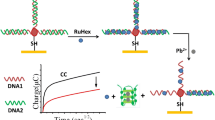
Abstract
Films of four different DNA quadruplex-forming (G4) sequences (c-KIT, c-MYC, HTelo, and BCL2) on gold surfaces were investigated by electrochemical impedance spectroscopy (EIS) to evaluate whether they evoke unique electrochemical responses that can be used for their identification. This could render EIS an alternative means for the determination of G4 sequences of unknown structure. Towards, this end, cation-dependent topology changes in the presence of either K+, K+ in combination with Li+, or Pb2+ in the presence of Li+ were first evaluated by circular dichroism (CD) spectroscopy, and electrochemical studies were performed subsequently. As a result, G4-sequence specific charge transfer resistance (RCT) patterns were in fact observed for each G4 sequence, allowing their discrimination by EIS.
Graphic abstract

Similar content being viewed by others
Introduction
Guanine-rich oligonucleotide sequences can adopt tetra-stranded structures referred to as quadruplex (G4) DNA [1]. The formation of G4 DNA is based on the self-assembly of four guanine (G) nucleobases in the form of tetrads, with a centrally positioned (mostly monovalent) cation coordinating the guanine keto groups (Fig. 1) [2]. Within G4 DNA, individual tetrads are located adjacent to each other, allowing the nucleobases to interact via π stacking. G4 DNA has been identified as a potential target in a novel approach to cancer therapy [3]. Moreover, the triggered formation of G4 DNA has been successfully applied to develop sensors for the respective trigger molecules [4]. Finally, it is an important player in DNA nanotechnology [5].
Structural studies have shown that G4 DNA can exist in a variety of topologies [6]. Circular dichroism (CD) spectroscopy represents a prominent tool to distinguish between these topologies [7]. The quadruplex topology is affected by the presence and the identity of its central cations. Among the alkali and alkaline earth metal ions, K+ ions exhibit the strongest G4 stabilization, whereas Li+ and Cs+ ions are only weakly stabilizing [8,9,10,11,12,13,14,15]. In addition, further cations including NH4+ [16], Tb3+ [17] or transition metal ions such as Co2+, Zn2+ and other divalent ions [12, 18] have been investigated. Amongst these, Pb2+ ions have attracted particular attention. Several studies have revealed a high affinity of Pb2+ towards G4 DNA [19,20,21,22].
The aim of this study is to evaluate whether electrochemical impedance spectroscopy (EIS) may act as a means to determine the identity of a given G4 DNA sequence. Towards this end, the charge transfer resistance (RCT) of a film of surface-deposited G4 DNA was measured under different experimental conditions and correlated with the corresponding CD spectroscopic results.
Materials and methods
G4 DNA oligonucleotides were synthesized using automated solid-phase synthesis and purified as reported previously [23]. All phosphoramidites were purchased from Glen Research. The identity of the desired G4 sequences was confirmed by MALDI-TOF mass spectrometry.
CD-spectroscopic studies were performed on a J-815 CD spectrometer (Jasco) using a quartz cuvette with 1 cm path length at 5 °C. The G4 solutions contained 1 μM DNA, 5 mM 3-(N-morpholino)propanesulfonic acid (MOPS), adjusted with either KOH or LiOH to pH 6.8, and 60 mM KCl or LiClO4, respectively. For measurements in the presence of Pb2+, Pb(NO3)2 was used.
Electrochemical impedance spectroscopy studies and cleaning of the electrodes were performed as reported previously [24,25,26,27]. The measurement solution comprised 2 mM K4[Fe(CN)6] and 2 mM K3[Fe(CN)6], 5 mM MOPS (pH 6.8), and 60 mM KCl or LiClO4. Freshly cleaned gold electrodes were incubated in a 25 μM aqueous solution of G4 DNA in either K+- or Li+-containing buffer (5 mM MOPS (pH 6.8), 60 mM KCl or LiClO4). After incubation for 3 d at 5 °C, electrodes were carefully washed with the respective buffer solution and subsequently incubated in MCH (1 mM in K+- or Li+-containing buffer) for 1 h at room temperature.
