Abstract
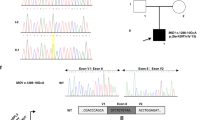
Rotor syndrome is caused by digenic loss-of-function variants in SLCO1B1 and SLCO1B3 but only a few studies have reported co-occurring inactivating variants from both genes. A rotor syndrome-causing long interspersed element-1 (LINE-1) insertion in SLCO1B3 had been reported to be highly prevalent in the Japanese population but there has been no additional report. In spite of its known association with various human diseases, LINE-1 is hard to detect with current sequencing technologies. In this study, we aimed to devise a method to screen the LINE-1 insertion variant and investigate the frequency of this variant in various populations. A chimeric sequence, that was generated by concatenating the reference sequence at the junction and a part of inserted LINE-1 sequence, was searched from 725 raw sequencing data files. In cases containing the chimeric sequence, confirmatory long-range PCR and gap-PCR were performed. In total, 95 (13.1%) of 725 patients were positive for the chimeric sequence, and all were confirmed to have the SLCO1B3 LINE-1 insertion by PCR-based tests. The same chimeric sequence was searched from the 1000 Genomes Project data repository and the carrier frequency was remarkably high in the East Asian populations (10.1%), especially in Southern Han Chinese (18.5%), but almost absent in other populations. This SLCO1B3 LINE-1 insertion should be screened in a population-specific manner under suspicion of Rotor syndrome and the methods proposed in this study would enable this in a simple way.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Ostertag EM, Kazazian HH Jr. Biology of mammalian L1 retrotransposons. Annu Rev Genet. 2001;35:501–38.
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921.
Scott AF, Schmeckpeper BJ, Abdelrazik M, Comey CT, O’Hara B, Rossiter JP, et al. Origin of the human L1 elements: proposed progenitor genes deduced from a consensus DNA sequence. Genomics. 1987;1:113–25.
Szak ST, Pickeral OK, Makalowski W, Boguski MS, Landsman D, Boeke JD. Molecular archeology of L1 insertions in the human genome. Genome Biol. 2002;3:1–18.
Sassaman DM, Dombroski BA, Moran JV, Kimberland ML, Naas TP, DeBerardinis RJ, et al. Many human L1 elements are capable of retrotransposition. Nat Genet. 1997;16:37–43.
Brouha B, Schustak J, Badge RM, Lutz-Prigge S, Farley AH, Moran JV, et al. Hot L1s account for the bulk of retrotransposition in the human population. Proc Natl Acad Sci USA. 2003;100:5280–5.
Hancks DC, Kazazian HH Jr. Roles for retrotransposon insertions in human disease. Mob DNA 2016;7:9.
Zhang X, Zhang R, Yu J. New understanding of the relevant role of LINE-1 retrotransposition in human disease and immune modulation. Front Cell Dev Biol. 2020;8:657.
Rotor AB. Familial non-hemolytic jaundice with direct van den Bergh reaction. Acta Med Philip. 1948;5:37–49.
van de Steeg E, Stránecký V, Hartmannová H, Nosková L, Hřebíček M, Wagenaar E, et al. Complete OATP1B1 and OATP1B3 deficiency causes human rotor syndrome by interrupting conjugated bilirubin reuptake into the liver. J Clin Investig. 2012;122:519–28.
Jirsa M, Knisely A, Schinkel A, Kmoch S. Rotor syndrome. In: GeneReviews®[Internet]. University of Washington, Seattle; 2019.
Kagawa T, Oka A, Kobayashi Y, Hiasa Y, Kitamura T, Sakugawa H, et al. Recessive inheritance of population-specific intronic LINE-1 insertion causes a rotor syndrome phenotype. Hum Mutat. 2015;36:327–32.
1000 Genomes Project Consortium, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, Korbel JO, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Stenson PD, Mort M, Ball EV, Evans K, Hayden M, Heywood S, et al. The human gene mutation database: towards a comprehensive repository of inherited mutation data for medical research, genetic diagnosis and next-generation sequencing studies. Hum Genet. 2017;136:665–77.
Thorvaldsdóttir H, Robinson JT, Mesirov JP. Integrative genomics viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 2013;14:178–92.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Kim YG, Kim MJ, Lee JS, Lee JA, Song JY, Cho SI, et al. SnackVar: an open-source software for sanger sequencing analysis optimized for clinical use. J Mol Diagn. 2021;23:140–8.
Steranka JP, Tang Z, Grivainis M, Huang CRL, Payer LM, Rego FOR, et al. Transposon insertion profiling by sequencing (TIPseq) for mapping LINE-1 insertions in the human genome. Mob DNA. 2019;10:8.
Zhou D, Qi S, Zhang W, Wu L, Xu A, Li X, et al. Insertion of LINE-1 retrotransposon inducing exon inversion causes a rotor syndrome phenotype. Front Genet. 2019;10:1399.
