Abstract
Because of the limitations inherent in conventional breeding of trees and clonally propagated crops, gene editing is of great interest. Dozens of published papers attest to the high efficiency of CRISPR-based systems in clonal crops and trees. The opportunity for “clean” edits is expected to avoid or reduce regulatory burdens in many countries and may improve market acceptance. To date, however, nearly all studies in trees and clonal crops retained all of the gene editing machinery in the genome. Despite high gene editing efficiency, technical and regulatory obstacles are likely to greatly limit progress toward commercial use. Technical obstacles include difficult and slow transformation and regeneration, delayed onset of flowering or clonal systems that make sexual segregation of CRISPR-associated genes difficult, inefficient excision systems to enable removal of functional (protein- or RNA-encoding) transgenic DNA, and narrow host range or limited gene-payload viral systems for efficient transient editing. Regulatory obstacles include those such as in the EU where gene-edited plants are regulated like GMO crops, and the many forms of method-based systems that regulate stringently based on the method vs. product novelty and thus are largely applied to each insertion event. Other major obstacles include the provisions of the Cartagena Protocol with respect to international trade and the need for compliance with the National Environmental Policy Act in the USA. The USDA SECURE act has taken a major step toward a more science- and risk-based—vs. method and insertion event based—system, but much further regulatory and legal innovation is needed in the USA and beyond.
Similar content being viewed by others
Introduction
Trees and clonally propagated plants represent a large portion of cultivated plant species. They are used for food, fiber, and fuel, and include some of the best-known crops in the world, including banana, apple, citrus, grape, stone fruits, nut trees, sugar cane, potato, mint, and numerous species of forest trees used for wood production. Many of these crops have ancient associations with agriculture, and some clones have had hundreds to thousands of years of continual propagation and cultivation, as is evidenced by recent genomic characterizations (Duan et al. 2017; Wu et al. 2018; Vondras et al. 2019). For clonally propagated plants, perpetual retainment of genotype-unique traits and heterosis is an important trait common to many cultivars. Coupled with delayed onset of reproduction, low tolerance of inbreeding, and natural sterility, many of these species are slow and difficult to breed. In addition, oligogenic and recessive traits of high value, such as those affecting product quality and pest resistance, are nearly impossible to breed for when they are rare in gene pools. Thus, by enabling the efficient production of loss of function traits, gene editing provides a powerful new tool that was essentially unavailable in tree and clonal crops.
The same traits that make sexual breeding difficult in tree and clonal crops, however, make gene editing and genetic engineering challenging. It is difficult to introgress new edits or transgenes into diverse lines and also difficult to remove editing machinery from the genome by sexual segregation, as is common in annual crops. In addition, many of these species are also notoriously difficult to genetically modify and regenerate (Birch 1997; Busov et al. 2005). Even in species where genetic transformation has been accomplished, such as poplars and eucalypts, most genotypes remain extremely difficult to transform (Nagle et al. 2018). Thus, new technologies that can speed genetic modification and gene editing, or reduce obstacles to their regulatory acceptance, are crucial.
There are several classes of technologies that can promote the production and acceptance of gene-edited trees and clonal crops. Technologies such as the use of developmental regulators (DRs) for example GROWTH REGULATORY FACTOR 4 (Debernardi et al. 2020), among many others, to aid regeneration of genetically modified cells have been reviewed elsewhere and will not be covered here (Nagle et al. 2018). However, there have been very few reports of their successful use in tree and clonal crops (Florez et al. 2015; Brand et al. 2019; Maher et al. 2020), thus appear to be important technologies for further development. The use of DRs is likely to require transient expression or excision from the genome (Hoerster et al. 2020), in common with the desire to avoid permanent insertion of gene editing machinery from genomes (to avoid off-target effects and ease regulatory or market approval). Thus, most of the technologies we discuss below can have benefits beyond facilitating the production and acceptance of gene-edited varieties. We first briefly review recent developments in CRISPR technology with special relevance to tree and clonal crops, then discuss options for transient editing, transgene excision, and accelerated flowering to enable segregation of gene editing components away where breeding methods allow it.
CRISPR/Cas Techniques: a Rapidly Evolving Toolset for Plant Genetic Modification
The previous decade has seen the rapid development of reliable and versatile molecular tools for the targeted modification of endogenous plant genomes, which has been a broad and rapid departure from mainly transgene-based tools such as overexpression, silencing, misexpression, and introduction of novel traits, which together dominated plant molecular biology throughout the 1990s and 2000s. While a number of effective site-directed nucleases such as TAL-like effector nucleases (TALENs) and designer zinc finger nucleases (ZFNs) have been successfully used in plants, in recent years, CRISPR/Cas9 systems have become most common (reviewed in Razzaq et al. 2019) Many recent variants of the original Cas9 system from Streptococcus pyogenes (spCas9) have expanded the toolbox of the types of modifications that can be accomplished, highlighting the versatility of Cas protein systems.
The main functional attributes of CRISPR systems are the capacity for mutations in target genes, directed single base changes, and directed sequence modifications (termed homology-directed replacement: HDR) (reviewed in Chen et al. 2019). Directed mutations lead to loss of function through frame-shift mutations or large deletions, which can be done in multiple genes simultaneously at high efficiency. There are many examples of such applications in trees and other clonal crops (summarized in Table 1). By fusion of catalytically dead Cas9 to natural or synthetic cytosine or adenine deaminases, targeted changes to individual bases can be accomplished without the use of a donor template (reviewed in Bharat et al.. 2020). Traits such as herbicide resistance traits have been modified using this method (Li et al. 2018) and were recently demonstrated in sugarcane (a clonal crop) (Oz et al. 2017). HDR was also recently shown in cassava to tag an endogenous gene with GFP to monitor expression levels (Veley et al. 2021); however, to our knowledge, no trait-focused non-herbicide-resistant traits have been shown in a tree or clonal crop using HDR. HDR in comparison is far less efficient than direct mutations or base editing, and as discussed below raises larger regulatory concerns as they are generally treated as regular GMOs.
Because rates of HDR are low compared with directed mutation, several methods have been adapted to improve efficiencies. These include the use of viral replicons that increase donor template copy number (Čermák et al. 2015). A second means to improve rates of directed sequence replacement is to increase the temperature during the early stages of transformation, as the efficiency of many Cas proteins seems to be tailored toward warmer temperatures than are commonly found in plant in vitro culture (22–25°C) (Malzahn et al. 2019; Milner et al. 2020). Still, many reports from stable integration of Cas9 and gRNAs in several species report 10% or lower rates of HDR incorporation of a donor template. Cas9 modified fusions to proteins such as virD2 may further increase these rates in addition to other potential strategies (Begemann et al., 2017; Čermák et al., 2017; Ali et al., 2020; Dong et al., 2020). Another technological improvement useful for directed sequence replacement is “prime” editing (Anzalone et al. 2019). It does not require a DNA template nor homology-driven recombination and thus should be more efficient than HDR. So far, this has been demonstrated in rice, wheat, and potato but at fairly low frequencies of 10–20% (Lin et al. 2020; Veillet et al. 2020b). Though inefficient at present, these approaches greatly increase the types of genomic modifications available, such as insertion of promoter elements to change endogenous expression and directing transgenes into “safe harbors” (i.e., well-characterized zones of DNA unlikely to be affected by transgene silencing or affecting the function of native genes).
Means to Produce Edited Trees/Clonal Crops Without Permanent Cas9/gRNA Integration
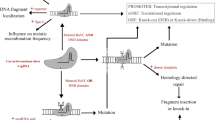
The dominant method for the development of CRISPR/Cas9-mediated edits in the genomes of trees and clonally propagated crops is through stable Agrobacterium-mediated transformation (see Table 1 for recent trends). Generally, this would involve delivery of Cas9, gRNAs, and plant resistance marker genes in a single T-DNA. While it is routine in many herbaceous species to self-pollinate such transgenic plants and then screen progeny for a combination of desired edits and absence of the CRISPR-related genes, it is very difficult to do in trees and clonally propagated crops (Nadakuduti et al. 2018). This is often due to the time delay to onset of flowering (especially in trees), intolerance of inbreeding, sterility of hybrid or polyploid varieties, and loss of clonal integrity due to sexual segregation. If the Cas9 and gRNA genes must be removed from edited lines due to regulatory needs or biological concerns (discussed below), methods for editing without integrated transgenes, or technologies for removing integrated transgenes, will be required. In the following sections, we outline the general methods that are available and the subject of active research (class I–IV, Fig. 1). We do not consider the case for retention of gene editing machinery in commercial plants, although evidence for extremely low rates of off-target mutation in plants (Young et al. 2019), reviewed in Graham et al. (2020) and Modrzejewski et al. (2020), and recent evidence for a virtual absence of vegetative consequences of continued CRISPR/Cas9 expression in eucalypts (Elorriaga et al. 2021), provides support for this option. If CRISPR machinery is retained, there may be regulatory or market concerns over potential gene drives affecting domesticated or wild/feral populations. However, because of the long generation time or low fertility/sterility of many tree and clonal crops, we expect that at least the biological concerns over possible gene drive to be negligible.
Methods for gene editing product development in trees and clonally propagated crops and anticipated regulatory outcomes in the USA under USDA SECURE. Gene editing via CRISPR/Cas9 can be accomplished through several approaches in trees and clonally propagated crops. These involve stable integration of an editing transgene in the genome (top arrow), viral delivery methods (left arrow), and transient delivery systems (right arrow). Under SECURE, methods detailed in the green region are likely to be exempt from regulation so long as the trait meets eligibility criteria and only single and simple edits are made that mimic potential natural variants. Methods shown in the yellow highlighted are likely to require a Regulatory Status Review (RSR) similar to that for other transgenic traits. Stable integration methods are the most common at present; in clonally propagated crops, somatic transgene removal systems mediated by site-specific recombinases or transposable elements have been demonstrated, though to date only recombinases have been used in conjunction with gene editing. If sexual segregation is possible, accelerated flowering systems can be used to remove stably integrated editing transgenes. Transient delivery methods can be accomplished by various DNA delivery systems (Agrobacterium, plasmids by physical delivery) or DNA-free methods (Cas9/-RNPs). These methods must be efficient enough to produce edited non-chimeric shoots, or protoplast regeneration from single cells, at a rate adequate to address breeding program demands. Viral-only delivery methods could be used if the viral genome has enough capacity for a Cas enzyme (rhabdoviruses, Potato virus X) and the gRNAs. If viral systems are used only to deliver gRNAs during systemic infection, mixed systems with stable Cas9 lines would be required.
