Abstract
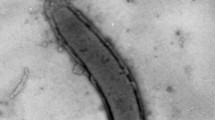
The diversity of anaerobic microorganisms in terrestrial mud volcanoes is largely unexplored. Here we report the isolation of a novel sulfate-reducing alkaliphilic bacterium (strain F-1T) from a terrestrial mud volcano located at the Taman peninsula, Russia. Cells of strain F-1T were Gram-negative motile vibrios with a single polar flagellum; 2.0–4.0 µm in length and 0.5 µm in diameter. The temperature range for growth was 6–37 °C, with an optimum at 24 °C. The pH range for growth was 7.0–10.5, with an optimum at pH 9.5. Strain F-1T utilized lactate, pyruvate, and molecular hydrogen as electron donors and sulfate, sulfite, thiosulfate, elemental sulfur, fumarate or arsenate as electron acceptors. In the presence of sulfate, the end products of lactate oxidation were acetate, H2S and CO2. Lactate and pyruvate could also be fermented. The major product of lactate fermentation was acetate. The main cellular fatty acids were anteiso-C15:0, C16:0, C18:0, and iso-C17:1ω8. Phylogenetic analysis revealed that strain F-1T was most closely related to Pseudodesulfovibrio aespoeensis (98.05% similarity). The total size of the genome of the novel isolate was 3.23 Mb and the genomic DNA G + C content was 61.93 mol%. The genome contained all genes essential for dissimilatory sulfate reduction. We propose to assign strain F-1T to the genus Pseudodesulfovibrio, as a new species, Pseudodesulfovibrio alkaliphilus sp. nov. The type strain is F-1T (= KCTC 15918T = VKM B-3405T).


Similar content being viewed by others
Data availability
The GenBank/EMBL accession number of 16S rRNA gene sequence of the strain F-1T is MN601397, the Whole Genome Shotgun project has been deposited at DDBJ/ENA/GenBank under the accession WODC00000000.
References
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Bale SJ, Goodman K, Rochelle PA et al (1997) Desulfovibrio profundus sp. nov., a novel barophilic sulfate-reducing bacterium from deep sediment layers in the Japan sea. Int J Syst Bacteriol 47:515–521. https://doi.org/10.1099/00207713-47-2-515
Bell E, Lamminmäki T, Alneberg J et al (2018) Biogeochemical cycling by a low-diversity microbial community in deep groundwater. Front Microbiol 9:2129. https://doi.org/10.3389/fmicb.2018.02129
Benson DA, Boguski MS, Lipman DJ et al (1999) GenBank. Nucleic Acids Res 27:12–17. https://doi.org/10.1093/nar/27.1.12
Beveridge TJ, Lawrence JR, Murray RGE (2014) Sampling and staining for light microscopy. In: Methods for general and molecular microbiology. Ed. 3. Ch. 2. pp 19–33 https://doi.org/10.1128/9781555817497.ch2
Bonch-Osmolovskaya EA, Sokolova TG, Kostrikina NA et al (1990) Desulfurella acetivorans gen. nov. and sp. nov. -a new thermophilic sulfur-reducing eubacterium. Arch Microbiol 153:151–155. https://doi.org/10.1007/BF00247813
Brettin T, Davis JJ, Disz T et al (2015) RASTtk: A modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep 5:8365. https://doi.org/10.1038/srep08365
Cao J, Gayet N, Zeng X et al (2016) Pseudodesulfovibrio indicus gen. nov., sp. nov., a piezophilic sulfate-reducing bacterium from the Indian ocean and reclassification of four species of the genus Desulfovibrio. Int J Syst Evol Microbiol 66:3904–3911. https://doi.org/10.1099/ijsem.0.001286
Caumette P, Cohen Y, Matheron R (1991) Isolation and Characterization of Desulfovibrio halophilus sp. nov., a halophilic sulfate-reducing bacterium isolated from solar lake (Sinai). Syst Appl Microbiol 14:33–38. https://doi.org/10.1016/S0723-2020(11)80358-9
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the Bootstrap. Evolution (n Y) 39(4):783–791. https://doi.org/10.2307/2408678
Fichtel K, Mathes F, Könneke M et al (2012) Isolation of sulfate-reducing bacteria from sediments above the deep-subseafloor aquifer. Front Microbiol 3:65. https://doi.org/10.3389/fmicb.2012.00065