Nyquist plots of the obtained EIS data were fitted by an equivalent circuit (R(CR(R(CPE))) with R = resistor, C = capacitor and CPE = constant phase element) which had been used previously for similar studies (Scheme S3-1, Supp. Inf.) [24,25,26]. Moreover, preliminary fits using other equivalent circuits (e.g. a Randles circuit without the Warburg element (Scheme S3-2a, Supp. Inf.) or an equivalent circuit in which the capacitor was replaced by a CPE (Scheme S3-2b, Supp. Inf.) have shown that the equivalent circuit considered in this work gives the best fit of experimental and calculated data (Tables S3-3–S3-5, Supp. Inf.). This is particularly true for the fit of the high-frequency region, as can clearly be discerned from the respective Bode plots (Fig. S3-2, Supp. Inf.). Although the equivalent circuit R(CR(R(CPE))) seems to be overfitting the data in a few cases (Tables S3-3 and S3-4, Supp. Inf.), this circuit was used since it is the only one with a good fit of the low- and high-frequency regions (Fig. S3-2, Supp. Inf.). Moreover, using the same model throughout allows a better comparison of the parameters.
Results and discussion
Choice of nucleic acid sequences
A total of four G4 DNA sequences were studied by electrochemical impedance spectroscopy. In particular, G-rich sequences from the promoter regions of the c-KIT [28, 29], c-MYC [30, 31] and BCL2 [32] genes as well as the human telomeric DNA sequence HTelo [33] were selected (Table 1), as the structures of these G4 DNA sequences are known from single-crystal X-ray diffraction analysis or from solution NMR studies. Moreover, they represent relevant therapeutic targets [34]. While parallel-stranded G4 topologies exist in c-KIT, c-MYC and HTelo in the presence of K+ [28,29,30,31, 33], a [3 + 1] strand orientation is known to be adopted by BCL2 [32]. Their respective experimental structures are depicted in Fig. 2.
CD-spectroscopic characterization
Prior to the electrochemical characterization of the surface-deposited G4 DNA films, their folding in solution was evaluated by CD spectroscopy. This allows a direct comparison of the charge transfer resistance of the DNA films with more common descriptors such as CD spectra. Due to the cation dependence of the G4 structures, it was decided to investigate the quadruplexes in the presence of different metal cations. In this context, it is important to consider that the redox probe used in the electrochemical studies consists of an equimolar mixture of 2 mM solutions of K4[Fe(CN)6] and K3[Fe(CN)6]. Hence, the presence of the excellent G4 binder K+ in significant amounts (14 mM) was inevitable. For that reason, a variety of experimental conditions were chosen for the CD spectroscopic and the EIS studies. These are a) the presence of K+ ions alone, b) the presence of the poor G4 binder Li+ alone, c) a combination of K+ and Li+ ions, and d) increasing amounts of the strong G4 binder Pb2+.
For the CD-spectroscopic characterization, G4 DNA oligonucleotides (c = 1 μM) were initially studied in the presence of either K+ (60 mM) or Li+ (60 mM), respectively, at pH 6.8 (5 mM MOPS buffer). Additional measurements were performed using a mixed solution of 14 mM K+ and 60 mM Li+. Furthermore, Pb2+ titrations were carried out with Li+-containing G4 DNA in the absence of K+ to identify the concentration required to induce a cation exchange within the G4 quadruplex. Their respective stabilities in the presence of 14 mM K+ alone were examined as well. As anticipated, different binding behaviors were observed depending on the G4 sequence.
The CD spectra of c-KIT are depicted in Fig. 3. The wavelengths of the minima and maxima are almost the same for all measurements in the absence of Pb2+, but the intensities vary. This suggests an altering stability but no change of the overall G4 topology. As expected, c-KIT DNA is most stable in the presence of K+ ions and is destabilized by Li+ ions [13]. If both cations are available (K+:Li+ = 14 mM:60 mM), the G4 structure is stabilized again. However, due to the equilibrium of K+ and Li+ ions, the quadruplex DNA does not reach the same stability as found for the G4 DNA in the presence of K+ only. Upon the addition of Pb2+ to the Li+-containing G4 DNA, an additional maximum appears at around 315 nm, indicating a cation exchange [19, 22, 25]. Based on this new maximum, it can be concluded that 50 μM Pb2+ are required to induce the changes in the case of c-KIT. In the presence of 150 μM Pb2+, a stable G4 DNA structure is formed (Fig. S2-1, Supp. Inf.), which is again slightly destabilized upon the addition of 14 mM K+. Nevertheless, even after four days the characteristic CD signature of G4 DNA with Pb2+ at ca. 315 nm remains clearly visible.
