Abstract
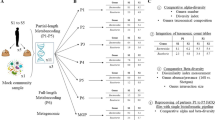
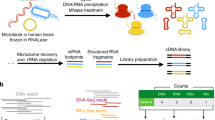
Ribosome profiling enables sequencing of ribosome-bound fragments of RNA, revealing which transcripts are being translated as well as the position of ribosomes along mRNAs. Although ribosome profiling has been applied to cultured bacterial isolates, its application to uncultured, mixed communities has been challenging. We present MetaRibo-Seq, a protocol that enables the application of ribosome profiling directly to the human fecal microbiome. MetaRibo-Seq is a benchmarked method that includes several modifications to existing ribosome profiling protocols, specifically addressing challenges involving fecal sample storage, purity and input requirements. We also provide a computational workflow to quality control and trim reads, de novo assemble a reference metagenome with metagenomic reads, align MetaRibo-Seq reads to the reference, and assess MetaRibo-Seq library quality (https://github.com/bhattlab/bhattlab_workflows/tree/master/metariboseq). This MetaRibo-Seq protocol enables researchers in standard molecular biology laboratories to study translation in the fecal microbiome in ~5 d.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The data presented in Fig. 3 were generated as part of ref. 27 and can be accessed under BioProject accession PRJNA510123.
Code availability
The pipeline implemented in this paper is available at https://github.com/bhattlab/bhattlab_workflows/tree/master/metariboseq (https://zenodo.org/record/4638134#.YGDevkhKgcg)46. This pipeline creates a de novo assembly from metagenomic reads, maps MetaRibo-Seq reads to the assembly, calls open reading frames across the assembly and creates visualizations such as metagene plots, triplet periodicity histograms and fragment length distribution for aligned MetaRibo-Seq reads. This serves as a baseline assessment of library quality and a starting point for future analysis. Though specifically designed for MetaRibo-Seq, this pipeline would also be generally useful for any situation in which both DNA-sequencing and Ribo-Seq data for an organism with no existing reference genome are available.
References
Zengel, J. M. & Lindahl, L. Diverse mechanisms for regulating ribosomal protein synthesis in Escherichia coli. Prog. Nucleic Acid Res. Mol. Biol. 47, 331–370 (1994).
Meyer, M. M. The role of mRNA structure in bacterial translational regulation. Wiley Interdiscip. Rev. RNA https://doi.org/10.1002/wrna.1370 (2017).
Lindahl, L., Jaskunas, S. R., Dennis, P. P. & Nomura, M. Cluster of genes in Escherichia coli for ribosomal proteins, ribosomal RNA, and RNA polymerase subunits. Proc. Natl Acad. Sci. USA 72, 2743–2747 (1975).
Fallon, A. M., Jinks, C. S., Strycharz, G. D. & Nomura, M. Regulation of ribosomal protein synthesis in Escherichia coli by selective mRNA inactivation. Proc. Natl Acad. Sci. USA 76, 3411–3415 (1979).
Dean, D. & Nomura, M. Feedback regulation of ribosomal protein gene expression in Escherichia coli. Proc. Natl Acad. Sci. USA 77, 3590–3594 (1980).
Li, G.-W., Burkhardt, D., Gross, C. & Weissman, J. S. Quantifying absolute protein synthesis rates reveals principles underlying allocation of cellular resources. Cell 157, 624–635 (2014).
Lalanne, J.-B. et al. evolutionary convergence of pathway-specific enzyme expression stoichiometry. Cell 173, 749–761.e38 (2018).
Bucca, G. et al. Translational control plays an important role in the adaptive heat-shock response of Streptomyces coelicolor. Nucleic Acids Res. 46, 5692–5703 (2018).
Sawyer, E. B., Grabowska, A. D. & Cortes, T. Translational regulation in mycobacteria and its implications for pathogenicity. Nucleic Acids Res. 46, 6950–6961 (2018).
Taylor, R. C. et al. Changes in translational efficiency is a dominant regulatory mechanism in the environmental response of bacteria. Integr. Biol. 5, 1393–1406 (2013).
Morita, Y., Gilmour, C., Metcalf, D. & Poole, K. Translational control of the antibiotic inducibility of the PA5471 gene required for mexXY multidrug efflux gene expression in Pseudomonas aeruginosa. J. Bacteriol. 191, 4966–4975 (2009).
Starosta, A. L., Lassak, J., Jung, K. & Wilson, D. N. The bacterial translation stress response. FEMS Microbiol. Rev. 38, 1172–1201 (2014).
Jeong, Y. et al. The dynamic transcriptional and translational landscape of the model antibiotic producer Streptomyces coelicolor A3(2). Nat. Commun. 7, 11605 (2016).
Mathieu, A. et al. Discovery and function of a general core hormetic stress response in E. coli induced by sublethal concentrations of antibiotics. Cell Rep. 17, 46–57 (2016).
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. S. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Becker, A. H., Oh, E., Weissman, J. S., Kramer, G. & Bukau, B. Selective ribosome profiling as a tool for studying the interaction of chaperones and targeting factors with nascent polypeptide chains and ribosomes. Nat. Protoc. 8, 2212–2239 (2013).
Oh, E. et al. Selective ribosome profiling reveals the cotranslational chaperone action of trigger factor in vivo. Cell 147, 1295–1308 (2011).
Li, G.-W., Oh, E. & Weissman, J. S. The anti-Shine-Dalgarno sequence drives translational pausing and codon choice in bacteria. Nature 484, 538–541 (2012).
Freeberg, L., Kuersten, S. & Syed, F. Isolate and sequence ribosome-protected mRNA fragments using size-exclusion chromatography. Nat. Methods 10, i–ii (2013).
