Abstract
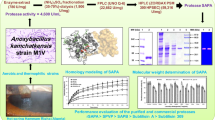
Extracellular protease SptA from the halophilic archaeon Natrinema sp. J7 was successfully expressed in Bacillus subtilis SCK6 using touchdown two-step PCR and plasmid multimer. Culture optimization was conducted to increase SptA protease activity 132-fold. It was shown that enzyme activity achieved the maximum activity at pH 8.0 and 55°C with casein as the substrate. SptA protease kept high stability and activity against many organic solvents and even maintained 60% activity in the presence of 8% NaCl. Meanwhile, SptA was redefined as a highly Ca2+-dependent subtilisin-like serine protease. Interestingly, SptA protease processed an autodigestion from 45 to 35 kDa in presence of Ca2+. Homology modeling illustrated the three binding sites of Ca2+ and the catalytic triad His231-Asp190-Ser384. In addition, SptA protease exhibited high compatibility with many surfactants and commercial detergents, showing 100% stability in the presence of commercial detergent Comfort. Washing performance analysis demonstrated remarkably high de-staining of the enzyme at 40°C for 20 min with a lower supplementation (500 U/mL). Accordingly, SptA protease could be considered as a promising bio-additive in the detergent industry.






Similar content being viewed by others
REFERENCES
Razzaq, A., Shamsi, S., Ali, A., Ali, Q., Sajjad, M., Malik, A., and Ashraf, M., Front. Bioeng. Biotechnol., 2019, vol. 7, p. 110.
Jaouadi, N., Jaouadi, B., Ben Hlima, H., Rekik, H., Belhoul, M., Hmidi, M., et al., PLoS One, 2014, vol. 9, e108367.
Abdelhedi, O., Jridi, M., Jemil, I., Mora, L., Toldra, F., Aristoy, M., et al., Food Res. Int., 2016, vol. 86, pp. 9–23.
Arbige, M., Shetty, J., and Chotani, G., Trends Biotechnol., 2019, vol. 37, vol. 12, pp. 1355–1366.
Rekik, H., Jaouadi, N., Gargouri, F., Bejar, W., Frikha, F., Jma,l N., et al., Int. J. Biol. Macromol., 2019, vol. 121, pp. 1227–1239.
Yang, H., Li, J., Shin, H., Du, G., Liu, L., and Chen, J., Appl. Microbiol. Biotechnol., 2014, vol. 98, no. 1, pp. 23–29.
Hadjidj, R., Badis, A., Mechri, S., Eddouaouda, K., Khelouia, L., Annane, R., et al., Int. J. Biol. Macromol., 2018, vol. 114, pp. 1033–1048.
Baweja, M., Nain, L., Kawarabayasi, Y., and Shukla, P., Front Microbiol., 2016, vol. 7, p. 965.
Wu, S. and Wong, S., J. Biotechnol., 1999, vol. 72, vol. 3, pp. 185–195.
You, C., Zhang, X., and Zhang, Y., Appl. Environ. Microbiol., 2012, vol. 78, no. 5, pp. 1593–1595.
Zhang, X.Z. and Zhang, Y.H.P., Microb. Biotechnol., 2011, vol. 4, no. 1, pp. 98–105.
Bradford, M.M., Anal. Biochem., 1976, vol. 72, nos. 1–2, pp. 248–254.
Natalia, Y., Hashim, R., Ali, A., and Chong, A., Aquaculture, 2004, vol. 233, nos. 1–4, pp. 305–320.
Tran, L.H., and Nagano, H., J. Food Sci., 2002, vol. 67, no. 3, pp. 1184–1187.
Smith, C.A., Toogood, H.S., Baker, H.M., Daniel, R.M., and Baker, E.N., J. Mol. Biol., 1999, vol. 294, no. 4, pp. 1027–1040.
Shi, W.L., Tang, X.F., Huang, Y.P., Gan, F., Tang, B., and Shen, P., Extremophiles, 2006, vol. 10, no. 6, pp. 599–606.
Gradišar, H., Friedrich, J., Križaj, I., and Jerala, R., Appl. Environ. Microbiol., 2005, vol. 71, no. 7, pp. 3420–3426.
Sun, F.D., Sun, Q.X., Zhang, H., Kong, B.H., and Xia, X.F., Int. J. Biol. Macromol., 2019, vol. 133, pp. 987–997.
Liu, S.Y., Selle, P.H., Court, S.G., and Cowieson, A.J., Anim. Feed Sci. Technol., 2013, vol. 183, nos. 3–4, pp. 175–183.
Baweja, M., Tiwari, R., Singh, P.K., Nain, L., and Shukla, P., Front Microbiol., 2016, vol. 7, p. 1195.
Lee, S. and Jang, D.J., Biophys. J., 2001, vol. 81, no. 5, pp. 2972–2978.
Xu, J.X., Zhuang, Y., Wu, B., Su, L., and He, B.F., J. Biol. Inorg. Chem., 2013, vol. 18, no. 2, pp. 211–221.
Gohel, S.D. and Singh, S.P., Int. J. Biol. Macromol., 2018, vol. 113, pp. 565–574.
Krishnamurthy, A. and Belur, P.D., Int. J. Biol. Macromol., 2018, vol. 112, pp. 110–118.
Yang, S., Zhai, L., Huang, L., Meng, D., Li, J., Hao, Z., et al., Int. J. Biol. Macromol., 2020, vol. 153, pp. 1220–1230.
Sareen, R. and Mishra, P., Appl. Microbiol. Biotechnol., 2008, vol. 79, no. 3, pp. 399–405.
Liew, K.J., Ngooi, C.Y., Shamsir, M.S., Sani, R.K., Chong, C.S., and Goh, K.M., Protein Expression Purif., 2019, vol. 164, p. 105464.
DasSarma, S. and DasSarma, P., Curr. Opin. Microbiol., 2015, vol. 25, pp. 120–126.
Gradišar, H., Friedrich, J., Križaj, I., and Jerala, R., Appl. Environ. Microbiol., 2005, vol. 71, no. 7, pp. 3420–3426.
Saravanan, P., Renitha, T., Gowthaman, M., and Kamini. N., J. Cleaner Prod., 2014, vol. 79, pp. 258–264.
George, N., Chauhan, P., Kumar, V., Puri, N., and Gupta, N., J. Cleaner Prod., 2014, vol. 79, pp. 249–257.
Jain, D., Pancha, I., Mishra, S., and Shrivastav, A., Bioresour. Technol., 2012, vol. 115, pp. 228–236.
Kumar, V., Marin-Navarro, J., and Shukla. P., World J. Microbiol. Biotechnol., 2016, vol. 32, no. 2, pp. 1–10.
ACKNOWLEDGMENTS
This work was supported by the National Key Research and Development Program of China (2017YFB0308401).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declared that there are no competing interests. This article does not contain any studies involving animals or human participants performed by any of the authors.
Rights and permissions
About this article
Cite this article
Wang, H., Cai, C., Gan, L. et al. Expression and Characterization of Surfactnt-Stable Calcium-Dependent Protease: a Potential Additive for Laundry Detergents. Appl Biochem Microbiol 57, 481–492 (2021). https://doi.org/10.1134/S0003683821040165
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0003683821040165