In contrast to c-KIT, the CD spectrum of c-MYC in the presence of Li+ and K+ ions more closely resembles that of c-MYC with only K+ ions (Fig. 4). This indicates higher stability of the K+-containing c-MYC quadruplex compared to the respective c-KIT quadruplex in the presence of a mixture of K+ and Li+. Interestingly, the maximum at 310 nm, which appears to be characteristic for the interaction between G4 DNA and Pb2+, is strongly pronounced immediately after the addition of Pb2+ but decreases over time until reaching a constant level after four hours (Fig. S2-2a, Supp. Inf.). This observation is in line with a structural transition via a kinetically favored G4 intermediate towards the thermodynamically favored product. Based on the decline of the maximum at 310 nm, it is likely that different G4 structures resulting from the presence of K+, Li+ and Pb2+ exist at the same time in equilibrium while the formation of the Pb2+-containing G4 topology is favored in initially. Such a concentration-dependent coexistence of different G4 DNA topologies with different cations as well as the formation of intermediates with mixed cation occupation has been shown before [37]. Initially, the subsequent addition of 14 mM K+ has a minor affect only, but already 15 min later a destabilization of the Pb2+-containing G4 is observed (Fig. S2-2b, Supp. Inf.). After 13 h, all Pb2+ ions appear to have been replaced by K+ and Li+ ions, as indicated by the lack of a CD signal at 310 nm and the similarity with the spectrum obtained in the presence of 14 mM K+ and 60 mM Li+.
CD spectra of c-MYC in the presence of 60 mM K+ (dotted black line), 60 mM Li+ (solid black line), or 14 mM K+ and 60 mM Li+ (red). The c-MYC G4 solution was measured immediately after the addition of 20 μM Pb2+ with 60 mM Li+ present (yellow) and again 8 h later (green–brown). Once no further changes were observed, 14 mM K+ was added and additional spectra were recorded immediately (green) and after 13 h (blue)
HTelo G4 DNA undergoes significant topological changes depending on the available cations (Fig. 5). The G4 topology adopted in the presence of Li+ is obviously different from the one in the presence of either only K+ or K+ and Li+. In contrast to the other three G4 DNA sequences under investigation, a complete change of conformation was observed for HTelo after the addition of 20 μM Pb2+ in the presence of Li+ and K+. Higher concentrations of Pb2+ do not affect the G4 structure further (Fig. S2-3, Supp. Inf.). The subsequent addition of 14 mM K+ destabilizes that G4 topology immediately, and the changes continue until a stable G4 conformation is reached after 15 h. The final CD spectrum neither fits that of HTelo in the presence of K+ or Li+ alone nor that obtained in the presence of more than one cation. It can therefore be concluded that in the concurrent presence of Pb2+, K+ and Li+, HTelo adopts a conformation different from those seen before. Based on what has been discussed before, it can be assumed that Li+ plays only a minor role here.
CD spectra of HTelo in the presence of 60 mM K+ (dotted black line), 60 mM Li+ (solid black line), or 14 mM K+ and 60 mM Li+ (red). Additional spectra were measured after the subsequent addition of 30 μM Pb2+ in the absence (yellow) or presence of 14 mM K+, either immediately after the addition of K+ (green) of 15 h later (blue)
The CD spectra of BCL2 (Fig. 6) in the absence of Pb2+ are similar to those of c-KIT. Again, the G4 conformation is affected by the presence of both K+ and Li+. Upon the addition of Pb2+, a time-dependent change of the maximum at 315 nm similar to that seen for c-MYC is observed. Furthermore, the changes start to occur after the addition of 10 μM Pb2+ for both quadruplexes. However, in contrast to the latter, the maximum is still present after 4 days, but significantly smaller than before.
