Abstract
The relationship between the number of available nutrients and community diversity is a central question in ecological research that remains unanswered. Here we studied the assembly of hundreds of soil-derived microbial communities on a wide range of well-defined resource environments, from single carbon sources to combinations of up to 16. We found that, while single resources supported multispecies communities varying from 8 to 40 taxa, mean community richness increased only one-by-one with additional resources. Cross-feeding could reconcile these seemingly contrasting observations, with the metabolic network seeded by the supplied resources explaining the changes in richness due to both the identity and the number of resources, as well as the distribution of taxa across different communities. By using a consumer–resource model incorporating the inferred cross-feeding network, we provide further theoretical support to our observations and a framework to link the type and number of environmental resources to microbial community diversity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Data on 16S amplicon sequencing and metadata files have been deposited in the NCBI SRA database under NCBI BioProject ID PRJNA715195.
Code availability
Data files and analysis/simulation codes to reproduce all figures are available at https://github.com/mdalbello/Resource-diversity-relationship.
Change history
15 September 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41559-021-01563-4
References
Hubbell, S. The Unified Neutral Theory of Biodiversity and Biogeography (Princeton Univ. Press, 2001).
Tilman, D. et al. Diversity and productivity in a long-term grassland experiment. Science 294, 843–845 (2001).
Hutchinson, G. E. The paradox of the plankton. Am. Nat. 95, 137–145 (1961).
Koskella, B., Hall, L. J. & Metcalf, C. J. E. The microbiome beyond the horizon of ecological and evolutionary theory. Nat. Ecol. Evol. 1, 1606–1615 (2017).
Flemming, H. C. & Wuertz, S. Bacteria and archaea on Earth and their abundance in biofilms. Nat. Rev. Microbiol. 17, 247–260 (2019).
Hug, L. A. et al. A new view of the tree of life. Nat. Microbiol. 1, 16048 (2016).
Salazar, G. & Sunagawa, S. Marine microbial diversity. Curr. Biol. 27, R489–R494 (2017).
Huttenhower, C. et al. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Falkowski, P. G., Fenchel, T. & Delong, E. F. The microbial engines that drive earth’s biogeochemical cycles. Science 320, 1034–1039 (2008).
Kau, A. L., Ahern, P. P., Griffin, N. W., Goodman, A. L. & Gordon, J. I. Human nutrition, the gut microbiome and the immune system. Nature 474, 327–336 (2011).
Cavicchioli, R. et al. Scientists’ warning to humanity: microorganisms and climate change. Nat. Rev. Microbiol. 17, 569–586 (2019).
Blasche, S. et al. Metabolic cooperation and spatiotemporal niche partitioning in a kefir microbial community. Nat. Microbiol. 6, 196–208 (2021).
Gude, S. et al. Bacterial coexistence driven by motility and spatial competition. Nature 578, 588–592 (2020).
Kommineni, S. et al. Bacteriocin production augments niche competition by enterococci in the mammalian gastrointestinal tract. Nature 526, 719–722 (2015).
Granato, E. T., Meiller-Legrand, T. A. & Foster, K. R. The evolution and ecology of bacterial warfare. Curr. Biol. 29, R521–R537 (2019).
Ratzke, C., Barrere, J. & Gore, J. Strength of species interactions determines biodiversity and stability in microbial communities. Nat. Ecol. Evol. 4, 376–383 (2020).
Hoek, T. A. et al. Resource availability modulates the cooperative and competitive nature of a microbial cross-feeding mutualism. PLoS Biol. 14, e1002540 (2016).
Goldford, J. E. et al. Emergent simplicity in microbial community assembly. Science 361, 469–474 (2018).
Tilman, D. Resource Competition and Community Structure Vol. 17 (Princeton Univ. Press, 1982).
Gause, G. F. The Struggle for Existence (Hafner Press, 1934).
MacArthur, R. Species packing and competitive equilibrium for many species. Theor. Popul. Biol. 1, 1–11 (1970).
Levin, S. A. Community equilibria and stability, and an extension of the competitive exclusion principle. Am. Nat. 104, 413–423 (1970).
