Abstract
World Health Organization articulated 9.8 million casualties globally in 2018 due to cancer. Cancer, as the world’s second most fatal disease, can be recuperated well if diagnosed at an early stage. In this work, a gradient-based impedance synthesis of normal and cancerous cells of breast and lungs, is demonstrated numerically for early-stage cancer detection. Low-voltage single-cell level examination is employed for indomitable diagnosis. MCF-7 and MCF-10A are utilized as breast cancer and breast normal cells, respectively; likewise, SK-MES and NL-20 are utilized as lung cancer and lung normal cell. Pre-examination numerical setup validity ensured with multiple test regimes. Micro-scaled planar and nano-structured electrodes are employed individually to witness the effect of the electrode’s structure during electrical impedance examination of cancer and non-cancer cell. Frequency range, at which differential impedance effect is found detectable, for breast and lung cancer cell pairs is determined to be 107 Hz and 108 Hz, respectively. By surpassing the conventional impedance spectroscopy with tedious data fitting formalities, the gradient synthesis technique for cancer detection is introduced. The gradient synthesis for cancer detection is found independent of electrode shape effect. Gradient for breast cancer cell is found to be 2 times greater than the normal breast cell while for lung cancer cell it is found to be 1.5 times greater than the normal lung cell. Our results suggest that as the frequency of applied electrical stimulus increases, impedance of cancerous cell falls at the rate almost double than its counterpart normal cell. This work provides a theoretical basis for further experimental exploration of gradient-based impedance synthesis in cancer therapy and serves as a design tool for performance optimization.
Graphical abstract

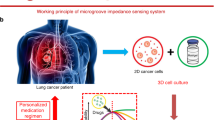
Figure 1 (a) Represents electrical Impedance analysis of breast normal cell MCF-10A and breast cancer cell MCF-7 using micro-scaled planar and nano-structured electrodes. (b) Gradient impedance synthesis performed, for breast normal cell (MCF-10A) and breast cancer cell (MCF-7) likewise for lungs normal cell (NL-20) and lungs cancer cell (SK-MES), which assures clear differential effect for cancer screening.
Surpassing the conventional and tedious data fitting impedance spectroscopies, a novel gradient-based impedance spectroscopy for early cancer detection is introduced. It clearly detects cancer without any data fitting formalities to find parameter of identification. Planar and nano structure electrodes are used to witness the impact of electrode shape on cell impedance. Breast normal MCF-10A and cancer cell MCF-7 as well as lungs normal NL-20 and cancer cell SK-MES are examined to reflect the efficacy of our work. Single cell level examination is performed for authenticated results.








Similar content being viewed by others
References
Elmore JG, Barton MB, Moceri VM et al (1998) Ten-year risk of false positive screening mammograms and clinical breast examinations. N Engl J Med 338:1089–1096
Hong J, Kandasamy K, Marimuthu M et al (2011) Electrical cell-substrate impedance sensing as a non-invasive tool for cancer cell study. Analyst 136:237–245
Vargas AJ, Harris CC (2016) Biomarker development in the precision medicine era: lung cancer as a case study. Nat Rev Cancer 16:525537
Mangé A, Dimitrakopoulos L, Soosaipillai A et al (2016) An integrated cell line-based discovery strategy identified follistatin and kallikrein 6 as serum biomarker candidates of breast carcinoma. J Proteomics 142:114–121. https://doi.org/10.1016/j.jprot.2016.04.050
Mao Y, Shin K-S, Wang X et al (2016) Semiconductor electronic label-free assay for predictive toxicology. Sci Rep 6:1–8
Abercrombie M, Ambrose E (1962) The surface properties of cancer cells: a review. Cancer Res 22:525–548
Schermelleh L, Carlton PM, Haase S et al (2008) Subdiffraction multicolor imaging of the nuclear periphery with 3D structured illumination microscopy. Science 80(320):1332–1336
Hong J, Lan K, Jang L (2012) Electrical characteristics analysis of various cancer cells using a microfluidic device based on single-cell impedance measurement. Sensors Actuators B Chem 173:927–934. https://doi.org/10.1016/j.snb.2012.06.046
Mansor MA, Ahmad MR (2015) Single cell electrical characterization techniques. 12686–12712. https://doi.org/10.3390/ijms160612686
Scharfetter H (1999) Structural modeling for impedance-based non-invasive diagnostic methods. Thesis for the habilitation at the faculty of Electrical Engineering, Technical University Graz.Graz
Ren D, Chui CO (2018) Feasibility of tracking multiple single-cell properties with impedance spectroscopy. ACS sensors 3:1005–1015
Wang M-H, Chang W-H (2015) Effect of electrode shape on impedance of single HeLa cell: a COMSOL simulation. Biomed Res Int, pp 1–9
Nanofiber C, Arrays E, Electrochemical U et al (2010) Characterization of carbon nanofiber electrode arrays using electrochemical impedance spectroscopy: effect of scaling down electrode size. ACS Nano 4:955–961