Electrochemical impedance spectroscopy
To investigate the differences in RCT through the quadruplex film, a solution containing the anionic redox couple [Fe(CN)6]3–/4– was used. Earlier studies by scanning electrochemical microscopy and EIS of DNA films on Au surfaces using the [Fe(CN)6]3–/4– redox couple demonstrated that metal interactions with DNA give rise to reproducible changes in the diffusion of the redox probe in and out of the thin film [38, 39]. The impedance of DNA films is highly sensitive to such changes and thus has been used extensively to study DNA films including their interaction with metal ions [26].
Thin films of quadruplex DNA on Au electrodes were prepared by immersing the Au electrodes into disulfide-modified DNA solution. The 5ʹ-disulfide group ensures chemisorption of the oligonucleotides onto the Au surface [24, 40,41,42,43,44,45,46]. Next, 6-mercaptohexanol (MCH) was used to backfill the DNA film to achieve a more uniform orientation of the oligonucleotides and to reduce potential film defects and pinholes [27, 43, 45]. The result is a heterogeneous thin film, and a schematic representation of the modified Au surface is depicted in Fig. 7. The fact that the DNA is indeed attached to the surface can be deduced from the observation that a DNA-free MCH-covered Au electrode shows an electrochemical response distinct from that of the DNA-covered and MCH-backfilled surface (Fig. S3-6, Supp. Inf.).
The trends observed for the different G4 DNA oligonucleotides in the EIS studies resemble those of the CD spectroscopic experiments. Representative Nyquist plots (− Zim vs. Zre) together with fits to an equivalent circuit for thin films of c-KIT are shown in Fig. 8 and show a significant change in the impedance as a result of K+, Li+, and Pb2+ interactions with the G4 film. Fitting of the electrochemical impedance using a modified Randles’ equivalent circuit allows extracting the charge transfer resistance RCT as well as the film capacitance Cfilm. The equivalent circuit contains a constant phase element acting as a non-ideal capacitor. This is commonly used to account for the heterogeneity of the film on polycrystalline electrodes [47]. Rx is attributed to residual pinholes and defects in the film structure. All Nyquist plots are given in the Supporting Information (Fig. S3-3–S3-5), the electrochemical impedance parameters are summarized in Table S3-1 (Supp. Inf.). Solution resistances RS are not included as they are negligible (RS = 10–60 Ω cm2). Changes in the RCT are responsible for the overall changes in the film impedance and in the following, the changes in RCT induced by the presence of different cations to the four G4 DNA sequences under investigation will be discussed (Fig. 9).
Representative Nyquist plots of a gold surface modified with c-KIT G4 DNA in the presence of K+-containing buffer solution (gray), of Li+-containing buffer solution before (light gray) and after incubation (black) of the latter in Pb2+-containing solution. Experimental data (■) are fitted with calculated impedance curves based on the circuit model shown in the inset (solid lines). For details, please see Table S3-1 (Supp. Inf.)
RCT values for G4 films on Au electrodes in the presence of K+-containing buffer solution (black), Li+-containing buffer solution (gray) or Li+-containing buffer solution in the presence of Pb2+ (dotted). The Li+-containing buffer inevitably comprises 14 mM K+ due to the presence of K4[Fe(CN)6] and K3[Fe(CN)6] in the measurement solution. Data represent an average of three individual measurements
The charge transfer resistance of DNA films composed of c-KIT, c-MYC, and BCL2 is reduced in the presence of both Li+ and K+ compared to K+ alone. Only the DNA films containing HTelo show almost identical RCT values in the absence and presence of Li+, taking into account the respective standard deviations. This matches the findings of the CD-spectroscopic studies. While the CD spectra of c-KIT, c-MYC, and BCL2 in the presence of Li+ and K+ differ from those recorded for only K+, they are comparable in the case of HTelo. This also confirms that the influence of the different anions (chloride vs. perchlorate) is negligible. Interestingly, RCT values for c-KIT and c-MYC are indistinguishable in the presence of buffers containing either K+ or K+ and Li+. BCL2 on the other hand shows a significantly higher charge transfer resistance RCT in both buffers.
