Abstract
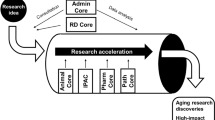
The USC-Buck Nathan Shock Center of Excellence in the Biology of Aging is a new and fully integrated multi-institutional center focused on training the next generation of geroscientists and providing access to cutting-edge geroscience technologies to investigators across the nation. The USC-Buck NSC is devoted to forging a deeper understanding of how and why aging processes cause disease in order to advance the translation of basic research on aging into effective preventions and therapies. Including more than 61 NIA-supported investigators, six NIA-funded research centers, four NIA T32s, and several additional aging research centers of excellence, the USC-Buck NSC constitutes one of the largest collections of leaders in geroscience research within the USA; the unique nature of the USC-Buck NSC research infrastructure ensures an integrated organization that is representative of the wide breadth of topics encompassed by the biology of aging field. By leveraging the 25-year-long relationship, current collaborations and joint administrational activities of the University of Southern California and the Buck Institute for Aging Research, the USC-Buck NSC aims to enhance and expand promising research in the biology of aging at both at the and to make a positive impact across California, the nation and throughout the world. Specialized cores provide services to all Shock Center members, as well as provide support for services to the community at large.

Similar content being viewed by others
References
Miller B, et al. Human population genetics in aging studies for molecular biologists. In: aging. Springer; 2020. p. 67–76.
Arpawong TE, et al. Effects of recent stress and variation in the serotonin transporter polymorphism (5-HTTLPR) on depressive symptoms: a repeated-measures study of adults age 50 and older. Behav Genet. 2016;46(1):72–88.
Arpawong TE, et al. Genetic variant specific to aging-related verbal memory: insights from GWASs in a population-based cohort. PloS one. 2017;12(8):e0182448.
Levine ME, et al. A polygenic risk score associated with measures of depressive symptoms among older adults. Biodemography Soc Biol. 2014;60(2):199–211.
Mekli K, et al. Frailty Index associates with GRIN2B in two representative samples from the United States and the United Kingdom. PLoS one. 2018;13(11):e0207824.
Wiley CD, Campisi J. From ancient pathways to aging cells-connecting metabolism and cellular senescence. Cell Metab. 2016;23(6):1013–21.
Childs BG, et al. Senescence and apoptosis: dueling or complementary cell fates? EMBO Rep. 2014;15(11):1139–53.
Wiley CD, et al. SILAC analysis reveals increased secretion of hemostasis-related factors by senescent cells. Cell Rep. 2019;28(13):3329-3337 e5.
Hodes RJ, et al. Disease drivers of aging. Ann N Y Acad Sci. 2016;1386(1):45–68.
Campisi J, et al. From discoveries in ageing research to therapeutics for healthy ageing. Nature. 2019;571(7764):183–92.
Larose H, et al. Gametogenesis: a journey from inception to conception. Curr Top Dev Biol. 2019;132:257–310.
Lei L, Spradling AC. Mouse oocytes differentiate through organelle enrichment from sister cyst germ cells. Science. 2016;352(6281):95–9.
Mookerjee SA, et al. Plate-based measurement of respiration by isolated mitochondria. Methods Mol Biol. 2018;1782:301–13.
Qin Q, et al. Chronic treatment with the mitochondrial peptide humanin prevents age-related myocardial fibrosis in mice. Am J Physiol Heart Circ Physiol. 2018;315(5):H1127–36.
Lee C, et al. The mitochondrial-derived peptide MOTS-c promotes metabolic homeostasis and reduces obesity and insulin resistance. Cell Metab. 2015;21(3):443–54.
Kim KH, et al. The mitochondrial-encoded peptide MOTS-c Translocates to the nucleus to regulate nuclear gene expression in response to metabolic stress. Cell Metab. 2018;28(3):516-524 e7.