Class I. Stable Integration of Gene Editing Components Followed by Removal of Functional Components Via Gene Excision
The removal of parts of transgenes has been a key component of transgenic research in plants for several decades, where focus has been on removal of antibiotic or herbicide resistance markers for regulatory acceptance (reviewed in Tuteja et al. 2012). Another benefit of marker removal is so that elite-generated insertion events can be stacked with further transgenic traits using the same selectable marker. Generally, marker removal from integrated transgenes by recombination is not possible even in annuals with high fecundity because of the tight linkage between the desired trait and the selectable marker in the same transgene. This necessitated methods where parts of the transgene could be selectively excised after the initial transformation process, and these usually focused on the somatic removal of selectable marker genes using recombinases such as Cre from P1 bacteriophage and FLP from yeast (Kilby et al. 1995; Wang et al. 2005; Hu et al. 2008; Kim et al. 2012), although more exotic recombinases have been used such as R, ParA, and CinH (Schaart et al. 2004; Shao et al. 2017). In these strategies, recognition sites for the recombinases are included at flanking regions where parts of transgenes are desired to be deleted (the selectable marker and usually including the recombinase gene). These could be expressed from inducible promoters (e.g., heat shock or cold shock) or developmental promoters (e.g., meristematic or reproductive; Fig. 2(B)) to decrease the rates of premature marker removal during the initial phases of transformation, but allow for the excision to occur once integrated transgenic events are isolated or potentially characterized (Fig. 2). These methods have been adapted to the removal of CRISPR-related genes in two recent studies in apple and grape where the FLP recombinase was expressed from a heat shock promoter and flanking recognition sites were placed near the T-DNA borders (Pompili et al. 2020). After transgenic lines were isolated and identified in vitro for traits predicted to confer blight resistance, the CRISPR-related genes were excised using heat shock. The method was effective for removing the editing transgene but required recovery and further regeneration in vitro of the material due to the prolonged, high temperature required for adequate excision. A second study in apple and grape using the same approach and a novel second method using CRISPR/Cas9-mediated deletion of the transgene showed rapid but incomplete excision after 3 h of heat shock with FLP recombinase and significantly less excision using Cas9 (and additionally, issues with CRISPR target sites being irreversibly mutated to prevent excision) (Dalla Costa et al. 2020). When taking into account marker removal studies, recombinase excision approaches have been validated in several tree and clonally propagated species, including poplars, strawberry, potato, and apple, highlighting their versatility (Schaart et al. 2004; Kondrák et al. 2006; Fladung et al. 2010; Fladung and Becker 2010; Timerbaev et al. 2019). However, improvement of the typically low excision rates seen may require mitigation of target site methylation (Liu et al. 2021).
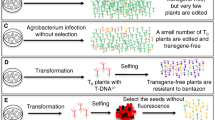
Method for coupling stable transgene integration followed by controlled recombinase-mediated excision to generate edited clonal plants with minimal remaining inserted DNA. (A) Transgenes delivered by Agrobacterium containing all the elements required for gene editing and transgene removal are stably integrated into the genome. When recombinases are expressed, they bind recognition sequences flanking the T-DNA region and excise the transgene through circularization and subsequent degradation. (B) Though many schemes for recombinase control after gene editing are possible (heat shock, reproductive development-triggered expression, chemical induction), developmental control of recombinase expression in shoot meristems would allow for the production of edited shoots with eliminated transgenes in a single step when used with common organogenic in vitro transformation systems. Shoots without fluorescent marker genes could be retained and analyzed for desired edits. For species and genotypes with low transformation rates, stable integration and recombinase-mediated excision methods are likely to be beneficial over transient or DNA-free methods, due to selection and positive evidence of transgenic insertion. The main challenge is the identification of developmental promoters and associated insulator elements that give desired and reliable expression levels in transgenic target tissues; a combination of developmental and environmentally or chemically induced expression may be needed for such a system to work reliably.
A drawback of this approach is that a small residual “footprint” from the original T-DNA insertion will still be present in the genome. If the original insertion event landed within the coding sequence of an existing gene, the function of that gene will likely still be affected after the transgene is excised. If a particular insertion event has multiple T-DNA integrations, the complete excision of all transgenes may be difficult to accomplish, and their resolution may give rise to complex rearrangements or deletions of large sections of native DNA (when the T-DNAs and associated recognition sites are located in close proximity on the same chromosome) (Wang et al. 2011). Thus, as discussed in depth below, under extant regulatory regimes, the plants, though lacking any transgenically derived functional (protein or RNA encoding) transgenes, may face similar regulatory hurdles to conventional GMOs (Fig. 1)—though of course chromosomal rearrangements are a common feature of conventional breeding that employs wide crosses, somaclonal variation, or radiation or chemical mutagenesis (Bradford et al. 2005; Fossi et al. 2019; Jo and Kim 2019).
Transposable elements can serve a similar function to recombinases. In this strategy, the transgene could be delivered with a functional transposase gene and recognition sites at the borders of the transgene to direct the transposase to excise. Over time, the integrated transgene might be excised completely and not be integrated elsewhere in the genome. Because the maize Ac transposon is very well characterized and functional in monocots and many dicot plants, several studies have used the Ac transposase and demonstrated its function in tobacco and poplar for the removal of transgenes (Ebinuma et al. 1997). Recent developments in animal transformation systems have developed excision only transposase enzymes such as piggyBac, which could be adapted to plant excision systems, but have only been demonstrated in a single recent study in rice (Nishizawa-Yokoi and Toki 2021). In demonstrated examples in plants, this strategy has a low frequency of excision and takes several months or longer to obtain fully non-chimeric, excised events. These drawbacks explain the increased focus on recombinase excision systems both for marker removal and editing transgene removal in trees and clonal crops.
Class II. Transient DNA Delivery (No Integration)
Stable integration of CRISPR-related genes in the plant genome has been repeatedly shown to give rise to non-chimeric, edited events at high frequency (Table 1). However, as discussed above, the complete removal of gene editing components without sexual reproduction is problematic in trees and many clonal crops. One obvious solution is to avoid integrating the transgene in the first place—relying on transient expression of editing agents. There are several methods that have been demonstrated or proposed to accomplish this, and they all share similar drawbacks when compared to editing by stable transgene integration. These include the difficulties of obtaining non-chimeric plants without strong selection using antibiotic or herbicide resistance genes, and overall low rates of editing in regenerated plants (Chen et al. 2018). The latter is a serious obstacle for species with low transformation rates, as normally tens to many hundreds of treated plants must be screened to find a small number of homozygous knockout edited plants (Bánfalvi et al. 2020). However, because the gene-edited plants generated using these approaches will be exempt in many countries (discussed below), there has been a flurry of interest in their development.
The first such technique is to borrow what is commonplace in animal CRISPR/Cas9 editing systems—i.e., simply to provide the Cas enzyme and associated guide RNAs as a pre-packaged ribonucleic protein complex (RNP) without any DNA (unless a donor template is required for HDR) (Chen et al. 2016). Using this approach requires a physical delivery method for the Cas9/sgRNA RNPs, similar to how fertilized zebrafish or mouse embryos are often injected (Hoshijima et al. 2019). In plants, the most common method employed has been to use polyethylene glycol (PEG) physical transformation with cultured protoplasts (which have the cell wall largely removed), then to regenerate the protoplasts into intact plantlets in vitro (Murovec et al. 2018; Wu et al. 2020). Generally, the use of DNA-free delivery and of single-celled source material greatly reduces the problem of chimeric regenerants, provides a high rate of editing, and eliminates the possibility of transgene integration (Metje-Sprink et al. 2019). The main drawbacks are that protoplast regeneration systems are very difficult to develop and reproduce in most plant species, especially trees, and protoplast regeneration methods are well known to cause a high rate of somaclonal mutation (Roest and Gilissen 1989; Bairu et al. 2011; Fossi et al. 2019). For example, when comparing protoplast and explant-based regeneration in potato, the former was found to cause a much higher rate of structural somatic mutations (Fossi et al. 2019). Another drawback is the frequent integration of small fragments of DNA at CRISPR target sites. In rice, a significant proportion of Cas9 RNP–derived lines contained DNA integrated at target cut sites whose origin was most likely explained by contamination from various sources of plasmid DNA (Banakar et al. 2019). More research in other species, and with contrasting transformation approaches, is needed to understand how widespread this problem is. A few examples of Cas9-RNP-mediated gene editing have been demonstrated in clonal crops and are discussed in greater detail below and found in Table 1.
A second method by which edited but transgene-free events can be obtained is using standard Agrobacterium transformation techniques but applying little or no antibiotic selection. This approach was demonstrated in tobacco (Chen et al. 2018); however, very low rates of successful edited and transgene-free events were obtained (0.4%). Potentially, modifications to Agrobacterium T-DNA function which increase infection and transient T-DNA delivery, but hinder the rate of transgene integration, may improve this approach. For example, modifications in proteins essential for T-DNA delivery such as VirD2, the protein which assists in processing and piloting the T-DNA via interaction with the T-DNA right border (RB) sequence, or plant factors involved in DNA repair may be important for reducing the rate of T-DNA integration (Mysore et al. 1998; van Kregten et al. 2009; van Kregten et al. 2016; Nishizawa-Yokoi et al. 2021).
Another option for improvement of transient editing is to have a short period of antibiotic selection when transient expression is highest. One study in potato saw no significant increase in edited, transgene-free plant recovery using this approach and was unable to recover non-chimeric edited shoots (Bánfalvi et al. 2020). Another study in apple succeeded in retaining transgene-free edited shoots using a visual editing marker, but at a 0.26% rate compared with all regenerated shoots (Charrier et al. 2019). At present, the rates of edited, transgene-free events may be too low to be useful in the many species of trees and clonally propagated crops with poor transformation rates.
Developmental genes that aid in regeneration of genetically modified cells could also improve the rate of transient editing. Regeneration aiding genes such as WUSCHEL (WUS) or BABY BOOM (BBM), whose proteins can move between cells, can have a negative or toxic impact on cells stably transformed but a positive effect on the regeneration of surrounding cells (Lowe et al. 2016; Hoerster et al. 2020). Such “altruistic” approaches typically employ a two-vector strategy where sacrificial cells retain the developmental gene encoding T-DNA, while neighboring cells are hopefully modified but do not see developmental gene integration (Hoerster et al. 2020). This approach is desirable because it may increase the rate of transformation in difficult to regenerate genotypes. This type of editing was recently shown for “in planta” transformation in tobacco, grape, tomato, and potato with Agrobacterium (Maher et al. 2020). In this study, a pair of developmental genes (IPT/WUS or BBM or SHOOT MERISTEMLESS (STM)) together with sgRNAs (all encoded by a single T-DNA) was applied to meristematic tissues of a stable Cas9-expressing plant under growth chamber or greenhouse conditions. Although only a few seeds and shoots were analyzed, most appeared to be edited but lacking stable T-DNA integration. Although the generation of a stable Cas9 expressing founder lines was an added complication of this approach, this might not be needed (i.e., Cas9 might be provided in the binary vector, as is common in gene editing studies). Additional studies to replicate this work, including in other plant species and in some of the notoriously difficult trees and clonal crops, are needed.