Latif, H. et al. A streamlined ribosome profiling protocol for the characterization of microorganisms. BioTechniques 58, 329–332 (2015).
Nakahigashi, K. et al. Comprehensive identification of translation start sites by tetracycline-inhibited ribosome profiling. DNA Res 23, 193–201 (2016).
Meydan, S. et al. Retapamulin-assisted ribosome profiling reveals the alternative bacterial proteome. Mol. Cell 74, 481–493.e6 (2019).
Mangano, K. et al. Genome-wide effects of the antimicrobial peptide apidaecin on translation termination in bacteria. eLife 9, e62655 (2020).
Hwang, J.-Y. & Buskirk, A. R. A ribosome profiling study of mRNA cleavage by the endonuclease RelE. Nucleic Acids Res. 45, 327–336 (2017).
Mohammad, F., Green, R. & Buskirk, A. R. A systematically-revised ribosome profiling method for bacteria reveals pauses at single-codon resolution. eLife 8, e42591 (2019).
Fremin, B. J., Sberro, H. & Bhatt, A. S. MetaRibo-Seq measures translation in microbiomes. Nat. Commun. 11, 3268 (2020).
Mohammad, F. & Buskirk, A. R. Protocol for ribosome profiling in bacteria. Bio Protoc. 9, e3468 (2019).
Tanca, A. et al. A straightforward and efficient analytical pipeline for metaproteome characterization. Microbiome 2, 49 (2014).
Ndah, E. et al. REPARATION: ribosome profiling assisted (re-)annotation of bacterial genomes. Nucleic Acids Res. 45, e168 (2017).
Clauwaerts, J., Menschaert, G. & Waegeman, W. DeepRibo: a neural network for precise gene annotation of prokaryotes by combining ribosome profiling signal and binding site patterns. Nucleic Acids Res. 47, e36 (2019).
Sberro, H. et al. Large-scale analyses of human microbiomes reveal thousands of small, novel genes. Cell 178, 1245–1259.e14 (2019).
Durrant, M. G. & Bhatt, A. S. Automated prediction and annotation of small open reading frames in microbial genomes. Cell Host Microbe 29, 121–131.e4 (2021).
Dar, D. et al. Term-seq reveals abundant ribo-regulation of antibiotics resistance in bacteria. Science 352, aad9822 (2016).
Fremin, B. J. & Bhatt, A. S. Structured RNA contaminants in bacterial Ribo-Seq. mSphere 5, e00855-20 (2020).
Mohammad, F., Woolstenhulme, C. J., Green, R. & Buskirk, A. R. Clarifying the translational pausing landscape in bacteria by ribosome profiling. Cell Rep. 14, 686–694 (2016).
Marks, J. et al. Context-specific inhibition of translation by ribosomal antibiotics targeting the peptidyl transferase center. Proc. Natl Acad. Sci. USA 113, 12150–12155 (2016).
Zinshteyn, B., Wangen, J. R., Hua, B. & Green, R. Nuclease-mediated depletion biases in ribosome footprint profiling libraries. RNA 26, 1481–1488 (2020).
O’Loughlin, S. et al. Polysomes bypass a 50-nucleotide coding gap less efficiently than monosomes due to attenuation of a 5′ mRNA stem-loop and enhanced drop-off. J. Mol. Biol. 432, 4369–4387 (2020).
Nurk, S., Meleshko, D., Korobeynikov, A. & Pevzner, P. A. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 27, 824–834 (2017).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Chung, B. Y. et al. The use of duplex-specific nuclease in ribosome profiling and a user-friendly software package for Ribo-seq data analysis. RNA 21, 1731–1745 (2015).
Di Tommasso, P. et al. Nextflow enables reproducible computational workflows. Nat. Biotechnol. 35, 316–319 (2017).
Kurtzer, G. M., Sochat, V. & Bauer, M. W. Scientific containers for mobility of compute. PLoS One 12, e0177459 (2017).
Siranosian, B. bhattlab/bhattlab_workflows: metariboseq_v1.0.0. https://doi.org/10.5281/zenodo.4638134 (2021).
Acknowledgements
We thank D. Maghini, E. Brooks and S. Vance for helpful comments on the manuscript. This work was funded by the Damon Runyon Clinical Investigator Award, grant nos. NIH R01AI148623 and NIH R01AI143757 as well as grant no. NIH P30 AG047366, which supports the Stanford ADRC. Computational work was supported by NIH S10 Shared Instrumentation grant no. 1S10OD02014101 and by NIH grant no. P30 CA124435.
Author information
Authors and Affiliations
Contributions
B.J.F. and A.S.B. conceived the study. B.J.F performed all experiments and data analysis. C.N. wrote the Nextflow computational pipeline. B.J.F. and A.S.B. wrote the paper with input from all authors. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Fremin, B. J. et al. Nat. Commun. 11, 3268 (2020): https://doi.org/10.1038/s41467-020-17081-z
Durrant, M. G. & Bhatt, A.S. Cell Host Microbe 29, 121–131.e4 (2021): https://doi.org/10.1016/j.chom.2020.11.002
Rights and permissions
About this article
Cite this article
Fremin, B.J., Nicolaou, C. & Bhatt, A.S. Simultaneous ribosome profiling of hundreds of microbes from the human microbiome. Nat Protoc 16, 4676–4691 (2021). https://doi.org/10.1038/s41596-021-00592-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00592-4
This article is cited by
-
Protein translation: biological processes and therapeutic strategies for human diseases
Signal Transduction and Targeted Therapy (2024)
-
Small proteins: overcoming size restrictions
Nature Reviews Microbiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.