Abstract
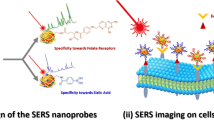
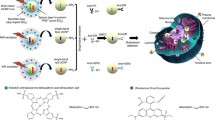
Highly selective nanoprobes have been developed based on SERS-active Au@Ag nanoparticles protected by a PEG coating and functionalized with monoclonal antibodies against human epidermal growth factor receptor 2 (HER2). The PEG coating allows to drastically reduce unspecific interactions during incubation on tissues, while the monoclonal antibodies allow a highly specific targeting of HER2. Using the designed SERS nanoprobes combined with a spectral imaging and data weighting approach, we demonstrate the proportionality between the SERS signal and the amount of HER2 antigen on the cell membranes as measured by digital image analysis of IHC staining in microscopic breast tumors (linear fit R2 = 0.87). We also show that the level of expression of HER2 measured by SERS is significantly different between several microscopic tumor parts of the same tissue slide. Therefore, SERS is proving to be a suitable technique for the localized quantitative measurement of specific markers in breast cancerous tissues. Owing to its high multiplexing capabilities, SERS could be a future tool of choice for characterizing the molecular heterogeneity of tumors at the microscopic scale.
Graphical abstract






Similar content being viewed by others
References
Pilot R, Signorini R, Durante C, Orian L, Bhamidipati M, Fabris L (2019) A review on surface-enhanced Raman scattering. Biosensors. 9:57. https://doi.org/10.3390/bios9020057
Cialla-May D, Zheng X-S, Weber K, Popp J (2017) Recent progress in surface-enhanced Raman spectroscopy for biological and biomedical applications: from cells to clinics. Chem Soc Rev 46:3945–3961. https://doi.org/10.1039/C7CS00172J
Langer J, Jimenez de Aberasturi D, Aizpurua J, Alvarez-Puebla RA, Auguié B, Baumberg JJ, Bazan GC, Bell SEJ, Boisen A, Brolo AG, Choo J, Cialla-May D, Deckert V, Fabris L, Faulds K, García de Abajo FJ, Goodacre R, Graham D, Haes AJ, Haynes CL, Huck C, Itoh T, Käll M, Kneipp J, Kotov NA, Kuang H, le Ru EC, Lee HK, Li JF, Ling XY, Maier SA, Mayerhöfer T, Moskovits M, Murakoshi K, Nam JM, Nie S, Ozaki Y, Pastoriza-Santos I, Perez-Juste J, Popp J, Pucci A, Reich S, Ren B, Schatz GC, Shegai T, Schlücker S, Tay LL, Thomas KG, Tian ZQ, van Duyne RP, Vo-Dinh T, Wang Y, Willets KA, Xu C, Xu H, Xu Y, Yamamoto YS, Zhao B, Liz-Marzán LM (2020) Present and future of surface-enhanced Raman scattering. ACS Nano 14:28–117. https://doi.org/10.1021/acsnano.9b04224
Guerrini L, Pazos-Perez N, Garcia-Rico E, Alvarez-Puebla R (2017) Cancer characterization and diagnosis with SERS-encoded particles. Cancer Nanotechnol 8. https://doi.org/10.1186/s12645-017-0031-3
Lane LA, Qian X, Nie S (2015) SERS nanoparticles in medicine: from label-free detection to spectroscopic tagging. Chem Rev 115:10489–10529. https://doi.org/10.1021/acs.chemrev.5b00265
Wang Z, Zong S, Wu L, Zhu D, Cui Y (2017) SERS-activated platforms for immunoassay: probes, encoding methods, and applications. Chem Rev 117:7910–7963. https://doi.org/10.1021/acs.chemrev.7b00027