Class III. Viral Delivery Approaches
There has been a recent increase in the use of viral delivery systems for plant gene editing. In its most common guise, plant viruses are used to systemically delivery gRNAs throughout an infected plant (Hu et al. 2019; Ghoshal et al. 2020). These could be used to accomplish gene editing in specific tissues (where viruses replicate) and thus to obtain non-chimeric edited and virus-free plants from those tissues. The major drawback of using viruses for editing, whether they be RNA or DNA viruses, is their limited genome size and thus modest gene delivery payload, if they are to maintaining the ability to move systemically throughout the plant. Widely studied RNA viruses such as Tobacco Mosaic Virus (TMV) lose the ability for systemic infection if Cas9 (~4kb protein-coding portion) is included in the viral genome (Ellison et al. 2020). In general, this means that mixed approaches with previously transformed Cas9-expressing plants are required to generate non-chimeric edited plants (Figs. 1 and 3).
Viral methods for gene editing component delivery to generate gene-edited lines in trees and clonally propagated plants. (A) gRNAs are delivered to Cas9 transgenic plants through systemic infection of a RNA virus such as TMV that supports a small genome payload. Infection of transgenic plants already expressing Cas9 results in many edited cells in tissues bearing viral infection. Regeneration of infected leaf tissue into shoots purges the virus and frequently results in fixation for edits/non-edits due to small founder cell populations. (B) gRNAs are delivered by systemic infection but are also tailored with mobile RNA elements which help improve gRNA movement into reproductive tissues (Using parts of the FT or AG transcript) to areas which are normally inaccessible to viral infection. When coupled with a transgenic plant with accelerated flowering and expressing Cas9, edited seeds can be obtained, and the transgene containing Cas9 and the FT expression construct can be segregated out of the plant genome. (C) Negative-strand RNA rhabdoviruses with large genome sizes can be used in a manner similar to the method outlined in (A), however, do not require a Cas9 transgenic plant to infect, simplifying the generation of edited plants.
More advanced viral editing strategies which increase gRNA mobility in reproductive tissues may expand the versatility of these systems. In contrast to the previous strategies, some plant negative-strand RNA viruses (NSVs) such as rhabdoviruses may have the requisite payload capacity to deliver both Cas9 and the gRNAs via systemic infection. Recently, the Sonchus yellow net virus (SYNV) and the Potato virus X (PVX) were shown to be capable of delivering both Cas9 and gRNAs throughout tobacco plants (Ariga et al. 2020; Ellison et al. 2020). These studies also caused successful editing of the PHYTOENE DESATURASE (PDS) endogene, a GFP visual marker transgene, and a few other genes of interest. Using this system, after infection and editing, virus-free materials can be produced through somatic regeneration. Although the host range of SYNV and PVX is limited, if similar viruses can be found for suitably wide taxonomic groups (family or genera) of trees and clonal crops, and adapted for use in routine gene editing, they could be powerful methods for gene editing. A great deal of research on the molecular virology of tree and clonal crops, and broadly compatible and better delivery vectors suited for them, is clearly needed.
Class IV. Accelerated Flowering to Enable Transgene Removal Via Sexual Segregation in Trees and Clonal Crops
Accelerated flowering in trees, and associated rapid breeding, has been demonstrated in several species (Zhang et al. 2010; Srinivasan et al. 2012; Ye et al. 2014; Klocko et al. 2016), enabling sexual segregation to remove gene editing components. Of course, such strategies could not be employed on events with engineered sterility for containment (e.g., some proposed plantation forest trees) or sterile genotypes (e.g., peppermint). In many cases due to the absence of transformability of desirable elite cultivars, editing will likely need to be performed in a transformable genotype, followed by successive backcrossing to the elite cultivar to obtain marketable lines. This has been demonstrated as a viable strategy in plum for the introgression of plum pox virus resistance traits (Petri et al. 2018), where there is a single gene being introgressed. As a result of rapid flowering, the desired introgression can be completed in less than a decade, as opposed to the multiple decades such an effort in plum would normally require. For more complex breeding programs where there are polygenic trait targets and the need for many gene donors, FT transformation of many genotypes is likely to be needed, and each sexual generation will need to be followed by phenotypic and/or genome marker evaluations—slowing breeding substantially. Gene editing would of course add to this complexity.
These techniques mostly depend on the florigen encoding gene FLOWERING LOCUS T (FT), or associated floral regulatory genes (e.g., BpMADS4 or CENTRORADIALIS/TERMINAL FLOWER 1), that are overexpressed or suppressed (Flachowsky et al. 2009; Zhang et al. 2010; Flachowsky et al. 2011; Petri et al. 2018). To combine this technology with gene editing for transgene segregation in trees and clonal crops, an FT overexpression gene could be contained within the CRISPR/Cas9 T-DNA (Fig. 1). Another approach could include the grafting of CRISPR/Cas9 transgenic material with desired edits onto an FT overexpressing rootstock, as further discussed below. A more advanced form of these methods could be the generation of a Cas9 and FTox stable integrated line coupled with the viral delivery of desired gRNAs carried by systemic infection of an RNA virus such as TMV (Fig. 3). In this hypothetical case, a single transgenic line could serve as source material for the introduction of many different engineered traits. This would be highly desirable for species or genotypes with very low transformation rates—as is the case with many trees, other woody perennials, and clonally propagated crops.
One major challenge of employing strategies dependent on the grafting of transgenic materials containing editing transgenes is the graft transmissibility of FT protein, which has been shown in several instances when expressed via the commonly used 35S promoter not to be graft transmissible (Tränkner et al. 2010; Zhang et al. 2010; Zhang et al. 2010; Wenzel et al. 2013), with notable exceptions found in blueberry, Jatropha curcas, and citrus (Ye et al. 2014; Song et al. 2019; Soares et al. 2020). Expression from phloem-companion cell-specific promoters (e.g., SUCROSE SYMPORTER 2 (SUC2)) rather than completely constitutive ones may increase the abundance of exportable FT protein to reach nearby grafted branches, as was recently shown in citrus (Soares et al. 2020). Presumably, this improvement is because although 35S overexpression of FT is able to induce early flowering through direct expression at the shoot apex, it is insufficient in the phloem-companion cells to be able to reach the thresholds required for flowering when transmitted through graft junctions. Further work is needed to establish the generality of such FT expression approaches for gene editing in trees and other woody perennial crops.
Recent Examples of Editing in Use in Trees and Clonally Propagated Crops, and Perspectives on Future Traits of Interest
A large number of engineered traits have recently been published in the literature or in conference abstracts in trees and clonally propagated crops. A summary of these recent demonstrations within the last 3 yr is available in Table 1 and includes members of 21 genera. At present, all reported demonstrations of CRISPR/Cas9 editing with the exception of two studies employing recombinase-mediated excision in apple and grape, and six instances of DNA-free Cas9-RNP delivery, had the editing transgenes stably integrated into the genome and there was no attempt to excise or segregate them away (Dalla Costa et al. 2020; Pompili et al. 2020). 19 of the 87 studies listed simply demonstrated the function of CRISPR/Cas9-mediated editing in a given species through mutation of a visual marker such as PDS. The types of traits targeted with the studied gRNAs included sexual reproduction, herbicide resistance, growth habit (dwarfism/semi-dwarfism), disease resistance, secondary xylem growth, lignin biosynthesis, onset of flowering, nodulation, concentration of toxic metabolites, starch or sugar accumulation, shelf life of fruits, biofortification, and metal transport. Of the 21 genera where editing has been validated, unsurprisingly, the most studied are in Populus (18), potato (15), Citrus (9), strawberry (9), and grape (6). Six of the 87 studies summarized in Table 1 used Cas9-RNPs for transgene-free gene editing, relying on protoplast regeneration and delivery via PEG or biolistic bombardment. These studies were in potato, grape, rubber tree, apple, and banana (Malnoy et al. 2016; Malnoy et al. 2016; Andersson et al. 2018; Fan et al. 2020; González et al. 2020; Wu et al. 2020). Two studies employed transient delivery of editing reagents by Agrobacterium without integration and only one succeeded to obtain edited, transgene-free, non-chimeric regenerants (Charrier et al. 2019; Bánfalvi et al. 2020). These studies were in potato and apple. No studies used virally delivered editing reagents in a tree or clonal crop. All the remaining studies used stable introduction of editing genes via Agrobacterium. The current method of choice for generation of transgene-free, edited lines is protoplast delivery of Cas9-RNPs, a technique which demands a large deal of technical expertise and is likely to be inaccessible for the vast majority of valuable tree and clonally propagated crop genotypes and species.
Regulatory Context for Gene-Edited Trees and Clonally Propagated Crops
US Regulation
In the USA, the 1986 Coordinated Framework for the Regulation of Biotechnology (“Coordinated Framework”) focused on regulating the process of recombinant DNA (“rDNA”) when used in plant and animal breeding (OSTP 1986). At first, the USDA-APHIS lightly regulated gene editing of crops under a “Am I Regulated” process (APHIS 81 FR 65414 2017). The focus was on whether the DNA had a plant pest gene and without such DNA sequences (i.e., most of the crops submitted through this process), there was no legal hook for the USDA to regulate gene-edited crops at all (Martínez-Fortún et al. 2017).
In May 2020, the USDA-APHIS announced the Final Rule for its biotechnology regulations 7 CFR part 340, called the Sustainable, Ecological, Consistent, Uniform, Responsible, Efficient (SECURE) rule (APHIS, 2020). Thus, going forward, the USDA’s Biotechnology Regulatory Services (“BRS”) will implement this rule in assessing the environmental safety of biotech crops. The 2020 Rule defines genetic engineering broadly as “[t]echniques that use recombinant, synthesized, or amplified nucleic acids to modify or create a genome” (7 CFR § 340.3., 2021). Under USDA’s new “Regulatory Status Review (RSR)” that is part of SECURE, developers can ask APHIS to evaluate novel plants and decide if those plants fall within the 2020 Rule’s permit scope (but requires less data than the petition process it replaces, including an explicit statement that field trial data are not generally required). In a historically significant move, the USDA in this Rule focused on the product, not the process, used to make the organism.
The 2020 Rule also exempts certain new and old organisms from regulation. These include certain types of innovative plant breeding methods (of most note, genome editing when the change could have been obtained by conventional breeding), plant-trait-mechanisms of action that have already been approved (i.e., MOAs, which are combinations of plant genera, gene functions, and traits) rather the individual gene insertions, and all “Am I Regulated” plants that were allowed under the past USDA process. In the future, the USDA could also exempt any plants whose modifications could have been achieved via conventional breeding. APHIS may grant an exemption itself or parties can request an exemption. APHIS will then provide public notice and opportunity for public comment if it agrees with the proposed exemption.