Estrela, S. et al. Metabolic rules of microbial community assembly. Preprint at bioRxiv https://doi.org/10.1101/2020.03.09.984278 (2020).
Enke, T. N. et al. Modular assembly of polysaccharide-degrading marine microbial communities. Curr. Biol. 29, 1528–1535 (2019).
Fu, H., Uchimiya, M., Gore, J. & Moran, M. A. Ecological drivers of bacterial community assembly in synthetic phycospheres. Proc. Natl Acad. Sci. USA 117, 3656–3662 (2020).
Gralka, M., Szabo, R., Stocker, R. & Cordero, O. X. Trophic interactions and the drivers of microbial community assembly. Curr. Biol. 30, R1176–R1188 (2020).
Enke, T. N. et al. Modular assembly of polysaccharide-degrading marine microbial communities. Curr. Biol. 29, 1528–1535.e6 (2019).
Martiny, J. B. H., Jones, S. E., Lennon, J. T. & Martiny, A. C. Microbiomes in light of traits: a phylogenetic perspective. Science 350, aac9323 (2015).
Naylor, D. et al. Deconstructing the soil microbiome into reduced-complexity functional modules. mBio 11, e01349-20 (2020).
MacArthur, R. H. Geographical Ecology. Patterns in the Distribution of Species (Harper & Row, 1972) .
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes. Nucleic Acids Res. 46, D633–D639 (2018).
Wang, T., Goyal, A., Dubinkina, V. & Maslov, S. Evidence for a multi-level trophic organization of the human gut microbiome. PLoS Comput. Biol. 15, e1007524 (2019).
Goyal, A. & Maslov, S. Diversity, stability, and reproducibility in stochastically assembled microbial ecosystems. Phys. Rev. Lett. 120, 158102 (2018).
Marsland, R. et al. Available energy fluxes drive a transition in the diversity, stability, and functional structure of microbial communities. PLoS Comput. Biol. 15, e1006793 (2019).
Levine, J. M. & HilleRisLambers, J. The importance of niches for the maintenance of species diversity. Nature 461, 254–257 (2009).
Tromas, N. et al. Niche separation increases with genetic distance among bloom-forming Cyanobacteria. Front. Microbiol. 9, 438 (2018).
Sriswasdi, S., Yang, C. C. & Iwasaki, W. Generalist species drive microbial dispersion and evolution. Nat. Commun. 8, 1162 (2017).
Logares, R. et al. Biogeography of bacterial communities exposed to progressive long-term environmental change. ISME J. 7, 937–948 (2013).
Monard, C., Gantner, S., Bertilsson, S., Hallin, S. & Stenlid, J. Habitat generalists and specialists in microbial communities across a terrestrial-freshwater gradient. Sci. Rep. 6, 37719 (2016).
Székely, A. J. & Langenheder, S. The importance of species sorting differs between habitat generalists and specialists in bacterial communities. FEMS Microbiol. Ecol. 87, 102–112 (2014).
Pandit, S. N., Kolasa, J. & Cottenie, K. Contrasts between habitat generalists and specialists: an empirical extension to the basic metacommunity framework. Ecology 90, 2253–2262 (2009).
Muscarella, M. E., Boot, C. M., Broeckling, C. D. & Lennon, J. T. Resource heterogeneity structures aquatic bacterial communities. ISME J. 13, 2183–2195 (2019).
Roller, B. R. K., Stoddard, S. F. & Schmidt, T. M. Exploiting rRNA operon copy number to investigate bacterial reproductive strategies. Nat. Microbiol. 1, 16160 (2016).
Goldfarb, K. C. et al. Differential growth responses of soil bacterial taxa to carbon substrates of varying chemical recalcitrance. Front. Microbiol. 2, 94 (2011).
Klappenbach, J. A., Dunbar, J. M. & Schmidt, T. M. rRNA operon copy number reflects ecological strategies of bacteria. Appl. Environ. Microbiol. 66, 1328–1333 (2000).
Rojo, F. Carbon catabolite repression in Pseudomonas: optimizing metabolic versatility and interactions with the environment. FEMS Microbiol. Rev. 34, 658–684 (2010).