Wang Y, Zhang Y, Schlücker S (2020) Chapter 17 - Immuno-SERS: from nanotag design to assays and microscopy. In: Ozaki Y, Baranska M, Lednev IK, B.R.B.T-V.S. in P.R. Wood (eds) . Academic Press, pp 485–528. https://doi.org/10.1016/B978-0-12-818610-7.00017-7
Salehi M, Schneider L, Ströbel P, Marx A, Packeisen J, Schlücker S (2014) Two-color SERS microscopy for protein co-localization in prostate tissue with primary antibody–protein A/G–gold nanocluster conjugates. Nanoscale. 6:2361–2367. https://doi.org/10.1039/C3NR05890E
Sun L, Sung K-B, Dentinger C, Lutz B, Nguyen L, Zhang J, Qin H, Yamakawa M, Cao M, Lu Y, Chmura AJ, Zhu J, Su X, Berlin AA, Chan S, Knudsen B (2007) Composite organic−inorganic nanoparticles as Raman labels for tissue analysis. Nano Lett 7:351–356. https://doi.org/10.1021/nl062453t
Porter MD, Lipert RJ, Siperko LM, Wang G, Narayanan R (2008) SERS as a bioassay platform: fundamentals, design, and applications. Chem Soc Rev 37:1001–1011. https://doi.org/10.1039/B708461G
Yuan Y, Panwar N, Yap SHK, Wu Q, Zeng S, Xu J, Tjin SC, Song J, Qu J, Yong K-T (2017) SERS-based ultrasensitive sensing platform: an insight into design and practical applications. Coord Chem Rev 337:1–33. https://doi.org/10.1016/j.ccr.2017.02.006
Wallace G, Masson J-F (2020) From single cells to complex tissues in applications of surface-enhanced Raman scattering. Analyst. https://doi.org/10.1039/D0AN01274B
Indrasekara ASDS, Meyers S, Shubeita S, Feldman LC, Gustafsson T, Fabris L (2014) Gold nanostar substrates for SERS-based chemical sensing in the femtomolar regime. Nanoscale. 6:8891–8899. https://doi.org/10.1039/C4NR02513J
Schütz M, Steinigeweg D, Salehi M, Kömpe K, Schlücker S (2011) Hydrophilically stabilized gold nanostars as SERS labels for tissue imaging of the tumor suppressor p63 by immuno-SERS microscopy. Chem Commun 47:4216–4218. https://doi.org/10.1039/C0CC05229A
Bhamidipati M, Lee G, Kim I, Fabris L (2018) SERS-based quantification of PSMA in tissue microarrays allows effective stratification of patients with prostate cancer. ACS Omega 3:16784–16794. https://doi.org/10.1021/acsomega.8b01839
Salehi M, Steinigeweg D, Ströbel P, Marx A, Packeisen J, Schlücker S (2013) Rapid immuno-SERS microscopy for tissue imaging with single-nanoparticle sensitivity. J Biophotonics 6:785–792. https://doi.org/10.1002/jbio.201200148
Feng J, Wu X, Ma W, Kuang H, Xu L, Xu C (2015) A SERS active bimetallic core–satellite nanostructure for the ultrasensitive detection of Mucin-1. Chem Commun 51:14761–14763. https://doi.org/10.1039/C5CC05255F
Wu L, Wang Z, Zong S, Huang Z, Zhang P, Cui Y (2012) A SERS-based immunoassay with highly increased sensitivity using gold/silver core-shell nanorods. Biosens Bioelectron 38:94–99. https://doi.org/10.1016/j.bios.2012.05.005
Monici MBT-BAR (2005) Cell and tissue autofluorescence research and diagnostic applications. Elsevier, pp 227–256. https://doi.org/10.1016/S1387-2656(05)11007-2
Schlücker S, Küstner B, Punge A, Bonfig R, Marx A, Ströbel P (2006) Immuno-Raman microspectroscopy: in situ detection of antigens in tissue specimens by surface-enhanced Raman scattering. J Raman Spectrosc 37:719–721. https://doi.org/10.1002/jrs.1534