Gam ZBA, Oueslati R, Abdelkafi S et al (2009) Desulfovibrio tunisiensis sp. nov., a novel weakly halotolerant, sulfate-reducing bacterium isolated from exhaust water of a Tunisian oil refinery. Int J Syst Evol Microbiol 59:1059–1063. https://doi.org/10.1099/ijs.0.000943-0
Gilmour CC, Soren AB, Gionfriddo CM et al (2019) Pseudodesulfovibrio mercurii sp. nov., a mercury-methylating bacterium isolated from sediment. Int J Syst Evol Microbiol 71:3. https://doi.org/10.1099/ijsem.0.004697
Hazkani-Covo E, Graur D (2007) A comparative analysis of numt evolution in human and chimpanzee. Mol Biol Evol 24:13–18. https://doi.org/10.1093/molbev/msl149
Khelaifia S, Fardeau ML, Pradel N et al (2011) Desulfovibrio piezophilus sp. nov., a piezophilic, sulfate-reducing bacterium isolated from wood falls in the Mediterranean Sea. Int J Syst Evol Microbiol 61:2706–2711. https://doi.org/10.1099/ijs.0.028670-0
Krekeler D, Sigalevich P, Teske A et al (1997) A sulfate-reducing bacterium from the oxic layer of a microbial mat from Solar Lake (Sinai) Desulfovibrio Oxyclinae Sp Nov. Arch Microbiol 167:369–375. https://doi.org/10.1007/s002030050457
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Mall A, Sobotta J, Huber C et al (2018) Reversibility of citrate synthase allows autotrophic growth of a thermophilic bacterium. Science 359:563–567. https://doi.org/10.1126/science.aao2410
Mazzini A, Etiope G (2017) Mud volcanism: An updated review. Earth-Science Rev 168:81–112. https://doi.org/10.1016/j.earscirev.2017.03
Meier-Kolthoff JP, Auch AF, Klenk HP et al (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics 14:60. https://doi.org/10.1186/1471-2105-14-60
Motamedi M, Pedersen K (1998) Desulfovibrio aespoeensis sp. nov., a mesophilic sulfate-reducing bacterium from deep groundwater at Aspo hard rock laboratory. Sweden Int J Syst Bacteriol 48:311–315. https://doi.org/10.1099/00207713-48-1-311
Nunoura T, Chikaraishi Y, Izaki R et al (2018) A primordial and reversible TCA cycle in a facultatively chemolithoautotrophic thermophile. Science 359:559–563. https://doi.org/10.1126/science.aao3407
Oren A, Garrity GM (2021) Validation list no. 197. List of new names and new combinations previously effectively, but not validly, published. Int J Syst Evol Microbiol 71:1–6
Overbeek R, Olson R, Pusch GD et al (2014) The SEED and the rapid annotation of microbial genomes using Subsystems technology (RAST). Nucleic Acids Res 42:206–214. https://doi.org/10.1093/nar/gkt1226
Parte AC, Carbasse JS, Meier-Kolthoff JP et al (2020) List of prokaryotic names with standing in nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol 70:5607–5612. https://doi.org/10.1099/ijsem.0.004332
Pereira IAC, Ramos AR, Grein F et al (2011) A comparative genomic analysis of energy metabolism in sulfate reducing bacteria and archaea. Front Microbiol 2:69. https://doi.org/10.3389/fmicb.2011.00069
Rabus R, Venceslau SS, Wöhlbrand L et al (2015) A post-genomic view of the ecophysiology, catabolism and biotechnological relevance of sulphate-reducing prokaryotes. Adv Microb Physiol 66:55–321. https://doi.org/10.1016/bs.ampbs.2015.05.002
Ranchou-Peyruse M, Goñi-Urriza M, Guignard M et al (2018) Pseudodesulfovibrio hydrargyri sp. nov., a mercury-methylating bacterium isolated from a brackish sediment. Int J Syst Evol Microbiol 68:1461–1466. https://doi.org/10.1099/ijsem.0.002173
Rodriguez-R L, Konstantinidis K (2016) The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Prepr. https://doi.org/10.7287/peerj.preprints.1900
Rzhetsky A, Nei M (1992) A simple method for estimating and testing minimum-evolution trees. Mol Biol Evol 9:945–967. https://doi.org/10.1093/oxfordjournals.molbev.a040771
