Abstract
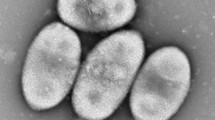
A novel moderately thermophilic and halophilic bacterium, designated strain M0105T, was isolated from mangrove sediment collected in the Beibu Gulf, south China. The isolate is Gram-negative, non-motile and rod-shaped bacterium with smooth colonies of pale-yellow appearance. Growth occurs at 15–46 °C (optimum 37–40 °C) and pH range of 6.0–10.0 (optimum pH 8.0–9.0). It required 1–7% NaCl (optimum 3–5%) for growth. Strain M0105T was affiliated to the family ‘Rhodobacteraceae’, sharing the highest 16S rRNA gene sequence similarity with Limibaculum halophilum CAU 1123T (96.8%). The major menaquinone Q-10 and the dominant unsaturated fatty acid (C18:1ω7) in this family were also detected in the strain M0105T. The genome sequence possesses a circular 4.1 Mb chromosome with a G + C content of 67.9%. Strain M0105T encoded many genes for cellular stress resistance and nutrient utilization, which could improve its adaptive capacity to the mangrove environment. Values of conserved proteins (POCP), average nucleotide identity, average amino acid identity (AAI) and DNA–DNA hybridization (dDDH) between the isolate and closely related species were below the proposed threshold for species discrimination. Information from phenotypic, chemotaxonomic and phylogenetic analyses proposed that strain M0105T should be assigned to a novel genus within the family ‘Rhodobacteraceae’. Thus, we suggested that the strain M0105T represents a novel species in a new genus, for which the name Thermohalobaculum xanthum gen. nov., sp. nov. is proposed. The type strain of the type species is M0105T (= BGMRC 2019T = KCTC 52118T = MCCC 1K03767T = NBRC 112057T).


Similar content being viewed by others
References
Aziz A, Agamuthu P, Alaribe FO, Fauziah SH (2018) Biodegradation of benzo[a]pyrene by bacterial consortium isolated from mangrove sediment. Environ Technol 39:527–535
Barrow GI, Feltham RKA (1993) Cowan and Steel’s manual for the identification of medical bacteria, 3rd edn. Cambridge University Press, Cambridge
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY et al (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res 47:81–87
Giri C, Long J, Abbas S, Murali RM, Qamer FM, Pengra B et al (2015) Distribution and dynamics of mangrove forests of South Asia. J Environ Manage 148:101–111
González JM, Simó R, Massana R, Covert JS, Casamayor EO, Pedrós-Alió C et al (2000) Bacterial community structure associated with a dimethylsulfoniopropionate-producing North Atlantic algal bloom. Appl Environ Microbiol 66:4237–4246
Goris J, Konstantinidis KT, Klappenbach JA, Coenye T, Vandamme P, Tiedje JM (2007) DNA-DNA hybridization values and their relationship to whole-genome sequence similarities. Int J Syst Evol Microbiol 57:81–91
Kathiresan K, Selvam MM (2006) Evaluation of beneficial bacteria from mangrove soil. Bot Mar 1:86–88
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lam MQ, Oates NC, Thevarajoo S, Tokiman L, Goh KM, McQueen-Mason SJ et al (2020) Genomic analysis of a lignocellulose degrading strain from the underexplored genus Meridianimaribacter. Genomics 112:952–960
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–174
Lin X, Hetharua B, Lin L, Xu H, Zheng T, He Z et al (2019) Mangrove sediment microbiome: adaptive microbial assemblages and their routed biogeochemical processes in Yunxiao mangrove national nature reserve, China. Microb Ecol 78:57–69
Liu Y, Lai Q, Wang W, Shao Z (2017) Defluviimonas nitratireducens sp. nov., isolated from surface seawater. Int J Syst Evol Microbiol 67:2752–2757
Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J et al (2012) SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience 1:18
Marcon J, Taketani RG, Dini-Andreote F, Mazzero GI, Soares FLJ, Melo IS et al (2014) Draft genome sequence of Bacillus thuringiensis strain BrMgv02-JM63, a chitinolytic bacterium isolated from oil-contaminated mangrove soil in Brazil. Genome Announc 2.