After incubation with Pb2+, an increase in RCT is observed for c-KIT, HTelo and BCL2, albeit rather small in the latter case. This increase in RCT is in good agreement with prior studies [25, 26, 48]. However, no change is observed for c-MYC. This finding is not unexpected, because the G4 topology in the presence of Pb2+ is relatively unstable, leading to an almost full recovery of the G4 topology present in Li+-containing buffer solution with K+ (Fig. 4). While it took 13 h to reach this conformation in solution (as derived from the CD spectra), a similar effect was observed electrochemically after a shorter period of time. It can be speculated that the immobilization of c-MYC DNA on Au surface affects either the binding affinity or the binding kinetics. This could also be an explanation for the unexpectedly low increase in RCT for BCL2. If the respective metal-binding capabilities are indeed the reason for the differences in the electrochemical response, then it can be assumed that either the films of c-KIT and HTelo quadruplexes release their Pb2+ cations more slowly or that the G4 conformations adopted in the presence of Pb2+ are more stable for films of c-KIT and HTelo compared to c-MYC and BCL2. Overall, these sequence-dependent differences in RCT enable electrochemical discrimination of c-KIT, c-MYC, HTelo and BCL2 with a set of three measurements (K+, Li+, and Pb2+).
Conclusion
We have shown that cation-related conformational changes of the G4-forming oligonucleotide sequences c-KIT, c-MYC, HTelo, and BCL2 cannot only be followed by CD spectroscopy, but also by electrochemical impedance spectroscopy using films of G4 DNA attached to a gold surface. Each G4 sequence shows an individual characteristic binding behavior towards K+, Li+ or Pb2+, resulting in sequence-specific changes of the charge transfer resistance RCT. In addition, significant differences in RCT in the presence of Pb2+ were observed. While c-MYC has a high affinity towards K+, leading to a fast cation exchange of Pb2+ by K+ and hence no change in RCT, CD-spectroscopic and EIS studies of c-KIT, HTelo and BCL2 confirm that their G4 conformations are reasonably affected by the presence of Pb2+. As less Pb2+ was needed to induce conformational changes in HTelo compared to c-KIT, the former sequence may be more sensitive towards Pb2+, even though the percentaged change of RCT upon the addition of Pb2+ was higher for c-KIT. As summarized in Table 2, a combination of three measurements of RCT in the presence of different combinations of cations represents a helpful tool for identifying the G4 sequence attached to the Au surface.
References
Gellert M, Lipsett MN, Davies DR (1962) Helix formation by guanylic acid. Proc Natl Acad Sci USA 48:2013–2018
Müller J (2010) Functional metal ions in nucleic acids. Metallomics 2:318–327
Neidle S (2017) Quadruplex nucleic acids as targets for anticancer therapeutics. Nat Rev Chem 1:0041
Chiorcea-Paquim A-M, Eritja R, Oliveira-Brett AM (2018) Electrochemical and AFM characterization of G-quadruplex electrochemical biosensors and applications. J Nucleic Acids 2018:5307106
Mergny J-L, Sen D (2019) DNA quadruple helices in nanotechnology. Chem Rev 119:6290–6325
Phan AT (2010) Human telomeric G-quadruplex: structures of DNA and RNA sequences. FEBS J 277:1107–1117
del Villar-Guerra R, Trent JO, Chaires JB (2018) G-quadruplex secondary structure obtained from circular dichroism spectroscopy. Angew Chem Int Ed 57:7171–7175
Hardin CC, Watson T, Corregan M, Bailey C (1992) Cation-dependent transition between the quadruplex and Watson-crick hairpin forms of d(CGCG3GCG). Biochemistry 31:833–841