Sánchez-Andrea I, Guedes IA, Hornung B et al (2020) The reductive glycine pathway allows autotrophic growth of Desulfovibrio desulfuricans. Nat Commun 11:5090. https://doi.org/10.1038/s41467-020-18906-7
Slobodkin AI, Tourova TP, Kuznetsov BB et al (1999) Thermoanaerobacter siderophilus sp.nov., a novel dissimilatory Fe(III)-reducing anaerobic thermophilic bacterium. Int J Syst Bacteriol 49:1471–1478. https://doi.org/10.1099/00207713-49-4-1471
Slobodkin AI, Reysenbach AL, Slobodkina GB et al (2012) Thermosulfurimonas dismutans gen. nov., sp. nov., an extremely thermophilic sulfur-disproportionating bacterium from a deep-sea hydrothermal vent. Int J Syst Evol Microbiol 62:2565–2571. https://doi.org/10.1099/ijs.0.034397-0
Slobodkina GB, Merkel AY, Novikov AA et al (2020) Pelomicrobium methylotrophicum gen. nov., sp. nov. a moderately thermophilic, facultatively anaerobic, lithoautotrophic and methylotrophic bacterium isolated from a terrestrial mud volcano. Extremophiles 24:177–185. https://doi.org/10.1007/s00792-019-01145-0
Song Y, Lee JS, Shin J et al (2020) Functional cooperation of the glycine synthasereductase and Wood-Ljungdahl pathways for autotrophic growth of Clostridium drakei. Proc Natl Acad Sci USA 117:7516–7523. https://doi.org/10.1073/pnas.1912289117
Suzuki D, Ueki A, Amaishi A et al (2009) Desulfovibrio portus sp. nov., a novel sulfate-reducing bacterium in the class Deltaproteobacteria isolated from an estuarine sediment. J Gen Appl Microbiol 55:125–133. https://doi.org/10.2323/jgam.55.125
Trüper HG, Schlegel HG (1964) Sulphur metabolism in Thiorhodaceae I. Quantitative measurements on growing cells of Chromatium okenii. Antonie Van Leeuwenhoek 30:225–238. https://doi.org/10.1007/BF02046728
Waite DW, Chuvochina M, Pelikan C et al (2020) Proposal to reclassify the proteobacterial classes Deltaproteobacteria and Oligoflexia, and the phylum Thermodesulfobacteria into four phyla reflecting major functional capabilities. Int J Syst Evol Microbiol 70:5972–6016. https://doi.org/10.1099/ijsem.0.004213
Weisburg WG, Barns SM, Pelletier DA et al (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703.1991
Wolin EA, Wolin MJ, Wolfe RS (1963) Formation of methane. J Franklin Inst 176:737. https://doi.org/10.1016/s0016-0032(13)90081-8
Yoon SH, Ha S, min, Lim J, et al (2017) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek, Int J Gen Mol Microbiol 110:1281–1286. https://doi.org/10.1007/s10482-017-0844-4
Zaunmüller T, Kelly DJ, Glöckner FO, Unden G (2006) Succinate dehydrogenase functioning by a reverse redox loop mechanism and fumarate reductase in sulphate-reducing bacteria. Microbiology 152:2443–2453. https://doi.org/10.1099/mic.0.28849-0
Funding
The work on isolation, physiological and taxonomic studies was supported by the Russian Science Foundation, project no. 17–74-30025. The work on genome sequencing, assembly and annotation was supported by the Ministry of Science and Higher Education of the Russian Federation. The work of A. A. K. (lipid analysis) was supported by the State Assignment 0768–2020-0007 in scientific activities in Gubkin University.
Author information
Authors and Affiliations
Contributions
AF conceived, planned and carried out experiments and wrote the original draft. AF and AS wrote the paper. AM performed genomic sequencing, assembly and annotation. AK performed lipid analysis. EB-O allowed funding acquisition. All authors provided critical feedback for the review of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Frolova, A.A., Merkel, A.Y., Kuchierskaya, A.A. et al. Pseudodesulfovibrio alkaliphilus, sp. nov., an alkaliphilic sulfate-reducing bacterium isolated from a terrestrial mud volcano. Antonie van Leeuwenhoek 114, 1387–1397 (2021). https://doi.org/10.1007/s10482-021-01608-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01608-5