The rule also exempts minor DNA changes: (1) cellular repair of a targeted DNA break without an externally provided repair template, (2) a single deletion of any size, (3) natural DNA repair mechanisms, (4) targeted single base-pair substitutions (7 CFR § 340.1(b) (2), 2021), and (5) insertions from compatible plant relatives. Starting in August 2020, the USDA started letting developers request confirmation that their organism is exempt (normally within 120 d). Such confirmation letters will be posted on the APHIS website to “help them market their products domestically and overseas” (APHIS, 2020 at 29,799).
For regulated crops undergoing a RSR, if BRS finds no significant plant pest risk or other impacts after a review of the public comments under the National Environmental Policy Act (NEPA), the deregulation notice allows the developer to commercialize the biotech crop. The EPA also regulates those crops that resist herbicides (to approve herbicide uses and warnings) or pests covered under the Federal Insecticide, Fungicide, and Rodenticide Act (FIFRA). The FDA is expecting to continue to regulate genetically edited animals under its cumbersome veterinary drug approval process (Van Eenennaam et al. 2019) though in the final days of the Trump administration, the USDA attempted to transfer that regulatory role to them (Perdue 2021). After approximately 20 yr of regulatory requests, the FDA approved the AquaBounty® AquAdvantage® Salmon, highlighting the cumbersome nature of FDA regulation of animal biotechnology (Van Eenennaam and Muir 2011; FDA USF and DA 2019; Steinberg and Vittorio 2019). Many commentators are calling for a more reasonable approach than the FDA has taken with GE animals, particularly when there is no drug-related aspect (e.g., a gene to prevent allergy that does not influence the structure and function of the animal or intended eater).
International Regulation
The European Union’s High Court of Justice in July 2018 ruled that crops and other organisms produced through genetic editing will be regulated as if they were a “GMO” under its long-standing “precautionary approach” to regulatory approval. This means that approval times will take several years, sometimes longer, for crops to be approved in the EU after being approved by nations planting the gene-edited crops (Callaway 2018).
The EU’s “precautionary approach” is the law followed by the 171 nations that are parties to the Cartagena Protocol on Biosafety (“CPB”). These parties plan to meet again for the tenth time (COP-MOP 10) in early 2021 in Kunming, China. At MOP 9 in November 2018 in Egypt, the parties put gene editing under a “synthetic biology” descriptor. Since the USA, Canada, Australia, Argentina, and other grain exporting nations are not signatories to the CPB, we can expect parties to follow the lead of the EU, which is in many instances a key trading partner and source of foreign aid (e.g., in parts of Africa, where the EU’s influence on GMO policy is notably strong).
As part of the implementation of this law, nations that are parties to the Biosafety Protocol enact legislation, such as the European Traceability Directive, that imposes zero tolerance for the import of any GMO that lacks regulatory approval (The Cartagena Protocol on Biosafety to the Convention on Biological Diversity 2021). More nations are imposing regulatory approval requirements as the Biosafety Protocol is implemented. Thus, any biotech crop that could be exported, including gene-edited crops or derived foods, may also require approval in many of these overseas markets. Similar problems exist for the potential use and export of GMO or gene-edited forestry crops (Strauss et al. 2009)
Canada regulates all “novel foods” and includes gene editing in that category. This encompasses those crops created using non-rDNA methods. For example, herbicide-resistant crops created using older methods such as chemical radiation, while newer methods such as gene editing, would be regulated. It is clear that those older forms of plant and animal “mutagenesis” breeding carry greater risks of “pleiotropic” changes (Wolt et al. 2016) that are often raised by activists opposed to gene editing (FOE F of the E (FOE) 2018) who see risks in off-target effects in genes (the adverse nature of which remain unlinked to health concerns). This is in spite of strong evidence that off-target mutagenesis from gene editing in crop plants (vs. animals) is negligible compared to other sources of natural and breeding-induced mutagenesis (Graham et al. 2020) Since many of these crops have similar ecological effects (e.g., there are mutagenic, rDNA and genetically edited crops with herbicide resistance, all of which can outcross to wild relatives or cause problematic herbicide-resistant weeds to develop after widespread use), Canada’s regime at least has a consistent approach to similar risks (Ellens et al. 2019).
In 2018, Argentina and 12 other nations, including other major grain exporting nations like Canada, Australia, Brazil, Paraguay, and the USA, issued a joint statement supporting agricultural applications of gene editing in agriculture (“precision biotechnology”), stating that governments should “avoid arbitrary and unjustifiable distinctions between end products (crop traits) derived from precision biotechnology and similar end products, obtained through other production methods.” Signatories were Argentina, Australia, Brazil, Canada, Colombia, the Dominican Republic, Guatemala, Honduras, Jordan, Paraguay, the USA, Uruguay, Vietnam, and the Secretariat of the Economic Community of West African States (WTO 2018). We hope that common sense and good science will prevail over the arbitrary and capricious “precautionary approach” to regulating gene editing that may be applied to these products under the Cartagena Protocol on Biosafety (Nill et al. 2000).
In the USA, there are two routes for opponents of this technology to seek to stop the launch of a gene-edited crop, both of which relate to economic and environmental impacts related to the National Environmental Policy Act. First, injunctions to stop the launch of biotech crops have been granted against beets, eucalypt trees, and alfalfa for “interrelated economic effects” and forced environmental reviews after the USDA had conducted environmental assessments (but not a full environmental impact statement). Second, where there is an export-related economic interest involved, either an injunction under “anticipatory nuisance” (Grossman 2008) or post-marketing litigation seeking recovery for economic impacts to export-related interests (Ledbetter 2018). Depending on the trait modified and its connection to environmental and economic or market impacts, such routes may indeed be fruitful and delay their integration into the marketplace. To avoid such tactics, new laws that coordinate regulations across agencies in directing attention to comparative outcomes vs. use of recombinant methods, similar to what SECURE is hoping to achieve, might be needed in the USA. And to avoid trade confusion, similar new treaties are needed at the international level, whether inside or outside of the CPB.
Conclusions
Gene editing is growing rapidly in its application to trees and clonally propagated crops. The high efficiency that has been observed in annual crops clearly also applies to trees and clonal crops. The surge in publications and modified traits also suggests both a high scientific and a high commercial demand for its capabilities. However, the large majority of the publications to date have not had the gene editing machinery removed from the modified plants, confirming its difficulty in these crops. This is likely to be a result of their difficult breeding systems, delayed onset of reproduction, and general recalcitrance to transformation and regeneration of modified plants. Gene editing methods that do not insert transgenes, or seek to excise them after integration, though feasible and a subject of active research, have been rarely used and appear highly inefficient. We therefore believe that major innovations are needed both in methods for transient editing and excision, and in national and international regulation. Biological innovations should include the further development of widely effective viral systems for transient editing and efficient inducible recombinase systems for excision. Regulatory innovations should seek to focus on the products and their risk/benefit comparisons to conventional breeding methods, similar to that in the USDA SECURE system, thus not needlessly penalizing the use of recombinant DNA methods and the addition of inert (e.g., non-coding) and/or well-studied and harmless DNA elements (e.g., recombinase sites, T-DNA borders, common selectable markers) used when making new gene-edited varieties. Similar recommendations have been made many times with respect to transgenic crop regulation (e.g., Strauss 2003; Bradford et al. 2005) Indeed, CRISPR-associated genes, even if left in trees or clonal crops used for food or feed, appear to have little potential for food safety or allergy problems (Nakajima et al. 2016), nor as discussed above as gene drives should they enter wild or feral populations.
Although the new USDA SECURE regulations provide several avenues by which trees or clonally propagated and gene-edited crops could be exempted from regulatory oversight or minimally regulated through the RSR process, serious regulatory and legal obstacles may remain given the uncertainty in how the regulations actually become implemented in practice, and due to the limited harmonization among the USDA, EPA, and FDA in their gene editing rules (e.g., in pest-resistant vs. other types of crops by the EPA vs. USDA). In addition, the NEPA process provides an avenue for opponents to greatly delay or block products, and the CBD is capable of blocking or greatly delaying all products intended for international markets. Given the great functional similarity of many gene-edited products with those from conventional breeding, gene editing is challenging regulatory systems to rethink their foundations, with increased emphasis on product over process, which scientists have recommended for GMO crops since the 1980s (National Academies of Sciences, Engineering, and Medicine 2016). With a decades-long record of GMO food and tree safety (Walter et al. 2010) and a world increasingly unable to produce its food and fiber sustainably, we hope that leaders take full advantage of this moment to craft more scientifically based systems, rather than cling to outdated method-based laws.