Benayoun BA, et al. Remodeling of epigenome and transcriptome landscapes with aging in mice reveals widespread induction of inflammatory responses. Genome Res. 2019;29(4):697–709.
Woodward NC, et al. Exposure to nanoscale particulate matter from gestation to adulthood impairs metabolic homeostasis in mice. Sci Rep. 2019;9(1):1816.
Pomatto LCD, et al. Aging attenuates redox adaptive homeostasis and proteostasis in female mice exposed to traffic-derived nanoparticles ('vehicular smog’). Free Radic Biol Med. 2018;121:86–97.
Flynn JM, et al. Late-life rapamycin treatment reverses age-related heart dysfunction. Aging Cell. 2013;12(5):851–62.
Demaria M, et al. Cellular senescence promotes adverse effects of chemotherapy and cancer relapse. Cancer Discov. 2017;7(2):165–76.
Basisty N, Holtz A, Schilling B. Accumulation of “Old Proteins” and the critical need for ms-based protein turnover measurements in aging and longevity. Proteomics. 2020;20(5–6):e1800403.
Basisty N, et al. Simultaneous quantification of the acetylome and succinylome by “one-pot” affinity enrichment. Proteomics. 2018;18(17):e1800123.
Basisty N, Meyer JG, Schilling B. Protein turnover in aging and longevity. Proteomics. 2018;18(5–6):e1700108.
Furman D, et al. Systems analysis of sex differences reveals an immunosuppressive role for testosterone in the response to influenza vaccination. Proc Natl Acad Sci U S A. 2014;111(2):869–74.
Furman D, et al. Apoptosis and other immune biomarkers predict influenza vaccine responsiveness. Mol Syst Biol. 2013;9:659.
Mirzaei H, Suarez JA, Longo VD. Protein and amino acid restriction, aging and disease: from yeast to humans. Trends Endocrinol Metab. 2014;25(11):558–66.
Lee C, et al. Fasting cycles retard growth of tumors and sensitize a range of cancer cell types to chemotherapy. Sci Transl Med. 2012;4(124):124ra27.
Fabrizio P, Longo VD. The chronological life span of Saccharomyces cerevisiae. Aging Cell. 2003;2(2):73–81.
Fabrizio P, et al. Regulation of longevity and stress resistance by Sch9 in yeast. Science. 2001;292(5515):288–90.
Longo VD. Mutations in signal transduction proteins increase stress resistance and longevity in yeast, nematodes, fruit flies, and mammalian neuronal cells. Neurobiol Aging. 1999;20(5):479–86.
Steffen KK, et al. Yeast life span extension by depletion of 60s ribosomal subunits is mediated by Gcn4. Cell. 2008;133(2):292–302.
Kaeberlein M, Kennedy BK. Protein translation, 2007. Aging Cell. 2007;6(6):731–4.
Powers RW 3rd, et al. Extension of chronological life span in yeast by decreased TOR pathway signaling. Genes Dev. 2006;20(2):174–84.
Kaeberlein M, et al. Regulation of yeast replicative life span by TOR and Sch9 in response to nutrients. Science. 2005;310(5751):1193–6.
Spatola BN, et al. Nuclear and cytoplasmic WDR-23 isoforms mediate differential effects on GEN-1 and SKN-1 substrates. Sci Rep. 2019;9(1):11783.
Lucanic M, et al. Chemical activation of a food deprivation signal extends lifespan. Aging Cell. 2016;15(5):832–41.
Lang S, et al. A conserved role of the insulin-like signaling pathway in diet-dependent uric acid pathologies in Drosophila melanogaster. PLoS Genet. 2019;15(8):e1008318.
Akagi K, et al. Dietary restriction improves intestinal cellular fitness to enhance gut barrier function and lifespan in D. melanogaster. PLoS Genet. 2018;14(11):e1007777.
Nelson CS, et al. Cross-phenotype association tests uncover genes mediating nutrient response in Drosophila. BMC Genomics. 2016;17(1):867.