Venczel EA, Sen D (1993) Parallel and antiparallel G-DNA structures from a complex telomeric sequence. Biochemistry 32:6220–6228
Deepa P, Kolandaivel P, Senthilkumar K (2011) Structural properties and the effect of interaction of alkali (Li+, Na+, K+) and alkaline earth (Be2+, Mg2+, Ca2+) metal cations with G and SG-tetrads. Comput Theor Chem 974:57–65
Chen F-M (1992) Sr2+ Facilitates intermolecular G-quadruplex formation of telomeric sequences. Biochemistry 31:3769–3776
Bhattacharyya D, Arachchilage GM, Basu S (2016) Metal cations in G-quadruplex folding and stability. Front Chem 4:38
Nový J, Böhm S, Králová J, Král V, Urbanová M (2008) Formation and temperature stability of G-quadruplex structures studied by electronic and vibrational circular dichroism spectroscopy combined with Ab initio calculations. Biopolymers 89:144–152
You J, Li H, Lu X-M, Li W, Wang P-Y, Dou S-X, Xi X-G (2017) Effects of monovalent cations on folding kinetics of G-quadruplexes. Biosci Rep 37:BSR20170771
Lee JS (1990) The stability of polypurine tetraplexes in the presence of mono- and divalent cations. Nucleic Acids Res 18:6057–6060
Nagesh N, Chatterji D (1995) Ammonium ion at low concentration stabilizes the G-quadruplex formation by telomeric sequence. J Biochem Biophys Methods 30:1–8
Nagesh N, Bhargava P, Chatterji D (1992) Terbium(III)-induced fluorescence of four-stranded G4-DNA. Biopolymers 32:1421–1424
Miyoshi D, Nakao A, Toda T, Sugimoto N (2001) Effect of divalent cations on antiparallel G-quartet structure of d(G4T4G4). FEBS Lett 496:128–133
Liu W, Zhu H, Zheng B, Cheng S, Fu Y, Li W, Lau T-C, Liang H (2012) Kinetics and mechanism of G-quadruplex formation and conformational switch in a G-quadruplex of PS2.M induced by Pb2+. Nucleic Acids Res 40:4229–4236
Kotch FW, Fettinger JC, Davis JT (2000) A lead-filled G-quadruplex: insight into the G-quartet’s selectivity for Pb2+ over K+. Org Lett 2:3277–3280
Smirnov I, Shafer RH (2000) Lead is unusually effective in sequence-specific folding of DNA. J Mol Biol 296:1–5
Li T, Wang E, Dong S (2009) Potassium-lead-switched G-quadruplexes: a new class of DNA logic gates. J Am Chem Soc 131:15082–15083
Megger DA, Fonseca Guerra C, Hoffmann J, Brutschy B, Bickelhaupt FM, Müller J (2011) Contiguous metal-mediated base pairs comprising two AgI ions. Chem Eur J 17:6533–6544
Léon JC, She Z, Kamal A, Shamsi MH, Müller J, Kraatz H-B (2017) DNA films containing the artificial nucleobase imidazole mediate charge transfer in a Silver(I)-responsive way. Angew Chem Int Ed 56:6098–6102
Lin Z, Chen Y, Li X, Fang W (2011) Pb2+ induced DNA conformational switch from hairpin to G-quadruplex: electrochemical detection of Pb2+. Analyst 136:2367–2372
Lin Z, Li X, Kraatz H-B (2011) Impedimetric immobilized DNA-based sensor for simultaneous detection of Pb2+, Ag+, and Hg2+. Anal Chem 83:6896–6901
Li C, Li X, Liu X, Kraatz H-B (2010) Exploiting the interaction of metal ions and peptide nucleic acids-DNA duplexes for the detection of a single nucleotide mismatch by electrochemical impedance spectroscopy. Anal Chem 82:1166–1169
Phan AT, Kuryavyi V, Burge S, Neidle S, Patel DJ (2007) Structure of an unprecedented G-quadruplex scaffold in the human c-kit promoter. J Am Chem Soc 129:4386–4392
Wei D, Parkinson GN, Reszka AP, Neidle S (2012) Crystal structure of a c-kit promoter quadruplex reveals the structural role of metal ions and water molecules in maintaining loop conformation. Nucleic Acids Res 40:4691–4700
Dai J, Carver M, Hurley LH, Yang D (2011) Solution structure of a 2:1 quindoline-c-MYC G-quadruplex: insights into G-quadruplex-interactive small molecule drug design. J Am Chem Soc 133:17673–17680