Meier-Kolthoff JP, Auch AF, Klenk HP, Goker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60
Minnikin DE, Patel PV, Alshamaony L, Goodfellow M (1977) Polar lipid composition in the classification of Nocardia and Related Bacteria. Int J Syst Bacteriol 27:104–117
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A et al (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Meth 2:233–241
Nagelkerken I, Blaber SJM, Bouillon S, Green P, Haywood M, Kirton LG et al (2008) The habitat function of mangroves for terrestrial and marine fauna: a review. Aquat Bot 2:155–185
Priya G, Lau N-S, Furusawa G, Dinesh B, Foong SY, Amirul AA-A (2018) Metagenomic insights into the phylogenetic and functional profiles of soil microbiome from a managed mangrove in Malaysia. Agri Gene 9:5–15
Qin QL, Xie BB, Zhang XY, Chen XL, Zhou BC, Zhou J, Oren A, Zhang YZ (2014) A proposed genus boundary for the prokaryotes based on genomic insights. J Bacteriol 196:2210–2215
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci U S A 106:19126–19131
Rodriguez-R LM, Konstantinidis K (2014) Bypassing cultivation to identify bacterial species: culture-independent genomic approaches identify credibly distinct clusters, avoid cultivation bias, and provide true insights into microbial species. Microbe Magn 9:111–118
Sasser M (2006) Bacterial identification by gas chromatographic analysis of fatty acid methyl esters GC-FAME. MIDI Technical Note.
Seppey M, Manni M, Zdobnov EM (2019) BUSCO: assessing genome assembly and annotation completeness. In: Kollmar M (ed) Gene prediction: methods and protocols. Springer, New York, pp 227–245
Shin YH, Kim JH, Suckhoom A, Kantachote D, Kim W (2017) Limibaculum halophilum gen. nov., sp. nov., a new member of the family Rhodobacteraceae. Int J Syst Evol Microbiol 67:3812–3818
Yin D, Xiao J, Ao J, Ai C, Chen X (2012) Albidovulum xiamenense sp. nov., a moderately thermophilic bacterium from a terrestrial hot spring. Int J Syst Evol Microbiol 62:1609–1612
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H et al (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Zhang R, Ju Z, Han S, Hou X, Yu Y, Zhang X et al (2019) Alkalilacustris brevis gen. nov., sp. nov., isolated from a soda lake. Int J Syst Evol Microbiol 69:1669–1675
Zhao H, Yan B, Mo S, Nie S, Li Q, Ou Q et al (2019) Carbohydrate metabolism genes dominant in a subtropical marine mangrove ecosystem revealed by metagenomics analysis. J Microbiol 7:575–586
Funding
This study was supported by the Guangxi Natural Science Foundation for Youth Scholar (No. 2018GXNSFBA050021), The Fundamental Research Funds for Guangxi Academy of Sciences (No. 2019YBJ101 and No. 2018YBJ303), Guangxi Key Laboratory of Marine Natural Products and Combinatorial Biosynthesis Chemistry (No. 17-259-74), The Special Project for the Base of Guangxi Science and Technology and Talents (Nos. AD17129019 and AD18281066).
Author information
Authors and Affiliations
Contributions
ZHL and XP: characterized the bacterium and prepare the manuscript; ZHL, FL, YH, QZH performed the phenotypic and biochemical characterization; SHH helped the analysis of taxonomic data; WH and MJ supervised the study and edited the original draft.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Ethical standards
This study does not describe any experimental work related to human.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The whole genome sequence and the 16S rRNA gene sequence of strain M0105T were deposited in GenBank/EMBL/DDBJ/PIR under Accession Numbers JAEHHL000000000 and MT613384, respectively.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pan, X., Li, Z., Li, F. et al. Thermohalobaculum xanthum gen. nov., sp. nov., a moderately thermophilic bacterium isolated from mangrove sediment. Antonie van Leeuwenhoek 114, 1819–1828 (2021). https://doi.org/10.1007/s10482-021-01641-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01641-4