Abstract
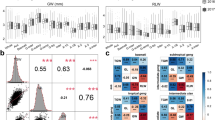
Grain morphology is an important agronomic trait since its strongly associated with thousand kernel weight, which is one of the main objectives for high–yield wheat breeding. The grain morphology–related traits were controlled by QTLs which were pyramided in wheat cultivars. To detect more genetic variants for grain morphology–related traits in improved varieties, a panel of 543 wheat accessions were phenotyped at three locations over 2 years with seven grain morphology–related traits: grain length (GL), grain width (GW), ratio of GL to GW, grain diameter, grain roundness, grain circumference, and grain surface area. Combined with 90 K Illumina iSelect single nucleotide polymorphism (SNP) array, a total of 47 significance SNPs on chromosomes 1A, 1B, 2A, 2B, 2D, 3A, 3B, 3D, 4B, 5B, 6A, 6B, 6D, and 7B were detected. An analysis of all loci identified 400 candidate genes, 22 of which included at least one significant SNP. The functional characterization of these candidate genes provided new insights into the genetic basis of wheat grain morphology.



Similar content being viewed by others
References
Bates D, Mächler M, Bolker B, Walker S (2014) Fitting linear mixed–effects models using lme4. J Stat Softw arXiv:1406(1):133–199. https://doi.org/10.18637/jss.v067.i01
Botwright TL, Condon AG, Rebetzke GJ, Richards RA (2002) Field evaluation of early vigour for genetic improvement of grain yield in wheat. Aust J Agric Res 53:1137–1145. https://doi.org/10.1071/AR02007
Boussau B, Szöllősi GJ, Duret L, Gouy M, Tannier E, Daubin V (2013) Genome–scale coestimation of species and gene trees. Genome Res 23:323–330. https://doi.org/10.1101/gr.141978.112
Brachi B, Morris GP, Borevitz JO (2011) Genome–wide association studies in plants: the missing heritability is in the field. Genome Biol 12:1–8. https://doi.org/10.1186/gb-2011-12-10-232
Breseghello F, Sorrells ME (2007) QTL analysis of kernel size and shape in two hexaploid wheat mapping populations. Field crops res 101(2):172–179. https://doi.org/10.1016/j.fcr.2006.11.008
Campbell KG, Bergman CJ, Gualberto DG, Anderson JA, Giroux MJ, Hareland G, Fulcher RG, Sorrells ME, Finney PL (1999) Quantitative trait loci associated with kernel traits in a Soft × Hard wheat cross. Crop Sci 39:1275–1285. https://doi.org/10.2135/cropsci1999.0011183X003900040039x
Cavanagh CR, Chao S, Wang S, Huang BE, Stephen S, Kiani S, Forrest K, Saintenac C, Brown-Guedira GL, Akhunova A, See D, Bai G, Pumphrey M, Tomar L, Wong D, Kong S, Reynolds M, da Silva ML, Bockelman H, Talbert L, Anderson JA, Dreisigacker S, Baenziger S, Carter A, Korzun V, Morrell PL, Dubcovsky J, Morell MK, Sorrells ME, Hayden MJ, Akhunov E (2013) Genome–wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc Natl Acad Sci India A 110:8057–8062. https://doi.org/10.1073/pnas.1217133110
Chang JZ, Zhang JN, Mao XG, Li A, Jing RL (2013) Polymorphism of TaSAP1–A1 and its association with agronomic traits in wheat. Planta 237:1495–1508. https://doi.org/10.1007/s00425-013-1860-x
Chastian TG, Ward KJ, Wysocki DJ (1995) Stand estblishment response of soft white winter wheat to seedbed residue and seed size. Crop Sci 35:213–218. https://doi.org/10.2135/cropsci1995.0011183X003500010040x