Mills, C. G., Allen, R. J. & Blythe, R. A. Resource spectrum engineering by specialist species can shift the specialist-generalist balance. Theor. Ecol. 13, 149–163 (2020).
Bajic, D. & Sanchez, A. The ecology and evolution of microbial metabolic strategies. Curr. Opin. Biotechnol. 62, 123–128 (2020).
Basan, M. et al. A universal trade-off between growth and lag in fluctuating environments. Nature 584, 470–474 (2020).
Paczia, N. et al. Extensive exometabolome analysis reveals extended overflow metabolism in various microorganisms. Microb. Cell Fact. 11, 122 (2012).
Pinu, F. R. et al. Metabolite secretion in microorganisms: the theory of metabolic overflow put to the test. Metabolomics 14, 43 (2018).
Douglas, A. E. The microbial exometabolome: ecological resource and architect of microbial communities. Phil. Trans. R. Soc. B 375, 20190250 (2020).
Machado, D., Andrejev, S., Tramontano, M. & Patil, K. R. Fast automated reconstruction of genome-scale metabolic models for microbial species and communities. Nucleic Acids Res. 46, 7542–7553 (2018).
Zelezniak, A. et al. Metabolic dependencies drive species co-occurrence in diverse microbial communities. Proc. Natl Acad. Sci. USA 112, 6449–6454 (2015).
Diener, C., Gibbons, S. M. & Resendis-Antonio, O. MICOM: metagenome-scale modeling to infer metabolic interactions in the gut microbiota. mSystems 5, e00606-19 (2020).
Garza, D. R., van Verk, M. C., Huynen, M. A. & Dutilh, B. E. Towards predicting the environmental metabolome from metagenomics with a mechanistic model. Nat. Microbiol. 3, 456–460 (2018).
Pacheco, A. R., Osborne, M. L. & Segrè, D. Non-additive microbial community responses to environmental complexity. Nat Commun. 12, 2365 (2021).
Wang, X., Xia, K., Yang, X. & Tang, C. Growth strategy of microbes on mixed carbon sources. Nat. Commun. 10, 1279 (2019).
Louca, S. et al. High taxonomic variability despite stable functional structure across microbial communities. Nat. Ecol. Evol. 1, 0015 (2017).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Callahan, B. J., Sankaran, K., Fukuyama, J. A., McMurdie, P. J. & Holmes, S. P. Bioconductor workflow for microbiome data analysis: from raw reads to community analyses. F1000Research 5, 1492 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2019).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Chao, A. et al. Rarefaction and extrapolation with Hill numbers: a framework for sampling and estimation in species diversity studies. Ecol. Monogr. 84, 45–67 (2014).
Chao, A., Chiu, C. H. & Jost, L. Phylogenetic diversity measures based on Hill numbers. Phil. Trans. R. Soc. B 365, 3599–3609 (2010).
Underwood, A. J. Experiments in Ecology (Cambridge Univ. Press, 1996); https://doi.org/10.1017/cbo9780511806407
Saeedghalati, M. et al. Quantitative comparison of abundance structures of generalized communities: from B-cell receptor repertoires to microbiomes. PLoS Comput. Biol. 13, e1005362 (2017).
Goldford, J. E., Hartman, H., Smith, T. F. & Segrè, D. Remnants of an ancient metabolism without phosphate. Cell 168, 1126–1134.e9 (2017).
Raymond, J. & Segrè, D. The effect of oxygen on biochemical networks and the evolution of complex life. Science 311, 1764–1767 (2006).
Handorf, T., Ebenhoh, O. E. & Heinrich, R. Expanding metabolic networks: scopes of compounds, robustness, and evolution. J. Mol. Evol. 61, 498–512 (2005).
Ebenhoh, O., Handorf, T. & Heinrich, R. Structural analysis of expanding metabolic networks. Genome Inform. 15, 35–45 (2004).
Orth, J. D., Thiele, I. & Palsson, B. O. What is flux balance analysis? Nat. Biotechnol. 28, 245–248 (2010).