References
Akagi T, Pilkington SM, Varkonyi-Gasic E, Henry IM, Sugano SS, Sonoda M, Firl A, McNeilage MA, Douglas MJ, Wang T et al (2019) Two Y-chromosome-encoded genes determine sex in kiwifruit. Nat Plants 5:801–809
Ali Z, Shami A, Sedeek K, Kamel R, Alhabsi A, Tehseen M, Hassan N, Butt H, Kababji A, Hamdan SM et al (2020) Fusion of the Cas9 endonuclease and the VirD2 relaxase facilitates homology-directed repair for precise genome engineering in rice. Commun Biol 3:44
Alvin C-HL (2020) Gene editing of cultivated and wild strawberry (Fragaria x ananassa and Fragaria vesca) with CRISPR/Cas9 knocking out F3H. Norwegian University of Life Sciences, Ås, NO
An Y, Geng Y, Yao J, Fu C, Lu M, Wang C, Du J (2020) Efficient genome editing in Populus using CRISPR/Cas12a. Front Plant Sci 11:1776
Andersson M, Turesson H, Olsson N, Fält A-S, Ohlsson P, Gonzalez MN, Samuelsson M, Hofvander P (2018) Genome editing in potato via CRISPR-Cas9 ribonucleoprotein delivery. Physiol Plant 164:378–384
Anzalone AV, Randolph PB, Davis JR, Sousa AA, Koblan LW, Levy JM, Chen PJ, Wilson C, Newby GA, Raguram A et al (2019) Search-and-replace genome editing without double-strand breaks or donor DNA. Nature 576:149–157
APHIS 81 FR 65414 (2017) Modernizing the regulatory system for biotechnology products: final version of the 2017 update to the coordinated framework for the regulation of biotechnology. https://www.aphis.usda.gov/biotechnology/downloads/2017_coordinated_framework_update.pdf
Ariga H, Toki S, Ishibashi K (2020) Potato Virus X vector-mediated DNA-free genome editing in plants. Plant Cell Physiol 61:1946–1953
Awasthi P, Kocábek T, Mishra AK, Nath VS, Shrestha A, Matoušek J (2021) Establishment of CRISPR/Cas9 mediated targeted mutagenesis in hop (Humulus lupulus). Plant Physiol Biochem 160:1–7
Azeez A, Busov V (2021) CRISPR/Cas9-mediated single and biallelic knockout of poplar STERILE APETALA (PopSAP) leads to complete reproductive sterility. Plant Biotechnol J 19:23–25
Bairu MW, Aremu AO, Van Staden J (2011) Somaclonal variation in plants: causes and detection methods. Plant Growth Regul 63:147–173
Banakar R, Eggenberger AL, Lee K, Wright DA, Murugan K, Zarecor S, Lawrence-Dill CJ, Sashital DG, Wang K (2019) High-frequency random DNA insertions upon co-delivery of CRISPR-Cas9 ribonucleoprotein and selectable marker plasmid in rice. Sci Rep 9:19902
Bánfalvi Z, Csákvári E, Villányi V, Kondrák M (2020) Generation of transgene-free PDS mutants in potato by Agrobacterium-mediated transformation. BMC Biotechnol 20:25
Begemann M, Gray B, January E, Gordon G, He Y, Liu H, Wu X, Brutnell T, Mockler T, Oufattole M (2017) Precise insertion and guided editing of higher plant genomes using Cpf1 CRISPR nucleases. Sci Rep 7:11606
Bharat SS, Li S, Li J, Yan L, Xia L (2020) Base editing in plants: current status and challenges. Crop Genome Ed Big Step Breed Des 8:384–395
Birch RG (1997) Plant transformation: problems and Strategies for Practical Application. Annu Rev Plant Physiol Plant Mol Biol 48:297–326
Bradford KJ, Van Deynze A, Gutterson N, Parrott W, Strauss SH (2005) Regulating transgenic crops sensibly: lessons from plant breeding, biotechnology and genomics. Nat Biotechnol 23:439–444
Brand A, Quimbaya M, Tohme J, Chavarriaga-Aguirre P (2019) Arabidopsis LEC1 and LEC2 orthologous genes are key regulators of somatic embryogenesis in cassava. Front Plant Sci 10:673
Breitler J-C, Dechamp E, Campa C, Zebral Rodrigues LA, Guyot R, Marraccini P, Etienne H (2018) CRISPR/Cas9-mediated efficient targeted mutagenesis has the potential to accelerate the domestication of Coffea canephora. Plant Cell Tissue Organ Cult PCTOC 134:383–394
Busov VB, Brunner AM, Meilan R, Filichkin S, Ganio L, Gandhi S, Strauss SH (2005) Genetic transformation: a powerful tool for dissection of adaptive traits in trees. New Phytol 167:9–18
Callaway E (2018) CRISPR plants now subject to tough GM laws in European Union. Nature 560:16–16
Čermák T, Baltes NJ, Čegan R, Zhang Y, Voytas DF (2015) High-frequency, precise modification of the tomato genome. Genome Biol 16:232
Čermák T, Curtin S, Gil-Humanes J, Čegan R, Kono T, Konečná E, Belanto J, Starker C, Mathre J, Greenstein R, Voytas D (2017) A multipurpose toolkit to enable advanced genome engineering in plants. The Plant Cell 29 (6):1196–1217
Charrier A, Vergne E, Dousset N, Richer A, Petiteau A, Chevreau E (2019) Efficient targeted mutagenesis in apple and pear using the CRISPR-Cas9 system. Front Plant Sci 10:40
Chatukuta P, Rey MEC (2020) A cassava protoplast system for screening genes associated with the response to South African cassava mosaic virus. Virol J 17:184
Chen L, Li W, Katin-Grazzini L, Ding J, Gu X, Li Y, Gu T, Wang R, Lin X, Deng Z et al (2018) A method for the production and expedient screening of CRISPR/Cas9-mediated non-transgenic mutant plants. Hortic Res 5:13
Chen K, Wang Y, Zhang R, Zhang H, Gao C (2019) CRISPR/Cas Genome editing and precision plant breeding in agriculture. Annu Rev Plant Biol 70:667–697
Chen S, Lee B, Lee AY-F, Modzelewski AJ, He L (2016) Highly efficient mouse genome editing by CRISPR ribonucleoprotein electroporation of zygotes. J Biol Chem 291:14457–14467
Dai Y, Hu G, Dupas A, Medina L, Blandels N, San Clemente H, Ladouce N, Badawi M, Hernandez-Raquet G, Mounet F et al (2020) Implementing the CRISPR/Cas9 technology in Eucalyptus hairy roots using wood-related genes. Int J Mol Sci
Dalla Costa L, Piazza S, Pompili V, Salvagnin U, Cestaro A, Moffa L, Vittani L, Moser C, Malnoy M (2020) Strategies to produce T-DNA free CRISPRed fruit trees via Agrobacterium tumefaciens stable gene transfer. Sci Rep 10:20155
De Meester B, Madariaga Calderón B, de Vries L, Pollier J, Goeminne G, Van Doorsselaere J, Chen M, Ralph J, Vanholme R, Boerjan W (2020) Tailoring poplar lignin without yield penalty by combining a null and haploinsufficient CINNAMOYL-CoA REDUCTASE2 allele. Nat Commun 11:5020
Debernardi JM, Tricoli DM, Ercoli MF, Hayta S, Ronald P, Palatnik JF, Dubcovsky J (2020) A GRF–GIF chimeric protein improves the regeneration efficiency of transgenic plants. Nat Biotechnol 38:1274–1279
Della Maggiora L (2020) CRISPR/Cas9 – technology in poplar: example of an approach for the production of uniform knockout mutants for the metal transporter MTP1. Sant’Anna Scuola Universitaria Superiore Pisa, Pisa, IT
Ding L, Chen Y, Ma Y, Wang H, Wei J (2020) Effective reduction in chimeric mutants of poplar trees produced by CRISPR/Cas9 through a second round of shoot regeneration. Plant Biotechnol Rep 14:549–558
Dong O, Yu S, Jain R, Zhang N, Duong P, Butler C, Li Y, Lipzen A, Martin J, Barry K, Schmutz J, Tian L, Ronald P (2020) Marker-free carotenoid-enriched rice generated through targeted gene insertion using CRISPR-Cas9. Nature Communications 11 (1)
Duan N, Bai Y, Sun H, Wang N, Ma Y, Li M, Wang X, Jiao C, Legall N, Mao L et al (2017) Genome re-sequencing reveals the history of apple and supports a two-stage model for fruit enlargement. Nat Commun 8:249
Dutt M, Mou Z, Zhang X, Tanwir SE, Grosser JW (2020) Efficient CRISPR/Cas9 genome editing with Citrus embryogenic cell cultures. BMC Biotechnol 20:58
Ebinuma H, Sugita K, Matsunaga E, Yamakado M (1997) Selection of marker-free transgenic plants using the isopentenyl transferase gene. Proc Natl Acad Sci U S A 94:2117–2121
Ellens KW, Levac D, Pearson C, Savoie A, Strand N, Louter J, Tibelius C (2019) Canadian regulatory aspects of gene editing technologies. Transgenic Res 28:165–168
Ellison EE, Nagalakshmi U, Gamo ME, Huang P, Dinesh-Kumar S, Voytas DF (2020) Multiplexed heritable gene editing using RNA viruses and mobile single guide RNAs. Nat Plants 6:620–624
Elorriaga E, Klocko AL, Ma C, Strauss SH (2018) Variation in mutation spectra among CRISPR/Cas9 mutagenized poplars. Front Plant Sci 9:594
Elorriaga E, Klocko A, Ma C, du Plessis M, An X, Myburg A, and Strauss S (2021) Genetic containment in vegetatively propagated forest trees: CRISPR disruption of LEAFY function in Eucalyptus gives sterile indeterminate inflorescences and normal juvenile development. Plant Biotechnol J
Enciso-Rodriguez F, Manrique-Carpintero NC, Nadakuduti SS, Buell CR, Zarka D, Douches D (2019) Overcoming self-incompatibility in diploid potato using CRISPR-Cas9. Front Plant Sci 10:376
FDA USF and DA (2019) AquAdvantage Salmon. https://www.fda.gov/animal-veterinary/animals-intentional-genomic-alterations/questions-and-answers-fdas-deactivation-import-alert-genetically-engineered-salmon
Fan D, Wang X, Tang X, Ye X, Ren S, Wang D, Luo K (2018) Histone H3K9 demethylase JMJ25 epigenetically modulates anthocyanin biosynthesis in poplar. Plant J 96:1121–1136
Fan Y, Xin S, Dai X, Yang X, Huang H, Hua Y (2020) Efficient genome editing of rubber tree (Hevea brasiliensis) protoplasts using CRISPR/Cas9 ribonucleoproteins. Ind Crop Prod 146:112146
Fellenberg C, Corea O, Yan L-H, Archinuk F, Piirtola E-M, Gordon H, Reichelt M, Brandt W, Wulff J, Ehlting J et al (2020) Discovery of salicyl benzoate UDP-glycosyltransferase, a central enzyme in poplar salicinoid phenolic glycoside biosynthesis. Plant J 102:99–115
Feng J, Dai C, Luo H, Han Y, Liu Z, Kang C (2019) Reporter gene expression reveals precise auxin synthesis sites during fruit and root development in wild strawberry. J Exp Bot 70:563–574
