 108(51):20467–20472
Sun L, Yang F, Sun HB, Zhu TC, Li XH, Li Y, Xu ZH, Zhang YP (2016) Synthetic pathway optimization for improved 1,2,4-butanetriol production. J Ind Microbiol Biotechnol 43:67–78
Taguchi S, Yamada M, Matsumoto K, Tajima K, Satoh Y, Munekata M, Ohno K, Kohda K, Shimamura T, Kambe H (2008) A microbial factory for lactate-based polyesters using a lactate-polymerizing enzyme. Proc Natl Acad Sci 105:17323–17327
Tetrahedron M (1990) Enantiomerically pure β, γ-epoxyesters from β-hydroxylactones: synthesis of β-hydroxyesters and (-)-GABOB. Tetrahedron 46:4277–4282
Valdehuesa KNG, Liu H, Ramos K, Si JP, Nisola GM, Lee WK, Chung WJ (2014) Direct bioconversion of d -xylose to 1,2,4-butanetriol in an engineered Escherichia coli. Process Biochem 49:25–32
Wang J, Shen X, Jain R, Wang J, Yuan Q, Yan Y (2017) Establishing a novel biosynthetic pathway for the production of 3,4-dihydroxybutyric acid from xylose in Escherichia coli. Metab Eng 41:39–45
Zhong WQ, Zhang Y, Wu WJ, Liu DH, Chen Z (2019) Metabolic engineering of a homoserine-derived non-natural pathway for the De Novo production of 1,3-propanediol from glucose. ACS Synth Biol 8:587–595
Acknowledgements
This work was supported by the Natural Science Foundation of Shanghai (No. 19ZR1412700), the Fundamental Research Funds for the Central Universities (No. 22221818014), and partially supported by the Open Funding Project of the State Key Laboratory of Bioreactor Engineering.
Supplementary Information
Supplementary Table 1—List of plasmids this study.
Supplementary Table 2—List of primer sequences this study.
Supplementary Figure 1—Schematic illustration of this fusion construct.
Supplementary Figure 2—HPLC-MS in negative ion mode for 3,4-DHBA verification by strain E0-E01-A01. 3,4-DHBA (C4H8O4) was corresponded to the retention time of 1.0 min and the peak of 119.03 Da.
Supplementary Figure 3—SDS-PAGE analysis of target proteins of the engineered strains. Lane 1: whole cell of E0-E01-A02; Lane 2: whole cell of E1-E01-A02; Lane 3: whole cell of E2-E01-A02; Lane 4: whole cell of E3-E01-A02; Lane 5: whole cell of E4-E01-A02; Lane 6: whole cell of E5-E01-A02. M: protein molecular weight marker.
Supplementary Figure 4—Cell growth of engineered strains.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, Y., Mao, X., Zhang, B. et al. Modification of an engineered Escherichia coli by a combinatorial strategy to improve 3,4-dihydroxybutyric acid production. Biotechnol Lett 43, 2035–2043 (2021). https://doi.org/10.1007/s10529-021-03169-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-021-03169-z