Johnson, M. et al. NCBI BLAST: a better web interface. Nucleic Acids Res. 36, 5–9 (2008).
Machado, D. et al. Polarization of microbial communities between competitive and cooperative metabolism. Nat. Ecol. Evol. 5, 195–203 (2021).
Stoddard, S. F., Smith, B. J., Hein, R., Roller, B. R. K. & Schmidt, T. M. rrnDB: improved tools for interpreting rRNA gene abundance in bacteria and archaea and a new foundation for future development. Nucleic Acids Res. 43, D593–D598 (2015).
Douglas, G. M. et al. PICRUSt2: an improved and extensible approach for metagenome inference. Preprint at bioRxiv https://doi.org/10.1101/672295 (2019).
Chen, T. & Guestrin, C. XGBoost: a scalable tree boosting system. In Proc. ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 785–794 (Association for Computing Machinery, 2016).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction 2nd edn. (Springer Series in Statistics, Springer, 2009).
Lundberg, S. M. et al. From local explanations to global understanding with explainable AI for trees. Nat. Mach. Intell. 2, 56–67 (2020).
Acknowledgements
We thank J. Grilli, M. Costantino-Lagomarsino, M. Corigliano and M. Barbier for feedback on models, the members of the Gore Lab for comments on the manuscript, and B. Stellato for help with the ensemble tree regression model. This work was supported by the Simons Collaboration: Principles of Microbial Ecosystems (PriME) award number 542395 and NIH (R01-GM102311). A.G. was supported by the Gordon and Betty Moore Foundation as a Physics of Living Systems Fellow through grant number GBMF4513.
Author information
Authors and Affiliations
Contributions
M.D.B. and J.G. conceived the study. M.D.B. performed the experiments and the sequencing analysis. H.L. performed theoretical modelling. A.G. performed metabolic and genomic analyses. All authors analysed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Ecology & Evolution thanks Marc Garcia-Garcera and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The majority of communities reached equilibrium before the end of the experiment.
Each panel shows the temporal trajectories of the composition of one community at the family level. The most prevalent 37 families are included. The first 16 plots depict communities grown on a single carbon source. The last four plots depict replicated communities grown on a media containing all the 16 carbon sources. The solid black line indicates the observed community richness.
Extended Data Fig. 2 Single carbon sources support microbial assemblages spanning a wide phylogenetic diversity.
The pool of ASVs found across all media supplied with single resources is phylogenetically diverse, encompassing 7 classes, indicated by colored lines on the right side of the plot (black lines indicate ASVs that could not be identified at any taxonomical level except the Domain, Bacteria). Families are indicated on the left side of the phylogenetic tree. Colored tiles indicate the media in which the ASV is found (for each carbon source, there are three replicated microcosms for a total of 48 communities.) Carbon sources are ordered based on the average richness they support.
Extended Data Fig. 3 All single resources supported multispecies communities, but richness varied with the identity of the resource.
Bars indicate, for each carbon source, the number of ASVs (mean ± s.e.m., N = 3).
Extended Data Fig. 4 Richness of two-resource communities is approximately the average richness of constituent single-resource communities.
a. Observed richness of each two-resource community is best approximated by the average richness of constituent single resources, compared to the maximum and the union. Both the average error for the three predictions, calculated from the absolute values of predicted minus observed richness, and the ratio between predicted and observed richness are shown. b. Average richness of two-resource communities does not differ from the average richness of single-resource communities (boxplots with median and 95 % confidence interval; number of 1-resource media = 16, number of 2-resource media = 24; each dot is obtained from the mean of 3 replicates, s.e.m. are not shown for clarity). The color of the dots indicates the supporting resource(s).
Extended Data Fig. 5 The estimated number of metabolites for combinations of carbon sources increases fast and tends to saturate with the number of supplied resources.
The number of metabolites has been computed from KEGG and MetaCyc databases (see Methods). Large colored dots indicate the average number of metabolites for each number of supplied resources (mean ± s.e.m.) while small grey dots indicate the average richness in each media containing a combination of resources (16 for single-resource, 24 for two-resource, 12 for four-resource, six for eight-resource, 16 for 15-resource and one for 16-resource combinations). Error bars are omitted for clarity. The dotted line was obtained by fitting a spline to the points.