Zhang Y, Wang X-P, Perner S, Bankfalvi A, Schlücker S (2018) Effect of antigen retrieval methods on nonspecific binding of antibody–metal nanoparticle conjugates on formalin-fixed paraffin-embedded tissue. Anal Chem 90:760–768. https://doi.org/10.1021/acs.analchem.7b03144
Schütz M, Müller CI, Salehi M, Lambert C, Schlücker S (2011) Design and synthesis of Raman reporter molecules for tissue imaging by immuno-SERS microscopy. J Biophotonics 4:453–463. https://doi.org/10.1002/jbio.201000116
Wang X-P, Zhang Y, König M, Papadopoulou E, Walkenfort B, Kasimir-Bauer S, Bankfalvi A, Schlücker S (2016) iSERS microscopy guided by wide field immunofluorescence: analysis of HER2 expression on normal and breast cancer FFPE tissue sections. Analyst. 141:5113–5119. https://doi.org/10.1039/C6AN00927A
Lutz B, Dentinger C, Sun L, Nguyen L, Zhang J, Chmura AJ, Allen A, Chan S, Knudsen B (2008) Raman nanoparticle probes for antibody-based protein detection in tissues. J Histochem Cytochem 56:371–379. https://doi.org/10.1369/jhc.7A7313.2007
Janiszewska M, Liu L, Almendro V, Kuang Y, Paweletz C, Sakr RA, Weigelt B, Hanker AB, Chandarlapaty S, King TA, Reis-Filho JS, Arteaga CL, Park SY, Michor F, Polyak K (2015) In situ single-cell analysis identifies heterogeneity for PIK3CA mutation and HER2 amplification in HER2-positive breast cancer. Nat Genet 47:1212–1219. https://doi.org/10.1038/ng.3391
Turashvili G, Brogi E (2017) Tumor heterogeneity in breast cancer. Front Med 4:227. https://doi.org/10.3389/fmed.2017.00227
Loibl S, Gianni L (2017) HER2-positive breast cancer. Lancet. 389:2415–2429. https://doi.org/10.1016/S0140-6736(16)32417-5
Arteaga CL, Sliwkowski MX, Osborne CK, Perez EA, Puglisi F, Gianni L (2012) Treatment of HER2-positive breast cancer: current status and future perspectives. Nat Rev Clin Oncol 9:16–32. https://doi.org/10.1038/nrclinonc.2011.177
Hosonaga M, Arima Y, Sampetrean O, Komura D, Koya I, Sasaki T, Sato E, Okano H, Kudoh J, Ishikawa S, Saya H, Ishikawa T (2018) HER2 heterogeneity is associated with poor survival in HER2-positive breast cancer. Int J Mol Sci 19. https://doi.org/10.3390/ijms19082158
Hildebrandt P, Stockburger M (1984) Surface-enhanced resonance Raman spectroscopy of Rhodamine 6G adsorbed on colloidal silver. J Phys Chem 88:5935–5944. https://doi.org/10.1021/j150668a038
Verdin A, Malherbe C, Müller WH, Bertrand V, Eppe G (2020) Multiplex micro-SERS imaging of cancer-related markers in cells and tissues using poly(allylamine)-coated Au@Ag nanoprobes. Anal Bioanal Chem 412:7739–7755. https://doi.org/10.1007/s00216-020-02927-8
Colpaert C, Salgado R (2007) Belgian guidelines for Her2/neu testing in breast cancer. Belgian J Med Oncol 1:22–29 http://scholar.google.com/scholar?q=related:LugmBk6Lf2UJ:scholar.google.com/&hl=en&num=20&as_sdt=0,5%5Cnpapers3://publication/uuid/63317367-27F3-4FAE-A8A8-BD8ACF8496C4
Crowe A, Yue W (2019) Semi-quantitative determination of protein expression using immunohistochemistry staining and analysis: an integrated protocol. Bio-Protocol 9. https://doi.org/10.21769/bioprotoc.3465