Riaz K, Zhao C, Lau TS, et al. (2015) Low-cost Nano-spike Bio-Impedance Sensor (NBIS) without surface functionalization for detection and phenotyping of cancer cells. In: 2015 Transducers-2015 18th International Conference on Solid-State Sensors, Actuators and Microsystems (TRANSDUCERS). IEEE, pp 367–370
Forbes RG, Edgcombe CJ, Valdre U (2003) Some comments on models for field enhancement. Ultramicroscopy 95:57–65
He H, Chang DC, Lee Y-K (2007) Using a micro electroporation chip to determine the optimal physical parameters in the uptake of biomolecules in HeLa cells. Bioelectrochemistry 70:363–368
Cho YH, Yamamoto T, Sakai Y et al (2006) Development of microfluidic device for electrical/physical characterization of single cell. J Microelectromechanical Syst 15:287–295
Riaz K, Leung S-F, Fan Z, Lee Y-K (2017) Electric field enhanced 3D scalable low-voltage nano-spike electroporation system. Sensors Actuators A Phys 255:10–20. https://doi.org/10.1016/j.sna.2016.12.022
Movahed S, Li D (2011) Microfluidics cell electroporation. Microfluid Nanofluid 10:703–734
Lee WG, Demirci U, Khademhosseini A (2009) Microscale electroporation: challenges and perspectives for clinical applications. Integr Biol 1:242–251
Geng T, Lu C (2013) Microfluidic electroporation for cellular analysis and delivery. Lab Chip 13:3803–3821
Potter H, Heller R (2018) Transfection by electroporation. Curr Protoc Mol Biol 121:3–9
Giaever I, Keese CR (1993) A morphological biosensor for mammalian cells. Nature 366:591–592
Jang L-S, Wang M-H (2007) Microfluidic device for cell capture and impedance measurement. Biomed Microdevices 9:737–743
Wang M-H, Jang L-S (2009) A systematic investigation into the electrical properties of single HeLa cells via impedance measurements and COMSOL simulations. Biosens Bioelectron 24:2830–2835
Tabasi A, Noorbakhsh A, Sharifi E (2017) Reduced graphene oxide-chitosan-aptamer interface as new platform for ultrasensitive detection of human epidermal growth factor receptor 2. Biosens Bioelectron. https://doi.org/10.1016/j.bios.2017.04.020
Egot-Lemaire S, Pijanka J, Sulé-Suso J, Semenov S (2009) Dielectric spectroscopy of normal and malignant human lung cells at ultra-high frequencies. Phys Med Biol 54:2341
An J, Lee J, Lee SH et al (2009) Separation of malignant human breast cancer epithelial cells from healthy epithelial cells using an advanced dielectrophoresis-activated cell sorter (DACS). Anal Bioanal Chem 394:801–809. https://doi.org/10.1007/s00216-009-2743-7
Nasir N, Al Ahmad M (2020) Cells electrical characterization: dielectric properties, mixture, and modeling theories. J Eng, https://doi.org/10.1155/2020/9475490
Guido I, Jaeger MS, Duschl C (2011) Dielectrophoretic stretching of cells allows for characterization of their mechanical properties. Eur Biophys J 40:281–288. https://doi.org/10.1007/s00249-010-0646-3
Hamaguchi K, Geiduschek EP (1962) The effect of electrolytes on the stability of the deoxyribonucleate helix. J Am Chem Soc 84:1329–1338. https://doi.org/10.1021/ja00867a001
RSC (2020) Phosphate-buffered saline solution. In: R. Soc. Chem. https://www.rsc.org/publishing/journals/prospect/ontology.asp?id=CMO:0001751&MSID=B103933B. Accessed 22 Jan 2021
Niemtzow RC (1985) Transmembrane potentials & characters immune & tumor cell. CRC-Press Inc., ed.1. ch.1:7–20, Boca Raton, Florida
Blad B, Baldetorp B (1996) Impedance spectra of tumour tissue in comparison with normal tissue; a possible clinical application for electrical impedance tomography. Physiol Meas 17:A105–A115
Stern RG, Milestone BN, Gatenby RA (1999) Carcinogenesis and the plasma membrane. Med Hypotheses 52:367–372
Martinsen OG, Grimnes S (2014) Bioimpedance and bioelectricity basics, ed. 3. ch.3 Dielectrics. Academic press, Cambridge
Haltiwanger S (2003) The electrical properties of cancer cells. In: The Rife 2003 International Conference in Seattle, Washington
Brown GC (1999) The energy of life: the science of what makes our minds and bodies work. ch.3–4, ppp 15–28. Free Press, New York
Ho M-W (1996) Bioenergetics and biocommunication. Comput Cell Mol Biol Syst 251–264
Aslam MA, Riaz K, Mahmood MQ, Zubair M (2019) Hybrid analytical-numerical approach for investigation of differential effects in normal and cancer cells under electroporation. RSC Adv 9:41518–41530
Grossi M, Riccò B (2017) Electrical impedance spectroscopy (EIS) for biological analysis and food characterization : a review 6:303–325
Randviir EP, Banks CE (2013) Electrochemical impedance spectroscopy: an overview of bioanalytical applications. Anal Methods 5:1098–1115. https://doi.org/10.1039/C3AY26476A
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Aslam, M.A., Riaz, K. & Saleem, M.M. Gradient-based impedance synthesis for breast and lung cancer cell screening deploying planar and nano-structured electrodes. Med Biol Eng Comput 59, 1709–1721 (2021). https://doi.org/10.1007/s11517-021-02382-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-021-02382-2