Vaupel JW, Johnson TE, Lithgow GJ. Rates of mortality in populations of Caenorhabditis elegans. Science. 1994;266(5186):826 (author reply 828).
Brooks A, Lithgow GJ, Johnson TE. Mortality rates in a genetically heterogeneous population of Caenorhabditis elegans. Science. 1994;263(5147):668–71.
Lithgow GJ, Kirkwood TB. Mechanisms and evolution of aging. Science. 1996;273(5271):80.
Babar P, et al. P13-kinase inhibition induces dauer formation, thermotolerance and longevity in C. elegans. Neurobiol Aging. 1999;20(5):513–9.
Melov S, et al. Extension of life-span with superoxide dismutase/catalase mimetics. Science. 2000;289(5484):1567–9.
Walker DW, et al. Evolution of lifespan in C. elegans. Nature. 2000;405(6784):296–7.
Barsyte D, Lovejoy DA, Lithgow GJ. Longevity and heavy metal resistance in daf-2 and age-1 long-lived mutants of Caenorhabditis elegans. FASEB J. 2001;15(3):627–34.
Sampayo JN, Olsen A, Lithgow GJ. Oxidative stress in Caenorhabditis elegans: protective effects of superoxide dismutase/catalase mimetics. Aging Cell. 2003;2(6):319–26.
McColl G, Vantipalli MC, Lithgow GJ. The C. elegans ortholog of mammalian Ku70, interacts with insulin-like signaling to modulate stress resistance and life span. Faseb J. 2005;19(12):1716–8.
Olsen A, Vantipalli MC, Lithgow GJ. Using Caenorhabditis elegans as a model for aging and age-related diseases. Ann N Y Acad Sci. 2006;1067:120–8.
Fisher AL, Lithgow GJ. The nuclear hormone receptor DAF-12 has opposing effects on Caenorhabditis elegans lifespan and regulates genes repressed in multiple long-lived worms. Aging Cell. 2006;5(2):127–38.
Pan KZ, et al. Inhibition of mRNA translation extends lifespan in Caenorhabditis elegans. Aging Cell. 2007;6(1):111–9.
Rogers AN, et al. Life span extension via eIF4G inhibition is mediated by posttranscriptional remodeling of stress response gene expression in C. elegans. Cell Metab. 2011;14(1):55–66.
Reis-Rodrigues P, et al. Proteomic analysis of age-dependent changes in protein solubility identifies genes that modulate lifespan. Aging cell. 2012;11(1):120–7.
Curran S, et al. Defective mitochondrial protein translocation precludes normal Caenorhabditis elegans development. J Biol Chem. 2004;279(52):54655–62.
Curran S, Ruvkun G. Lifespan regulation by evolutionarily conserved genes essential for viability. PLoS Genet. 2007;3(4):e56.
Curran SP, et al. A soma-to-germline transformation in long-lived Caenorhabditis elegans mutants. Nature. 2009;459(7250):1079–84.
Tacutu R, et al. Prediction of C. elegans longevity genes by human and worm longevity networks. PloS one. 2012;7(10):e48282.
Pang S, Curran SP. Adaptive capacity to bacterial diet modulates aging in C. elegans. Cell metabolism. 2014;19(2):221–31.
Pang S, et al. SKN-1 and Nrf2 couples proline catabolism with lipid metabolism during nutrient deprivation. Nat Commun. 2014;5:5048.
Khanna A, Johnson DL, Curran SP. Physiological roles for mafr-1 in reproduction and lipid homeostasis. Cell Rep. 2014;9(6):2180–91.
Lynn DA, et al. Omega-3 and -6 fatty acids allocate somatic and germline lipids to ensure fitness during nutrient and oxidative stress in Caenorhabditis elegans. Proc Natl Acad Sci U S A. 2015;112(50):15378–83.