Extended Data Fig. 6 The observed linear trend is sufficiently robust to the exclusion of low-abundance ASVs, coarse-graining at the family level, and the index used to measure microbial community diversity.
a. Richness was calculated as the number of ASVs after the exclusion of those with relative abundance lower than 0.1%. b. Richness was calculated as the number of unique families in the media. c, d. The increase in diversity, measured as Shannon Entropy and Inverse Simpson Index, with the number of carbon sources can still be approximated by a line. These indices give progressively more weight to abundant species, accounting, in this way, for the evenness of the communities. In each panel, large colored dots indicate the mean ± s.e.m. while small grey dots indicate the average richness in each media containing a combination of resources (16 for single-resource, 24 for two-resource, 12 for four-resource, six for eight-resource, 16 for 15-resource and one for 16-resource combinations). Error bars are omitted for clarity.
Extended Data Fig. 7 The evenness of the microbial communities increases with the number of supplied carbon sources.
a. Log-linear rank abundance distributions (RADs) are shown for all the experimental microbial microcosms (48 for single-resource, 72 for two-resource, 36 for four-resource, 18 for eight-resource, 16 for 15-resource and nine for 16-resource combinations) together with the fitted regression lines (black dashed lines). Going from one to 16 resources, RADs exhibit heavier tails. b. The average absolute value of the slope (bars indicate mean ± s.e.m. across replicates with the same number of supplied resources, while jittered dots represent the slope for each individual replicate) decreases with the number of supplied resources. c. Changes in evenness are independent from changes in richness, as revealed by RADs normalized for richness (mean RADs, dashed colored lines, ± SD, shaded colored ribbon, for each number of supplied resources).
Extended Data Fig. 8 Resource occupancy of the 275 ASVs found in media containing a single carbon source.
The histogram shows the number of single resources in which each ASV is found. Bars are colored depending on whether the ASV has been classified as a generalist (pink), a specialist (teal) or an intermediate (beige). The families to which ASVs belong are reported.
Extended Data Fig. 9 The fraction of habitat generalist decreases while the fraction of habitat specialists increases with community richness.
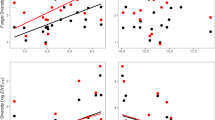
A. Percentage fraction of habitat generalists (mean, N = 3) as a function of community richness (mean ± s.e.m., N = 3). B. Percentage fraction of habitat specialists (mean, N = 3) as a function of community richness (mean ± s.e.m., N = 3). Fitted linear regression lines (black, dashed) are shown.
Extended Data Fig. 10 Richness from simulations grows modestly with the number of resources.
Richness obtained from simulations of the consumer-resource model with cross-feeding (blue triangles, mean ± s.e.m., N = 16 for single-resource, 24 for two-resource, 12 for four-resource, six for eight-resource, 16 for 15-resource and 1 for 16-resource combinations) as a linear function of the number of available resources (solid blue line, intercept = 22.7, slope = 2). Grey jittered triangles indicate the richness of communities grown on a particular resource combination.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Table 1.
Rights and permissions
About this article
Cite this article
Dal Bello, M., Lee, H., Goyal, A. et al. Resource–diversity relationships in bacterial communities reflect the network structure of microbial metabolism. Nat Ecol Evol 5, 1424–1434 (2021). https://doi.org/10.1038/s41559-021-01535-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41559-021-01535-8
This article is cited by
-
Genome-scale community modelling reveals conserved metabolic cross-feedings in epipelagic bacterioplankton communities
Nature Communications (2024)
-
Resource competition predicts assembly of gut bacterial communities in vitro
Nature Microbiology (2024)
-
A social niche breadth score reveals niche range strategies of generalists and specialists
Nature Ecology & Evolution (2023)
-
Stress-induced metabolic exchanges between complementary bacterial types underly a dynamic mechanism of inter-species stress resistance
Nature Communications (2023)
-
Carbohydrate complexity limits microbial growth and reduces the sensitivity of human gut communities to perturbations
Nature Ecology & Evolution (2023)