Fister AS, Landherr L, Maximova SN, Guiltinan MJ (2018) Transient expression of CRISPR/Cas9 machinery targeting TcNPR3 enhances defense response in Theobroma cacao. Front Plant Sci 9:268
Flachowsky H, Hanke M-V, Peil A, Strauss SH, Fladung M (2009) A review on transgenic approaches to accelerate breeding of woody plants. Plant Breed 128:217–226
Flachowsky H, Le Roux P-M, Peil A, Patocchi A, Richter K, Hanke M-V (2011) Application of a high-speed breeding technology to apple (Malus × domestica) based on transgenic early flowering plants and marker-assisted selection. New Phytol 192:364–377
Fladung M, Becker D (2010) Targeted integration and removal of transgenes in hybrid aspen (Populus tremula L. x P. tremuloides Michx.) using site-specific recombination systems. Plant Biol 12:334–340
Fladung M, Schenk TMH, Polak O, Becker D (2010) Elimination of marker genes and targeted integration via FLP/FRT recombination system from yeast in hybrid aspen (Populus tremula L. × P. tremuloides Michx.). Tree Genet Genomes 6:205–217
Florez SL, Erwin RL, Maximova SN, Guiltinan MJ, Curtis WR (2015) Enhanced somatic embryogenesis in Theobroma cacao using the homologous BABY BOOM transcription factor. BMC Plant Biol 15:121
FOE F of the E (FOE) (2018) New report: gene editing in agriculture poses new risks to health, environment. https://foe.org/news/new-report-gene-editing-agriculture-poses-new-risks-health-environment/
Fossi M, Amundson K, Kuppu S, Britt A, Comai L (2019) Regeneration of Solanum tuberosum plants from protoplasts induces widespread genome instability. Plant Physiol 180:78
Gao Q, Luo H, Li Y, Liu Z, Kang C (2020) Genetic modulation of RAP alters fruit coloration in both wild and cultivated strawberry. Plant Biotechnol J 18:1550–1561
Ghoshal B, Vong B, Picard CL, Feng S, Tam JM, Jacobsen SE (2020) A viral guide RNA delivery system for CRISPR-based transcriptional activation and heritable targeted DNA demethylation in Arabidopsis thaliana. PLoS Genet 16:e1008983
Gomez MA, Lin ZD, Moll T, Chauhan RD, Hayden L, Renninger K, Beyene G, Taylor NJ, Carrington JC, Staskawicz BJ et al (2019) Simultaneous CRISPR/Cas9-mediated editing of cassava eIF4E isoforms nCBP-1 and nCBP-2 reduces cassava brown streak disease symptom severity and incidence. Plant Biotechnol J 17:421–434
González MN, Massa GA, Andersson M, Turesson H, Olsson N, Fält A-S, Storani L, Décima Oneto CA, Hofvander P, Feingold SE (2020) Reduced enzymatic browning in potato tubers by specific editing of a polyphenol oxidase gene via ribonucleoprotein complexes delivery of the CRISPR/Cas9 system. Front Plant Sci 10:1649
Graham N, Patil GB, Bubeck DM, Dobert RC, Glenn KC, Gutsche AT, Kumar S, Lindbo JA, Maas L, May GD et al (2020) Plant genome editing and the relevance of off-target changes. Plant Physiol 183:1453
Grossman MR (2008) Anticipatory nuisance and the prevention of environmental harm and economic loss from GMOs in the United States. J Environ Law Pract 2:107–194
Hoerster G, Wang N, Ryan L, Wu E, Anand A, McBride K, Lowe K, Jones T, Gordon-Kamm B (2020) Use of non-integrating Zm-Wus2 vectors to enhance maize transformation. In Vitro Cell Dev Biol - Plant 56:265–279
Hoshijima K, Jurynec MJ, Klatt Shaw D, Jacobi AM, Behlke MA, Grunwald DJ (2019) Highly efficient CRISPR-Cas9-based methods for generating deletion mutations and F0 embryos that lack gene function in zebrafish. Dev Cell 51:645–657.e4
Hu C, Sheng O, Deng G, He W, Dong T, Yang Q, Dou T, Li C, Gao H, Liu S et al (2020) CRISPR/Cas9-mediated genome editing of MaACO1 (aminocyclopropane-1-carboxylate oxidase 1) promotes the shelf life of banana fruit. Plant Biotechnol J
Hu J, Li S, Li Z, Li H, Song W, Zhao H, Lai J, Xia L, Li D, Zhang Y (2019) A barley stripe mosaic virus-based guide RNA delivery system for targeted mutagenesis in wheat and maize. Mol Plant Pathol 20:1463–1474
Hu Q, Kononowicz-Hodges H, Nelson-Vasilchik K, Viola D, Zeng P, Liu H, Kausch AP, Chandlee JM, Hodges TK, Luo H (2008) FLP recombinase-mediated site-specific recombination in rice. Plant Biotechnol J 6:176–188
Huang X, Wang Y, Xu J, Wang N (2020) Development of multiplex genome editing toolkits for citrus with high efficacy in biallelic and homozygous mutations. Plant Mol Biol 104:297–307
Hummel AW, Chauhan RD, Cermak T, Mutka AM, Vijayaraghavan A, Boyher A, Starker CG, Bart R, Voytas DF, Taylor NJ (2018) Allele exchange at the EPSPS locus confers glyphosate tolerance in cassava. Plant Biotechnol J 16:1275–1282
Jia H, Orbović V, Wang N (2019) CRISPR-LbCas12a-mediated modification of citrus. Plant Biotechnol J 17:1928–1937
Jia H, Wang N (2020) Generation of homozygous canker-resistant citrus in the T0 generation using CRISPR-SpCas9p. Plant Biotechnol J 18:1990–1992
Jia H, Xu J, Orbović V, Zhang Y, Wang N (2017a) Editing Citrus genome via SaCas9/sgRNA System. Front Plant Sci 8:2135
Jia H, Zhang Y, Orbović V, Xu J, White FF, Jones JB, Wang N (2017b) Genome editing of the disease susceptibility gene CsLOB1 in citrus confers resistance to citrus canker. Plant Biotechnol J 15:817–823
Jo YD, Kim J-B (2019) Frequency and spectrum of radiation-induced mutations revealed by whole-genome sequencing analyses of plants. Quantum Beam Sci.
Johansen IE, Liu Y, Jørgensen B, Bennett EP, Andreasson E, Nielsen KL, Blennow A, Petersen BL (2019) High efficacy full allelic CRISPR/Cas9 gene editing in tetraploid potato. Sci Rep 9:17715
Kannan B, Jung JH, Moxley GW, Lee S-M, Altpeter F (2018) TALEN-mediated targeted mutagenesis of more than 100 COMT copies/alleles in highly polyploid sugarcane improves saccharification efficiency without compromising biomass yield. Plant Biotechnol J 16:856–866
Kaur N, Alok A, Shivani KN, Pandey P, Awasthi P, Tiwari S (2018) CRISPR/Cas9-mediated efficient editing in phytoene desaturase (PDS) demonstrates precise manipulation in banana cv. Rasthali genome. Funct Integr Genomics 18:89–99
Kaur N, Alok A, Shivani KP, Kaur N, Awasthi P, Chaturvedi S, Pandey P, Pandey A, Pandey AK et al (2020) CRISPR/Cas9 directed editing of lycopene epsilon-cyclase modulates metabolic flux for β-carotene biosynthesis in banana fruit. Metab Eng 59:76–86
Kilby NJ, Davies GJ, Snaith MR, Murray JAH (1995) FLP recombinase in transgenic plants: constitutive activity in stably transformed tobacco and generation of marked cell clones in Arabidopsis. Plant J 8:637–652
Kim H-B, Cho J-I, Ryoo N, Qu S, Wang G-L, Jeon J-S (2012) Development of a simple and efficient system for excising selectable markers in Arabidopsis using a minimal promoter::Cre fusion construct. Mol Cell 33:61–69
Klocko AL, Ma C, Robertson S, Esfandiari E, Nilsson O, Strauss SH (2016) FT overexpression induces precocious flowering and normal reproductive development in Eucalyptus. Plant Biotechnol J 14:808–819
Kondrák M, van der Meer IM, Bánfalvi Z (2006) Generation of marker- and backbone-free transgenic potatoes by site-specific recombination and a bi-functional marker gene in a non-regular one-border agrobacterium transformation vector. Transgenic Res 15:729–737
Kusano H, Ohnuma M, Mutsuro-Aoki H, Asahi T, Ichinosawa D, Onodera H, Asano K, Noda T, Horie T, Fukumoto K et al (2018) Establishment of a modified CRISPR/Cas9 system with increased mutagenesis frequency using the translational enhancer dMac3 and multiple guide RNAs in potato. Sci Rep 8:13753
Ledbetter, K. (2018) Corn producers receiving Syngenta litigation settlement notices have options, The Eagle, https://theeagle.com/landandlivestockpost/agrilifetoday/corn-producers-receiving-syngenta-litigation-settlement-notices-have-options/article_a73a0a6e-9e9f-56e7-bd73-342a58e35c33.html
Li C, Zong Y, Wang Y, Jin S, Zhang D, Song Q, Zhang R, Gao C (2018) Expanded base editing in rice and wheat using a Cas9-adenosine deaminase fusion. Genome Biol 19:59
Li M-Y, Jiao Y-T, Wang Y-T, Zhang N, Wang B-B, Liu R-Q, Yin X, Xu Y, Liu G-T (2020a) CRISPR/Cas9-mediated VvPR4b editing decreases downy mildew resistance in grapevine (Vitis vinifera L.). Hortic Res 7:149
Li Z, Wang Y, Lu X, Li R, Liu J, Fu S, Hu X, Guo J, Yao Y (2020b) Construction and verification of CRISPR/Cas9 gene editing vector for cassava MeSSIII gene. Mol Plant Breed
Lin Q, Zong Y, Xue C, Wang S, Jin S, Zhu Z, Wang Y, Anzalone AV, Raguram A, Doman JL et al (2020) Prime genome editing in rice and wheat. Nat Biotechnol 38:582–585
Liu R, Long Q, Zou X, Wang Y, Pei Y (2021) DNA methylation occurring in Cre-expressing cells inhibits loxP recombination and silences loxP-sandwiched genes. New Phytol 231:210–224
Lowe K, Wu E, Wang N, Hoerster G, Hastings C, Cho M-J, Scelonge C, Lenderts B, Chamberlin M, Cushatt J et al (2016) Morphogenic regulators baby boom and Wuschel improve monocot transformation. Plant Cell 28:1998–2015
Ma W, Kang X, Liu P, Zhang Y, Lin X, Li B, Chen Z (2021) The analysis of transcription factor CsHB1 effects on caffeine accumulation in tea callus through CRISPR/Cas9 mediated gene editing. Process Biochem 101:304–311
Maher MF, Nasti RA, Vollbrecht M, Starker CG, Clark MD, Voytas DF (2020) Plant gene editing through de novo induction of meristems. Nat Biotechnol 38:84–89
Malnoy M, Viola R, Jung M-H, Koo O-J, Kim S, Kim J-S, Velasco R, Nagamangala Kanchiswamy C (2016) DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front Plant Sci 7:1904–1904