Khanna A, Pradhan A, Curran SP. Emerging roles for Maf1 beyond the regulation of RNA polymerase III activity. J Mol Biol. 2015;427(16):2577–85.
Yen CA, Curran SP. Gene-diet interactions and aging in C. elegans. Exp Gerontol. 2016;86:106–12.
Lo JY, Spatola BN, Curran SP. WDR23 regulates NRF2 independently of KEAP1. PLoS Genet. 2017;13(4):e1006762.
Escorcia W, Ruter DL, Nhan J, Curran SP. Quantification of lipid abundance and evaluation of lipid distribution in Caenorhabditis elegans by Nile Red and Oil Red O staining. J Vis Exp. 2018;(133):57352.
Dalton HM, Curran SP. Hypodermal responses to protein synthesis inhibition induce systemic developmental arrest and AMPK-dependent survival in Caenorhabditis elegans. PLoS Genet. 2018;14(7):e1007520.
Haghani A, et al. Air pollution alters Caenorhabditis elegans development and lifespan: responses to traffic-related nanoparticulate matter. J Gerontol A Biol Sci Med Sci. 2019;74(8):1189–97.
Tower J. Transgenic methods for increasing Drosophila life span. Mech Ageing Dev. 2000;118(1–2):1–14.
Pomatto LCD, et al. The mitochondrial lon protease is required for age-specific and sex-specific adaptation to oxidative stress. Curr Biol. 2017;27(1):1–15.
Kapahi P, et al. Regulation of lifespan in Drosophila by modulation of genes in the TOR signaling pathway. Curr Biol. 2004;14(10):885–90.
Levine ME, et al. Low protein intake is associated with a major reduction in IGF-1, cancer, and overall mortality in the 65 and younger but not older population. Cell Metab. 2014;19(3):407–17.
Safdie F, et al. Fasting enhances the response of glioma to chemo- and radiotherapy. PloS one. 2012;7(9):e44603.
Brandhorst S, et al. Short-term calorie and protein restriction provide partial protection from chemotoxicity but do not delay glioma progression. Experimental gerontology. 2013;48(10):1120–8.
Parrella E, et al. Protein restriction cycles reduce IGF-1 and phosphorylated Tau, and improve behavioral performance in an Alzheimer’s disease mouse model. Aging Cell. 2013;12(2):257–68.
Yen K, et al. The emerging role of the mitochondrial-derived peptide humanin in stress resistance. J Mol Endocrinol. 2013;50(1):R11–9.
Lee C, Yen K, Cohen P. Humanin: a harbinger of mitochondrial-derived peptides? Trends Endocrinol Metab. 2013;24(5):222–8.
Lee C, Longo V. Dietary restriction with and without caloric restriction for healthy aging. F1000Res. 2016;5:F1000 Faculty Rev-117.
Lui JC, et al. Changes in gene expression associated with aging commonly originate during juvenile growth. Mechanisms of ageing and development. 2010;131(10):641–9.
Ntambi JM, et al. Loss of stearoyl-CoA desaturase-1 function protects mice against adiposity. Proceedings of the National Academy of Sciences of the United States of America. 2002;99(17):11482–6.
Birsoy K, et al. Cellular program controlling the recovery of adipose tissue mass: an in vivo imaging approach. Proceedings of the National Academy of Sciences of the United States of America. 2008;105(35):12985–90.
Li JZ, et al. Chronic overexpression of PNPLA3I148M in mouse liver causes hepatic steatosis. The Journal of clinical investigation. 2012;122(11):4130–44.
Valenzano DR, et al. The African Turquoise killifish genome provides insights into evolution and genetic architecture of lifespan. Cell. 2015;163(6):1539–54.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
About this article
Cite this article
Curran, S.P., Lithgow, G.J., Verdin, E. et al. University of Southern California and buck institute nathan shock center: multidimensional models of aging. GeroScience 43, 2119–2127 (2021). https://doi.org/10.1007/s11357-021-00416-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11357-021-00416-z