Ambrus A, Chen D, Dai J, Jones RA, Yang D (2005) Solution structure of the biologically relevant G-quadruplex element in the human c-MYC promoter implications for G-quadruplex stabilization. Biochemistry 44:2048–2058
Dai J, Chen D, Jones RA, Hurley LH, Yang D (2006) NMR solution structure of the major G-quadruplex structure formed in the human BCL2 promoter region. Nucleic Acids Res 34:5133–5144
Parkinson GN, Lee MPH, Neidle S (2002) Crystal structure of parallel quadruplexes from human telomeric DNA. Nature 417:876–880
Neidle S (2016) Quadruplex nucleic acids as novel therapeutic targets. J Med Chem 59:5987–6011
Dai J, Carver M, Punchihewa C, Jones RA, Yang D (2007) Structure of the hybrid-2 type intramolecular human telomeric G-quadruplex in K+ solution: insights into structure polymorphism of the human telomeric sequence. Nucleic Acids Res 35:4927–4940
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Yu Z, Zhou W, Ma G, Li Y, Fan L, Li X, Lu Y (2018) Insights into the competition between K+ and Pb2+ binding to a G-quadruplex and discovery of a novel k+−Pb2+−quadruplex intermediate. J Phys Chem B 122:9382–9388
Liu B, Bard AJ, Li C-Z, Kraatz H-B (2005) Scanning electrochemical microscopy. 51. Studies of self-assembled monolayers of DNA in the absence and presence of metal ions. J Phys Chem B 109:5193–5198
Diakowski PM, Kraatz H-B (2009) Detection of single-nucleotide mismatches using scanning electrochemical microscopy, Chem. Commun. 1189–1191
Grönbeck H, Curioni A, Andreoni W (2000) Thiols and disulfides on the Au(111) surface: the headgroup-gold interaction. J Am Chem Soc 122:3839–3842
Rivera-Gandía J, del Mar Maldonado M, De La Torre-Meléndez Y, Ortiz-Quiles EO, Vargas-Barbosa NM, Cabrera CR (2011) Electrochemical capacitance DNA sensing at Hairpin-Modified Au electrodes. J Sensors 2011:735279
Tevatia R, Prasad A, Saraf RF (2019) Electrochemical characteristics of a DNA modified electrode as a function of percent binding. Anal Chem 91:10501–10508
Herne TM, Tarlov MJ (1997) Characterization of DNA probes immobilized on gold surfaces. J Am Chem Soc 119:8916–8920
Ulman A (1996) Formation and structure of self-assembled monolayers. Chem Rev 96:1533–1554
Kummer K, Vyalikh DV, Gavrila G, Kade A, Weigel-Jech M, Mertig M, Molodtsov SL (2008) High-resolution photoelectron spectroscopy of self-assembled mercaptohexanol monolayers on gold surfaces. J Electron Spectros Relat Phenom 163:59–64
Kékedy-Nagy L, Shipovskov S, Ferapontova EE (2019) Electrocatalysis of ferricyanide reduction mediated by electron transfer through the DNA duplex: kinetic analysis by thin layer voltammetry. Electrochim Acta 318:703–710
Dijksma M, Boukamp BA, Kamp B, van Bennekom WP (2002) Effect of hexacyanoferrate(II/III) on self-assembled monolayers of thioctic acid and 11-mercaptoundecanoic acid on gold. Langmuir 18:3105–3112
Jarczewska M, Kierzkowska E, Ziółowski R, Górski Ł, Malinowska E (2015) Electrochemical oligonucleotide-based biosensor for the determinationof lead ion. Bioelectrochemistry 101:35–41
Acknowledgements
The authors gratefully acknowledge financial support from the DFG (GRK 2027) and NSERC (RGPIN-2016-06122).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Dedicated to Prof. Wolfgang Kaim on the occasion of his 70th birthday.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Escher, D., Hossain, M.N., Kraatz, HB. et al. Metal-dependent electrochemical discrimination of DNA quadruplex sequences. J Biol Inorg Chem 26, 659–666 (2021). https://doi.org/10.1007/s00775-021-01881-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-021-01881-9