Davidson RM, Reeves PA, Manosalva PM, Leach JE (2009) Germins: a diverse protein family important for crop improvement. Plant Sci 177:499–510. https://doi.org/10.1016/j.plantsci.2009.08.012
Dholakia BB, Ammiraju JSS, Singh H, Lagu MD, Röder MS, Rao VS, Dhaliwal HS, Ranjekar PK, Gupta VS, Weber WE (2003) Molecular marker analysis of kernel size and shape in bread wheat. Plant Breed 122:392–395. https://doi.org/10.1046/j.1439-0523.2003.00896.x
Gegas VC, Nazari A, Griffiths S, Simmonds J, Fish L, Orford S, Sayers L, Doonan JH, Snape JW (2010) A genetic framework for grain size and shape variation in wheat. Plant Cell 22:1046–1056. https://doi.org/10.1105/tpc.110.074153
Giura A, Saulescu NN (1996) Chromosomal location of genes controlling grain size in a large grained selection of wheat (Triticum aestivum L.). Euphytica 89:77–80. https://doi.org/10.1007/BF00015722
Guo Y, Sun JJ, Zhang GZ, Wang YY, Kong F, Zhao Y, Li SS (2013) Haplotype, molecular marker and phenotype effects associated with mineral nutrient and grain size traits of TaGS1a in wheat. Field Crops Res 154:119–125. https://doi.org/10.1016/j.fcr.2013.07.012
Huang XZ, Qian Q, Liu ZB, Sun HY, He SY, Luo D, Xia GM (2009) Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet 41:494–497. https://doi.org/10.1038/ng.352
Huang X, Wei X, Sang T, Zhao Q, Feng Q, Zhao Y, Li C, Zhu C, Lu T, Zhang Z, Li M, Fan D, Guo Y, Wang A, Wang L, Deng L, Li W, Lu Y, Weng Q, Liu K, Huang T, Zhou T, Jing Y, Li W, Lin Z, Buckler ES, Qian Q, Zhang QF, Li J, Han B (2010) Genome–wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42:961–967. https://doi.org/10.1038/ng.695
Ihmels J, Bergmann S, Barkai N (2004) Defining transcription modules using large–scale gene expression data. Bioinformatics 20:1993–2003. https://doi.org/10.1093/bioinformatics/bth166
Ishimaru K, Hirotsu N, Madoka Y, Murakami N, Hara N, Onodera H, Kashiwagi T, Ujiie K, Shimizu B-I, Onishi A (2013) Loss of function of the IAA–glucose hydrolase gene TGW6 enhances rice grain weight and increases yield. Nat Genet 45:707–711. https://doi.org/10.1038/ng.2612
Jiang QY, Hou J, Hao CY, Wang LF, Ge HM, Dong YS, Zhang XY (2011) The wheat (T. aestivum) sucrose synthase 2 gene (TaSus2) active in endosperm development is associated with yield traits. Funct Integr Genomics 11:49–61. https://doi.org/10.1007/s10142-010-0188-x
Li F, Chen BY, Xu K, Wu JF, Song WL, Bancroft I, Harper AL, Trick M, Liu SY, Gao GZ, Wang N, Yan GX, Qiao JW, Li J, Li H, Xiao X, Zhang TY, Wu XM (2014) Genome–wide association study dissects the genetic architecture of seed weight and seed quality in rapeseed (Brassica napus L.). DNA Res 21(4):355–367. https://doi.org/10.1093/dnares/dsu002
Li FJ, Wen W, He ZH, Liu JD, Jin H, Cao SH, Geng HW, Yan J, Zhang PZ, Wan YX, Xia XC (2018) Genome-wide linkage mapping of yield-related traits in three Chinese bread wheat populations using high-density SNP markers. Theor Appl Genet 131:1–22. https://doi.org/10.1007/s00122-018-3122-6
Li H, Peng ZY, Yang XH, Wang WD, Fu JJ, Wang JH, Han YJ, Chai YC, Guo TT, Yang N, Liu J, Warburton ML, Cheng YB, Hao XM, Zhang P, Zhao JY, Liu YJ, Wang GY, Li JS, Yan JB (2013) Genome–wide association study dissects the genetic architecture of oil biosynthesis in maize kernels. Nat Genet 45:43–50. https://doi.org/10.1038/ng.2484