Malzahn AA, Tang X, Lee K, Ren Q, Sretenovic S, Zhang Y, Chen H, Kang M, Bao Y, Zheng X et al (2019) Application of CRISPR-Cas12a temperature sensitivity for improved genome editing in rice, maize, and Arabidopsis. BMC Biol 17:9
Martínez-Fortún J, Phillips DW, Jones HD (2017) Potential impact of genome editing in world agriculture. Emerg Top Life Sci 1:117–133
Martín-Pizarro C, Triviño JC, Posé D (2019) Functional analysis of the TM6 MADS-box gene in the octoploid strawberry by CRISPR/Cas9-directed mutagenesis. J Exp Bot 70:885–895
Metje-Sprink J, Menz J, Modrzejewski D, Sprink T (2019) DNA-free genome editing: past, present and future. Front Plant Sci 9:1957–1957
Milner MJ, Craze M, Hope MS, Wallington EJ (2020) Turning up the temperature on CRISPR: increased temperature can improve the editing efficiency of wheat using CRISPR/Cas9. Front Plant Sci 11:1780
Modrzejewski D, Hartung F, Lehnert H, Sprink T, Kohl C, Keilwagen J, Wilhelm R (2020) Which factors affect the occurrence of off-target effects caused by the use of CRISPR/Cas: a systematic review in plants. Front Plant Sci 11:1838
Movahedi A, Wei H, Chen Z-H, Sun W, Zhang J, Li D, Yang L, Zhuge Q (2020) NHEJ deficiency develops homologous recombination in poplar meaningfully further than the overexpression of HDR factors. bioRxiv 2020.07.04.188219
Muhr M, Paulat M, Awwanah M, Brinkkötter M, Teichmann T (2018) CRISPR/Cas9-mediated knockout of Populus BRANCHED1 and BRANCHED2 orthologs reveals a major function in bud outgrowth control. Tree Physiol 38:1588–1597
Müller NA, Kersten B, Leite Montalvão AP, Mähler N, Bernhardsson C, Bräutigam K, Carracedo Lorenzo Z, Hoenicka H, Kumar V, Mader M et al (2020) A single gene underlies the dynamic evolution of poplar sex determination. Nat Plants 6:630–637
Murovec J, Guček K, Bohanec B, Avbelj M, Jerala R (2018) DNA-free genome editing of Brassica oleracea and B. rapa protoplasts using CRISPR-Cas9 ribonucleoprotein complexes. Front Plant Sci 9:1594–1594
Mysore KS, Bassuner B, Deng X, Darbinian NS, Motchoulski A, Ream W, Gelvin SB (1998) Role of the Agrobacterium tumefaciens VirD2 protein in T-DNA transfer and integration. Mol Plant-Microbe Interactions 11:668–683
Nadakuduti SS, Buell CR, Voytas DF, Starker CG, Douches DS (2018) Genome editing for crop improvement – applications in clonally propagated polyploids with a focus on potato (Solanum tuberosum L.). Front Plant Sci 9:1607
Nagle M, Déjardin A, Pilate G, Strauss SH (2018) Opportunities for innovation in genetic transformation of forest trees. Front Plant Sci 9:1443–1443
Naim F, Dugdale B, Kleidon J, Brinin A, Shand K, Waterhouse P, Dale J (2018) Gene editing the phytoene desaturase alleles of Cavendish banana using CRISPR/Cas9. Transgenic Res 27:451–460
Nakajima I, Ban Y, Azuma A, Onoue N, Moriguchi T, Yamamoto T, Toki S, Endo M (2017) CRISPR/Cas9-mediated targeted mutagenesis in grape. PLoS One 12:e0177966
Nakajima O, Nishimaki-Mogami T, Kondo K (2016) Cas9 in genetically modified food is unlikely to cause food allergy. Biol Pharm Bull 39:1876–1880
Nakayasu M, Akiyama R, Lee HJ, Osakabe K, Osakabe Y, Watanabe B, Sugimoto Y, Umemoto N, Saito K, Muranaka T et al (2018) Generation of α-solanine-free hairy roots of potato by CRISPR/Cas9 mediated genome editing of the St16DOX gene. Genome Ed Technol Plant Physiol 131:70–77
National Academies of Sciences, Engineering, and Medicine (2016) Genetically Engineered Crops: Experiences and Prospects. Washington, DC: The National Academies Press. https://doi.org/10.17226/23395
Nill K, Redick T, Drew N (2000) Precautionary priority in approving imports of genetically improved commodity crops. Biotechnol Law Rep - BIOTECHNOL LAW REP 19:546–559
Nishizawa-Yokoi A, Saika H, Hara N, Lee L-Y, Toki S, Gelvin SB (2021) Agrobacterium T-DNA integration in somatic cells does not require the activity of DNA polymerase θ. New Phytol 229:2859–2872
Nishizawa-Yokoi A, Toki S (2021) A piggyBac-mediated transgenesis system for the temporary expression of CRISPR/Cas9 in rice. Plant Biotechnol J
Ntui VO, Tripathi JN, Tripathi L (2020) Robust CRISPR/Cas9 mediated genome editing tool for banana and plantain (Musa spp.). Spec Issue Plant Genomics Bioinforma 21:100128
Odipio J, Alicai T, Ingelbrecht I, Nusinow DA, Bart R, Taylor NJ (2017) Efficient CRISPR/Cas9 genome editing of Phytoene desaturase in cassava. Front Plant Sci 8:1780
Omori M, Yamane H, Osakabe K, Osakabe Y, Tao R (2021) Targeted mutagenesis of CENTRORADIALIS using CRISPR/Cas9 system through the improvement of genetic transformation efficiency of tetraploid highbush blueberry. J Hortic Sci Biotechnol 96:153–161
Osakabe Y, Liang Z, Ren C, Nishitani C, Osakabe K, Wada M, Komori S, Malnoy M, Velasco R, Poli M et al (2018) CRISPR–Cas9-mediated genome editing in apple and grapevine. Nat Protoc 13:2844–2863
Osmani Z, Jin S, Mikami M, Endo M, Atarashi H, Fujino K, Yamada T, Nakahara KS (2019) CRISPR/Cas9-mediated editing of genes encoding rgs-CaM-like proteins in transgenic potato plants. K Kobayashi, M Nishiguchi, Antivir. Resist. Plants Methods Protoc. Springer New York, New York, NY, pp 153–165
OSTP (1986) Coordinated framework for regulation of biotechnology. https://www.aphis.usda.gov/brs/fedregister/coordinated_framework.pdf
Oz TM, Karan R, Merotto A, Altpeter F (2017) Precision nucleotide substitutions in sugarcane following CRISPR/Cas9 and template mediated genome editing
Peng A, Chen S, Lei T, Xu L, He Y, Wu L, Yao L, Zou X (2017) Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene CsLOB1 promoter in citrus. Plant Biotechnol J 15:1509–1519
Perdue S (2021) Secretary Perdue Statement on MOU on Animal Biotechnology
Petri C, Alburquerque N, Faize M, Scorza R, Dardick C (2018) Current achievements and future directions in genetic engineering of European plum (Prunus domestica L.). Transgenic Res 27:225–240
Pompili V, Dalla Costa L, Piazza S, Pindo M, Malnoy M (2020) Reduced fire blight susceptibility in apple cultivars using a high-efficiency CRISPR/Cas9-FLP/FRT-based gene editing system. Plant Biotechnol J 18:845–858
Poovaiah C, Phillips L, Geddes B, Reeves C, Sorieul M, Thorlby G (2020) Genome editing with CRISPR/Cas9 in Pinus Radiata (D. Don).
Qin S, Fan C, Li X, Li Y, Hu J, Li C, Luo K (2020) LACCASE14 is required for the deposition of guaiacyl lignin and affects cell wall digestibility in poplar. Biotechnol Biofuels 13:197
Ramos-Sánchez JM, Triozzi PM, Alique D, Geng F, Gao M, Jaeger KE, Wigge PA, Allona I, Perales M (2019) LHY2 integrates night-length information to determine timing of poplar photoperiodic growth. Curr Biol:29, 2402–2406.e4
Razzaq A, Saleem F, Kanwal M, Mustafa G, Yousaf S, Imran Arshad HM, Hameed MK, Khan MS, Joyia FA (2019) Modern trends in plant genome editing: an inclusive review of the CRISPR/Cas9 toolbox. Int J Mol Sci 20:4045
Ren C, Guo Y, Gathunga EK, Duan W, Li S, Liang Z (2019) Recovery of the non-functional EGFP-assisted identification of mutants generated by CRISPR/Cas9. Plant Cell Rep 38:1541–1549
Roest S, Gilissen LJW (1989) Plant regeneration from protoplasts: a literature review. Acta Bot Neerlandica 38:1–23
Schaart JG, Krens FA, Pelgrom KTB, Mendes O, Rouwendal GJA (2004) Effective production of marker-free transgenic strawberry plants using inducible site-specific recombination and a bifunctional selectable marker gene. Plant Biotechnol J 2:233–240
Sevestre F, Facon M, Wattebled F, Szydlowski N (2020) Facilitating gene editing in potato: a single-nucleotide polymorphism (SNP) map of the Solanum tuberosum L. cv. Desiree genome. Sci Rep 10:2045
Shao M, Blechl A, Thomson JG (2017) Small serine recombination systems ParA-MRS and CinH-RS2 perform precise excision of plastid DNA. Plant Biotechnol J 15:1577–1589
Shao X, Wu S, Dou T, Zhu H, Hu C, Huo H, He W, Deng G, Sheng O, Bi F et al (2020) Using CRISPR/Cas9 genome editing system to create MaGA20ox2 gene-modified semi-dwarf banana. Plant Biotechnol J 18:17–19
Soares JM, Weber KC, Qiu W, Stanton D, Mahmoud LM, Wu H, Huyck P, Zale J, Al Jasim K, Grosser JW et al (2020) The vascular targeted citrus FLOWERING LOCUS T3 gene promotes non-inductive early flowering in transgenic Carrizo rootstocks and grafted juvenile scions. Sci Rep 10:21404
Song G, Walworth A, Lin T, Chen Q, Han X, Irina Zaharia L, Zhong G (2019) VcFT-induced mobile florigenic signals in transgenic and transgrafted blueberries. Hortic Res 6:105
Srinivasan C, Dardick C, Callahan A, Scorza R (2012) Plum (Prunus domestica) trees transformed with poplar FT1 result in altered architecture, dormancy requirement, and continuous flowering. PLoS One 7:e40715–e40715
Steinberg J, Vittorio A (2019) ‘Frankenfish’ salmon can be overseen by FDA despite concerns
Strauss SH (2003) Genomics, genetic engineering, and domestication of crops. Science 300:61
Strauss SH, Tan H, Boerjan W, Sedjo R (2009) Strangled at birth? Forest biotech and the Convention on Biological Diversity. Nat Biotechnol 27:519–527
Sunitha S, Rock CD (2020) CRISPR/Cas9-mediated targeted mutagenesis of TAS4 and MYBA7 loci in grapevine rootstock 101-14. Transgenic Res 29:355–367
The Cartagena Protocol on Biosafety to the Convention on Biological Diversity (2021) https://bch.cbd.int/protocol
Timerbaev V, Mitiouchkina T, Pushin A, Dolgov S (2019) Production of marker-free apple plants expressing the supersweet protein gene driven by plant promoter. Front Plant Sci 10:388