Yamakawa Y, Hamada A, Nakashima R, Yuki M, Hirayama C, Kawaguchi T, et al. Association of genetic polymorphisms in the influx transporter SLCO1B3 and the efflux transporter ABCB1 with imatinib pharmacokinetics in patients with chronic myeloid leukemia. Ther Drug Monit. 2011;33:244–50.
Ren Q, Han X, Ren J, Liu X, Ji L. Influence of the SLCO1B3 gene on sulfonylurea failure in patients with type 2 diabetes in China. Exp Clin Endocrinol Diabetes. 2017;125:449–53.
Tounsi N, Trabelsi I, Kerkeni E, Grissa MH, Fredj N, Sekma A, et al. ABCB1 and SLCO1B3 gene polymorphisms and their impact on digoxin pharmacokinetics in atrial fibrillation patients among the tunisian population. Pharmacology. 2017;99:250–8.
Tague LK, Byers DE, Hachem R, Kreisel D, Krupnick AS, Kulkarni HS, et al. Impact of SLCO1B3 polymorphisms on clinical outcomes in lung allograft recipients receiving mycophenolic acid. Pharmacogenomics J. 2020;20:69–79.
Abe T, Unno M, Onogawa T, Tokui T, Kondo TN, Nakagomi R, et al. LST-2, a human liver-specific organic anion transporter, determines methotrexate sensitivity in gastrointestinal cancers. Gastroenterology. 2001;120:1689–99.
Smith NF, Acharya MR, Desai N, Figg WD, Sparreboom A. Identification of OATP1B3 as a high-affinity hepatocellular transporter of paclitaxel. Cancer Biol Ther. 2005;4:815–8.
Lee HH, Leake BF, Teft W, Tirona RG, Kim RB, Ho RH. Contribution of hepatic organic anion-transporting polypeptides to docetaxel uptake and clearance. Mol Cancer Ther. 2015;14:994–1003.
Lancaster CS, Sprowl JA, Walker AL, Hu S, Gibson AA, Sparreboom A. Modulation of OATP1B-type transporter function alters cellular uptake and disposition of platinum chemotherapeutics. Mol Cancer Ther. 2013;12:1537–44.
Yamaguchi H, Kobayashi M, Okada M, Takeuchi T, Unno M, Abe T, et al. Rapid screening of antineoplastic candidates for the human organic anion transporter OATP1B3 substrates using fluorescent probes. Cancer Lett. 2008;260:163–9.
Hu S, Franke RM, Filipski KK, Hu C, Orwick SJ, de Bruijn EA, et al. Interaction of imatinib with human organic ion carriers. Clin Cancer Res. 2008;14:3141–8.
Sun R, Ying Y, Tang Z, Liu T, Shi F, Li H, et al. The emerging role of the SLCO1B3 protein in cancer resistance. Protein Pept Lett. 2020;27:17–29.
de Morrée ES, Böttcher R, van Soest RJ, Aghai A, de Ridder CM, Gibson AA, et al. Loss of SLCO1B3 drives taxane resistance in prostate cancer. Br J Cancer. 2016;115:674–81.
Kagawa T, Adachi Y, Hashimoto N, Mitsui H, Ohashi T, Yoneda M, et al. Loss of organic anion transporting polypeptide 1B3 function causes marked delay in indocyanine green clearance without any clinical symptoms. Hepatology. 2017;65:1065–8.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
10038_2021_967_MOESM1_ESM.xlsx
Supplemental Table S1. List of individuals and files included from 1000 Genomes Project (File access : ftp.1000genomes.ebi.ac.uk/vol1/ftp/phase3/data, last accessed 10 June, 2021)
10038_2021_967_MOESM3_ESM.xlsx
Supplementary Table S3. BAM file findings and confirmatory PCR results for the detecion of intronic LINE-1 insertion in SLCO1B3 (95 positive cases from SNUH repository data)
10038_2021_967_MOESM5_ESM.xlsx
Supplementary Table S5. SLCO1B1 c.1738C>T, p.R580* variant positive individuals in 1000 Genomes Project and their predicted status of SLCO1B3 LINE-1 insertion revealed in this study
Rights and permissions
About this article
Cite this article
Kim, Yg., Sung, H., Shin, H.S. et al. Intronic LINE-1 insertion in SLCO1B3 as a highly prevalent cause of rotor syndrome in East Asian population. J Hum Genet 67, 71–77 (2022). https://doi.org/10.1038/s10038-021-00967-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-021-00967-1
This article is cited by
-
Prevalence and founder effect of DRC1 exon 1–4 deletion in Korean patients with primary ciliary dyskinesia
Journal of Human Genetics (2023)