Li N, Li Y (2014) Ubiquitin–mediated control of seed size in plants. Front Plant Sci 5:332. https://doi.org/10.3389/fpls.2014.00332
Li YB, Fan CC, Xing YZ, Jiang YH, Luo LJ, Sun L, Shao D (2011) Natural variation in GS5 plays an important role in regulating grain size and yield in rice. Nat Genet 43:1266–1269. https://doi.org/10.1038/ng.977
Lipka AE, Tian F, Wang QS, Peiffer J, Li M, Bradbury PJ, Gore MA, Buckler ES, Zhang ZW (2012) GAPIT: genome association and prediction integrated tool. Bioinformatics 28:2397–2399. https://doi.org/10.1093/bioinformatics/bts444
Ma DY, Yan J, He ZH, Wu L, Xia XC (2012) Characterization of a cell wall invertase gene TaCwi–A1 on common wheat chromosome 2A and development of functional markers. Mol Breed 29:43–52. https://doi.org/10.1007/s11032-010-9524-z
Mao HL, Sun SY, Yao JL, Wang CR, Yu SB, Xu CG, Li XH (2010) Linking differential domain functions of the GS3 protein to natural variation of grain size in rice. Proc Natl Acad Sci India A 107:19579–19584. https://doi.org/10.1073/pnas.1014419107
Mir RR, Kumar N, Jaiswal V, Girdharwal N, Prasad M, Balyan HS, Gupta PK (2012) Genetic dissection of grain weight in bread wheat through quantitative trait locus interval and association mapping. Mol Breed 29:963–972. https://doi.org/10.1007/s00122-019-03418-w
Petersen G, Seberg O, Yde M, Berthelsen K (2006) Phylogenetic relationships of Triticum and Aegilops and evidence for the origin of the A, B, and D genomes of common wheat (Triticum aestivum). Mol Phylogenet Evol 39:70–82. https://doi.org/10.1016/j.ympev.2006.01.023
Qi P, Lin YS, Song XJ, Shen JB, Huang W, Shan JX, Zhu MZ, Jiang LW, Gao JP, Lin HX (2012) The novel quantitative trait locus GL3. 1 controls rice grain size and yield by regulating Cyclin–T1; 3. Cell Res 22(12):1666–1680. https://doi.org/10.1038/cr.2012.151
Ramya P, Chaubal A, Kulkarni K, Gupta L, Gupta V (2010) QTL mapping of 1000–kernel weight, kernel length, and kernel width in bread wheat (Triticum aestivum L.). J Appl Genet 51:421–429. https://doi.org/10.1007/BF03208872
Shomura A, Izawa T, Ebana K, Ebitani T, Kanegae H, Konishi S, Yano M (2008) Deletion in a gene associated with grain size increased yields during rice domestication. Nat Genet 40:1023–1028. https://doi.org/10.1038/ng.169
Simmonds J, Scott P, Brinton J, Mestre TC, Bush M, Del Blanco A, Dubcovsky J, Uauy C (2016) A splice acceptor site mutation in TaGW2–A1 increases thousand grain weight in tetraploid and hexaploid wheat through wider and longer grains. Theor Appl Genet 129:1099–1112. https://doi.org/10.1007/s00122-016-2686-2
Song XJ, Huang W, Shi M, Zhu MZ, Lin HX (2007) A QTL for rice grain width and weight encodes a previously unknown RING–type E3 ubiquitin ligase. Nat Genet 39:623–630. https://doi.org/10.1038/ng2014
Sun C, Zhang F, Yan X, Zhang X, Dong Z, Cui D, Chen F (2017) Genome–wide association study for 13 agronomic traits reveals distribution of superior alleles in bread wheat from the Yellow and Huai Valley of China. Plant Biotechnol J 15:953–969. https://doi.org/10.1111/pbi.12690
Su ZQ, Hao CY, Wang LF, Dong YC, Zhang XY (2011) Identification and development of a functional marker of TaGW2 associated with grain weight in bread wheat (Triticum aestivum L.). Theor Appl Genet 122:211–223. https://doi.org/10.1007/s00122-010-1437-z