Kenny PA, Lee GY, Myers CA, Neve RM, Semeiks JR, Spellman PT, Lorenz K, Lee EH, Barcellos-Hoff MH, Petersen OW, Gray JW, Bissell MJ (2007) The morphologies of breast cancer cell lines in three-dimensional assays correlate with their profiles of gene expression. Mol Oncol 1:84–96. https://doi.org/10.1016/j.molonc.2007.02.004
Kato Y, Ozawa S, Miyamoto C, Maehata Y, Suzuki A, Maeda T, Baba Y (2013) Acidic extracellular microenvironment and cancer. Cancer Cell Int 13:89. https://doi.org/10.1186/1475-2867-13-89
Rahme K, Chen L, Hobbs RG, Morris MA, O’Driscoll C, Holmes JD (2013) PEGylated gold nanoparticles: polymer quantification as a function of PEG lengths and nanoparticle dimensions. RSC Adv 3:6085–6094. https://doi.org/10.1039/C3RA22739A
Nassar A, Radhakrishnan A, Cabrero IA, Cotsonis GA, Cohen C (2010) Intratumoral heterogeneity of immunohistochemical marker expression in breast carcinoma: a tissue microarray-based study. Appl Immunohistochem Mol Morphol 18. https://doi.org/10.1097/PAI.0b013e3181dddb20
Giesen C, Wang HAO, Schapiro D, Zivanovic N, Jacobs A, Hattendorf B, Schüffler PJ, Grolimund D, Buhmann JM, Brandt S, Varga Z, Wild PJ, Günther D, Bodenmiller B (2014) Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat Methods 11:417–422. https://doi.org/10.1038/nmeth.2869
Lewis JT, Ketterling RP, Halling KC, Reynolds C, Jenkins RB, Visscher DW (2005) Analysis of intratumoral heterogeneity and amplification status in breast carcinomas with equivocal (2+) HER-2 immunostaining. Am J Clin Pathol 124:273–281. https://doi.org/10.1309/J9VXABUGKC4Y07DL
Matkowskyj KA, Cox R, Jensen RT, Benya RV (2003) Quantitative immunohistochemistry by measuring cumulative signal strength accurately measures receptor number. J Histochem Cytochem 51:205–214. https://doi.org/10.1177/002215540305100209
Tadrous PJ, Siegel J, French PMW, Shousha S, Lalani E-N, Stamp GWH (2003) Fluorescence lifetime imaging of unstained tissues: early results in human breast cancer. J Pathol 199:309–317. https://doi.org/10.1002/path.1286
A. Keikhosravi, J.S. Bredfeldt, A.K. Sagar, K.W. Eliceiri, Chapter 28 - Second-harmonic generation imaging of cancer, in: J.C. Waters, T. B. T-M. in C.B. Wittman (Eds.), Quant. Imaging Cell Biol., Academic Press, 2014: pp. 531–546. https://doi.org/10.1016/B978-0-12-420138-5.00028-8.
Acknowledgements
The authors acknowledge the University Hospital Biobank of the University of Liège for providing the samples, performing the IHC staining, and obtaining high resolution images of the IHC slides.
Funding
Cedric Malherbe acknowledges the F.R.S-FNRS for funding (Research Associate fellowship).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval
All samples were provided with the approval of the University Hospital Ethics Committee (Ref 2017/236).
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Nanomaterials for biomedical imaging and targeting.
Supplementary Information
ESM 1
(DOCX 71.8 mb)
Rights and permissions
About this article
Cite this article
Verdin, A., Malherbe, C. & Eppe, G. Spatially resolved determination of the abundance of the HER2 marker in microscopic breast tumors using targeted SERS imaging. Microchim Acta 188, 288 (2021). https://doi.org/10.1007/s00604-021-04943-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-021-04943-6