Tränkner C, Lehmann S, Hoenicka H, Hanke M-V, Fladung M, Lenhardt D, Dunemann F, Gau A, Schlangen K, Malnoy M et al (2010) Over-expression of an FT-homologous gene of apple induces early flowering in annual and perennial plants. Planta 232:1309–1324
Tripathi JN, Ntui VO, Ron M, Muiruri SK, Britt A, Tripathi L (2019) CRISPR/Cas9 editing of endogenous banana streak virus in the B genome of Musa spp. overcomes a major challenge in banana breeding. Commun Biol 2:46
Tsai C-J, Xu P, Xue L-J, Hu H, Nyamdari B, Naran R, Zhou X, Goeminne G, Gao R, Gjersing E et al (2020) Compensatory guaiacyl lignin biosynthesis at the expense of syringyl lignin in 4CL1-knockout poplar. Plant Physiol 183:123
Tuncel A, Corbin KR, Ahn-Jarvis J, Harris S, Hawkins E, Smedley MA, Harwood W, Warren FJ, Patron NJ, Smith AM (2019) Cas9-mediated mutagenesis of potato starch-branching enzymes generates a range of tuber starch phenotypes. Plant Biotechnol J 17:2259–2271
Tuteja N, Verma S, Sahoo RK, Raveendar S, Reddy INBL (2012) Recent advances in development of marker-free transgenic plants: regulation and biosafety concern. J Biosci 37:167–197
Van Eenennaam AL, Muir WM (2011) Transgenic salmon: a final leap to the grocery shelf? Nat Biotechnol 29:706–710
Van Eenennaam AL, Wells KD, Murray JD (2019) Proposed U.S. regulation of gene-edited food animals is not fit for purpose. Npj Sci Food 3:3
van Kregten M, de Pater S, Romeijn R, van Schendel R, Hooykaas PJJ, Tijsterman M (2016) T-DNA integration in plants results from polymerase-θ-mediated DNA repair. Nat Plants 2:16164
van Kregten M, Lindhout BI, Hooykaas PJJ, van der Zaal BJ (2009) Agrobacterium-mediated T-DNA transfer and integration by minimal VirD2 consisting of the relaxase domain and a type IV secretion system translocation signal. Mol Plant-Microbe Interactions® 22:1356–1365
van Zeijl A, Wardhani TAK, Seifi Kalhor M, Rutten L, Bu F, Hartog M, Linders S, Fedorova EE, Bisseling T, Kohlen W et al (2018) CRISPR/Cas9-mediated mutagenesis of four putative symbiosis genes of the tropical tree Parasponia andersonii reveals novel phenotypes. Front Plant Sci 9:284
Varkonyi-Gasic E, Wang T, Cooney J, Jeon S, Voogd C, Douglas MJ, Pilkington SM, Akagi T, Allan AC (2021) Shy Girl, a kiwifruit suppressor of feminization, restricts gynoecium development via regulation of cytokinin metabolism and signalling. New Phytol
Varkonyi-Gasic E, Wang T, Voogd C, Jeon S, Drummond RSM, Gleave AP, Allan AC (2019) Mutagenesis of kiwifruit CENTRORADIALIS-like genes transforms a climbing woody perennial with long juvenility and axillary flowering into a compact plant with rapid terminal flowering. Plant Biotechnol J 17:869–880
Veillet F, Perrot L, Chauvin L, Kermarrec M-P, Guyon-Debast A, Chauvin J-E, Nogué F, Mazier M (2019) Transgene-free genome editing in tomato and potato plants using Agrobacterium-mediated delivery of a CRISPR/Cas9 cytidine base editor. Int J Mol Sci
Veillet F, Kermarrec M-P, Chauvin L, Chauvin J-E, Nogué F (2020a) CRISPR-induced indels and base editing using the Staphylococcus aureus Cas9 in potato. PLoS One 15:e0235942
Veillet F, Kermarrec M-P, Chauvin L, Guyon-Debast A, Chauvin J-E, Gallois J-L, Nogué F (2020b) Prime editing is achievable in the tetraploid potato, but needs improvement. bioRxiv 2020.06.18.159111
Veley KM, Okwuonu I, Jensen G, Yoder M, Taylor NJ, Meyers BC, Bart RS (2021) Gene-tagging via CRISPR-mediated homology-directed repair in cassava. G3 Genes Genomes Genetics.
Vondras AM, Minio A, Blanco-Ulate B, Figueroa-Balderas R, Penn MA, Zhou Y, Seymour D, Ye Z, Liang D, Espinoza LK et al (2019) The genomic diversification of grapevine clones. BMC Genomics 20:972
Walter C, Fladung M, Boerjan W (2010) The 20-year environmental safety record of GM trees. Nat Biotechnol 28:656–658
Wan S, Li C, Ma X, Luo K (2017) PtrMYB57 contributes to the negative regulation of anthocyanin and proanthocyanidin biosynthesis in poplar. Plant Cell Rep 36:1263–1276
Wang H, Wu Y, Zhang Y, Yang J, Fan W, Zhang H, Zhao S, Yuan L, Zhang P (2019) CRISPR/Cas9-based mutagenesis of starch biosynthetic genes in sweet potato (Ipomoea Batatas) for the Improvement of Starch Quality. Int J Mol Sci 20:4702
Wang J, Wu H, Chen Y, Yin T (2020) Efficient CRISPR/Cas9-mediated gene editing in an interspecific hybrid poplar with a highly heterozygous genome. Front Plant Sci 11:996
Wang L, Ran L, Hou Y, Tian Q, Li C, Liu R, Fan D, Luo K (2017) The transcription factor MYB115 contributes to the regulation of proanthocyanidin biosynthesis and enhances fungal resistance in poplar. New Phytol 215:351–367
Wang X, Tu M, Wang D, Liu J, Li Y, Li Z, Wang Y, Wang X (2018a) CRISPR/Cas9-mediated efficient targeted mutagenesis in grape in the first generation. Plant Biotechnol J 16:844–855
Wang Y, Chen B, Hu Y, Li J, Lin Z (2005) Inducible excision of selectable marker gene from transgenic plants by the Cre/lox site-specific recombination system. Transgenic Res 14:605–614
Wang Y, Yau Y-Y, Perkins-Balding D, Thomson JG (2011) Recombinase technology: applications and possibilities. Plant Cell Rep 30:267–285
Wang Z, Wang S, Li D, Zhang Q, Li L, Zhong C, Liu Y, Huang H (2018b) Optimized paired-sgRNA/Cas9 cloning and expression cassette triggers high-efficiency multiplex genome editing in kiwifruit. Plant Biotechnol J 16:1424–1433
Wenzel S, Flachowsky H, Hanke M-V (2013) The fast-track breeding approach can be improved by heat-induced expression of the FLOWERING LOCUS T genes from poplar (Populus trichocarpa) in apple (Malus × domestica Borkh.). Plant Cell Tissue Organ Cult PCTOC.
Wilson FM, Harrison K, Armitage AD, Simkin AJ, Harrison RJ (2019) CRISPR/Cas9-mediated mutagenesis of phytoene desaturase in diploid and octoploid strawberry. Plant Methods 15:45
Wolt JD, Wang K, Yang B (2016) The regulatory status of genome-edited crops. Plant Biotechnol J 14:510–518
WTO (2018) International statement on agricultural applications of precision biotechnology. https://docs.wto.org/dol2fe/Pages/FE_Search/FE_S_S009-DP.aspx?language=E&CatalogueIdList=249321
Wu GA, Terol J, Ibanez V, López-García A, Pérez-Román E, Borredá C, Domingo C, Tadeo FR, Carbonell-Caballero J, Alonso R et al (2018) Genomics of the origin and evolution of Citrus. Nature 554:311–316
Wu S, Zhu H, Liu J, Yang Q, Shao X, Bi F, Hu C, Huo H, Chen K, Yi G (2020) Establishment of a PEG-mediated protoplast transformation system based on DNA and CRISPR/Cas9 ribonucleoprotein complexes for banana. BMC Plant Biol 20:425
Xing S, Chen K, Zhu H, Zhang R, Zhang H, Li B, Gao C (2020) Fine-tuning sugar content in strawberry. Genome Biol 21:230
Xing S, Jia M, Wei L, Mao W, Abbasi UA, Zhao Y, Chen Y, Cao M, Zhang K, Dai Z et al (2018) CRISPR/Cas9-introduced single and multiple mutagenesis in strawberry. J Genet Genomics 45:685–687
Yang L, Zhao X, Ran L, Li C, Fan D, Luo K (2017) PtoMYB156 is involved in negative regulation of phenylpropanoid metabolism and secondary cell wall biosynthesis during wood formation in poplar. Sci Rep 7:41209
Yasumoto S, Umemoto N, Lee HJ, Nakayasu M, Sawai S, Sakuma T, Yamamoto T, Mizutani M, Saito K, Muranaka T (2019) Efficient genome engineering using platinum TALEN in potato. Plant Biotechnol 36:167–173
Ye J, Geng Y, Zhang B, Mao H, Qu J, Chua N-H (2014) The Jatropha FT ortholog is a systemic signal regulating growth and flowering time. Biotechnol Biofuels 7:91
Young J, Zastrow-Hayes G, Deschamps S, Svitashev S, Zaremba M, Acharya A, Paulraj S, Peterson-Burch B, Schwartz C, Djukanovic V et al (2019) CRISPR-Cas9 editing in maize: systematic evaluation of off-target activity and its relevance in crop improvement. Sci Rep 9:6729
Zhang H, Harry DE, Ma C, Yuceer C, Hsu C-Y, Vikram V, Shevchenko O, Etherington E, Strauss SH (2010) Precocious flowering in trees: the FLOWERING LOCUS T gene as a research and breeding tool in Populus. J Exp Bot 61:2549–2560
Zhou J, Wang G, Liu Z (2018) Efficient genome editing of wild strawberry genes, vector development and validation. Plant Biotechnol J 16:1868–1877
Zhou H, Bai S, Wang N, Sun X, Zhang Y, Zhu J, Dong C (2020a) CRISPR/Cas9-mediated mutagenesis of MdCNGC2 in apple callus and VIGS-mediated silencing of MdCNGC2 in fruits improve resistance to Botryosphaeria dothidea. Front Plant Sci 11:575477–575477
Zhou J, Sittmann J, Guo L, Xiao Y, Huang X, Pulapaka A, Liu Z (2020b) Gibberellin and auxin signaling genes RGA1 and ARF8 repress accessory fruit initiation in diploid strawberry. Plant Physiol
Zhu C, Zheng X, Huang Y, Ye J, Chen P, Zhang C, Zhao F, Xie Z, Zhang S, Wang N et al (2019) Genome sequencing and CRISPR/Cas9 gene editing of an early flowering Mini-Citrus (Fortunella hindsii). Plant Biotechnol J 17:2199–2210
Zou X, Fan D, Peng A, He Y, Xu L, Lei T, Yao L, Li Q, Luo K (2019) CRISPR/Cas9-mediated editing of multiple sites in the citrus CsLOB1 promoter. Acta Hortic Sin 46:337–344
Author information
Authors and Affiliations
Corresponding author
Additional information
Editor: David Songstad
Supplementary Information
ESM 1
(DOCX 106 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Goralogia, G.S., Redick, T.P. & Strauss, S.H. Gene editing in tree and clonal crops: progress and challenges. In Vitro Cell.Dev.Biol.-Plant 57, 683–699 (2021). https://doi.org/10.1007/s11627-021-10197-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11627-021-10197-x