Sun LJ, Li XJ, Fu YC, Zhu ZF, Tan LB, Liu FX, Sun XY (2013) GS6, a member of the GRAS gene family, negatively regulates grain size in rice. J Integr Plant Biol 55:938–949. https://doi.org/10.1111/jipb.12062
Sun XY, Wu K, Zhao Y, Kong FM, Han GZ, Jiang HM, Huang XJ, Li RJ, Wang HG, Li SS (2009) QTL analysis of kernel shape and weight using recombinant inbred lines in wheat. Euphytica 165:615. https://doi.org/10.1007/s10681-008-9794-2
Wang SK, Wu K, Yuan QB, Liu XY, Liu ZB, Lin XY, Zeng RZ (2012) Control of grain size, shape and quality by OsSPL16 in rice. Nat Genet 44:950–954. https://doi.org/10.1038/ng.2327
Williams K, Sorrells ME (2014) Three‐dimensional seed size and shape QTL in hexaploid wheat (Triticum aestivum L.) populations. Crop Sci 54(1):98–110. https://doi.org/10.2135/cropsci2012.10.0609
Wu QH, Chen YX, Zhou SH, Fu L, Chen JJ, Xiao Y et al (2015) High-density genetic linkage map construction and QTL mapping of grain shape and size in the wheat population yansda 1817 × Beinong6. PLoS ONE 10(2):e0118144. https://doi.org/10.1371/journal.pone.0118144
Xu F, Park MR, Kitazumi A, Herath V, Mohanty B, Yun S, De lR, Benildo G (2012) Cis–regulatory signatures of orthologous stress–associated bZIP transcription factors from rice, sorghum and Arabidopsis based on phylogenetic footprints. BMC Genomics 13:497. https://doi.org/10.1186/1471-2164-13-497
Yu JM, Pressoir G, Briggs WH, Irie VB, Yamasaki M, Doebley JF, McMullen MD, Gaut BS, Nielsen DM, Holland JB, Kresovich S, Buckler ES (2006) A unified mixed–model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208. https://doi.org/10.1038/ng1702
Zhang L, Zhao YL, Gao LF, Zhao GY, Zhou RH, Zhang BS, Jia JZ (2012) TaCKX6-D1, the ortholog of rice OsCKX2, is associated with grain weight in hexaploid wheat. New Phytol 195:574–584. https://doi.org/10.1111/j.1469-8137.2012.04194.x
Zhang LY, Liu DC, Guo XL, Yang WL, Sun JZ, Wang DW, Zhang AM (2010) Genomic distribution of quantitative trait loci for yield and yield-related traits in common wheat. J Integr Plant Biol 52:996–1007. https://doi.org/10.1111/j.1744-7909.2010.00967.x
Zhao Y, Li JH, Zhao RL, Xu K, Xiao YR, Zhang SH, Tian JC, Yang XJ (2020) Genome–wide association study reveals the genetic basis of cold tolerance in wheat. Mol Breed 40:36. https://doi.org/10.1007/s11032-020-01115-x
Zuo JR, Li JY (2014) Molecular genetic dissection of quantitative trait loci regulating rice grain size. Annu Rev Genet 48:99–118. https://doi.org/10.1146/annurev-genet-120213-092138
Funding
This work was supported by the Science and Technology Planning Project of Hebei province (16226320D), and the National Key Research and Development Program of China (2017YFD0100600).
Author information
Authors and Affiliations
Contributions
Y.Z. planned and designed the research. L.G., S.J.S., K.X., H.D.L. and S.H.Z. performed the experiments. L.G. and J.Y. analysed the data. L.G. and J.Y. wrote the text. All authors read, commented on and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gao, L., Yang, J., Song, Sj. et al. Genome–wide association study of grain morphology in wheat. Euphytica 217, 170 (2021). https://doi.org/10.1007/s10681-021-02900-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-021-02900-1